"Image": models, code, and papers
Enhancing Breast Cancer Classification Using Transfer ResNet with Lightweight Attention Mechanism
Aug 25, 2023
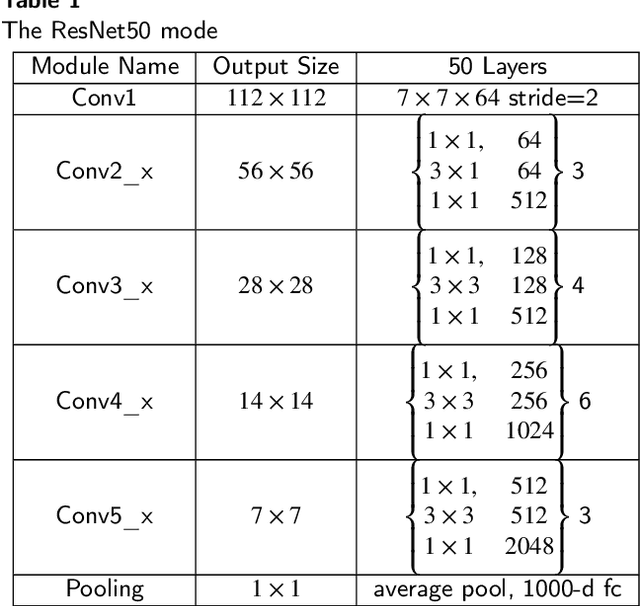
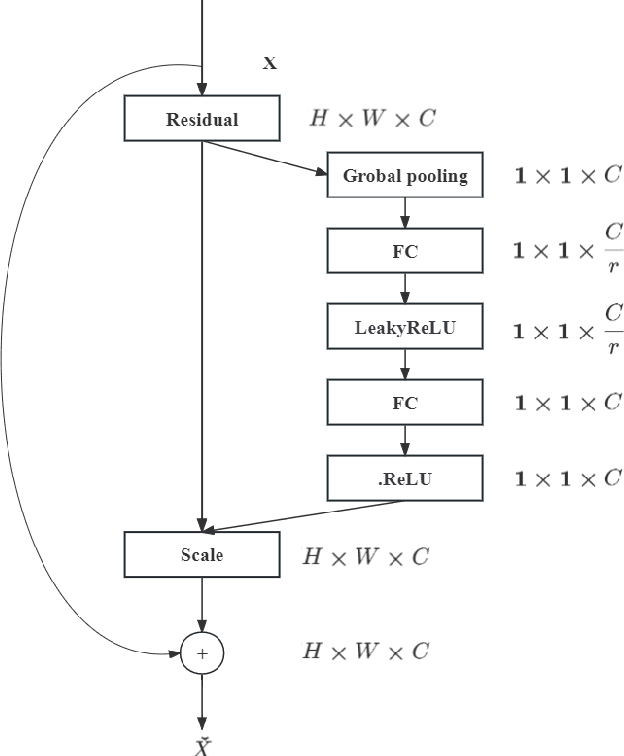
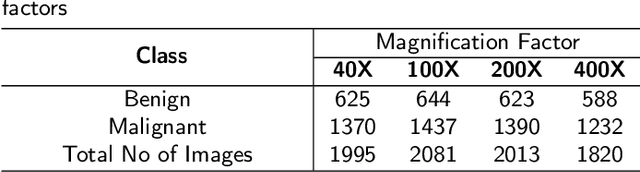
Deep learning models have revolutionized image classification by learning complex feature hierarchies in raw pixel data. This paper introduces an image classification method based on the ResNet model, and introduces a lightweight attention mechanism framework to improve performance. The framework optimizes feature representation, enhances classification capabilities, and improves feature discriminativeness. We verified the effectiveness of the algorithm on the Breakhis dataset, showing its superior performance in many aspects. Not only in terms of conventional models, our method also shows advantages on state-of-the-art methods such as contemporary visual transformers. Significant improvements have been achieved in metrics such as precision, accuracy, recall, F1-score, and G-means, while also performing well in terms of convergence time. These results strengthen the performance of the algorithm and solidify its application prospects in practical image classification tasks. Keywords: ResNet model, Lightweight attention mechanism
How You Split Matters: Data Leakage and Subject Characteristics Studies in Longitudinal Brain MRI Analysis
Sep 01, 2023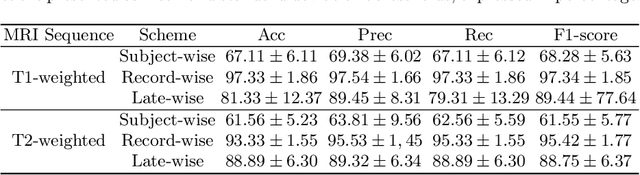
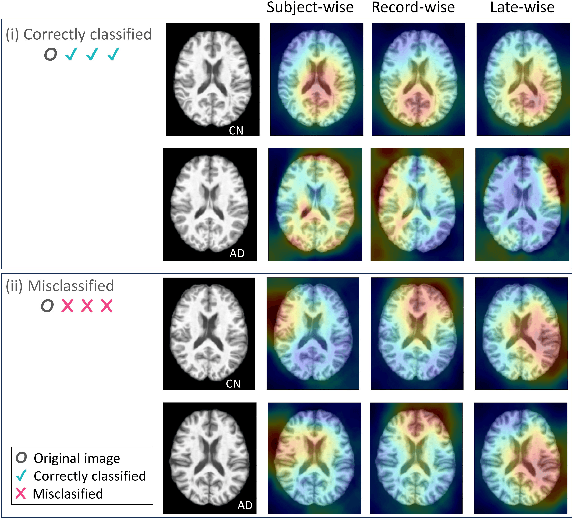
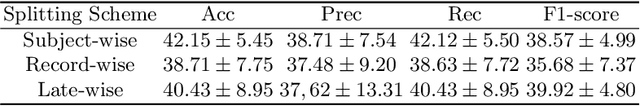
Deep learning models have revolutionized the field of medical image analysis, offering significant promise for improved diagnostics and patient care. However, their performance can be misleadingly optimistic due to a hidden pitfall called 'data leakage'. In this study, we investigate data leakage in 3D medical imaging, specifically using 3D Convolutional Neural Networks (CNNs) for brain MRI analysis. While 3D CNNs appear less prone to leakage than 2D counterparts, improper data splitting during cross-validation (CV) can still pose issues, especially with longitudinal imaging data containing repeated scans from the same subject. We explore the impact of different data splitting strategies on model performance for longitudinal brain MRI analysis and identify potential data leakage concerns. GradCAM visualization helps reveal shortcuts in CNN models caused by identity confounding, where the model learns to identify subjects along with diagnostic features. Our findings, consistent with prior research, underscore the importance of subject-wise splitting and evaluating our model further on hold-out data from different subjects to ensure the integrity and reliability of deep learning models in medical image analysis.
Shape of my heart: Cardiac models through learned signed distance functions
Aug 31, 2023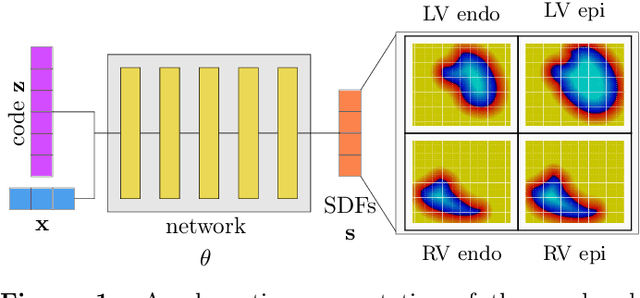
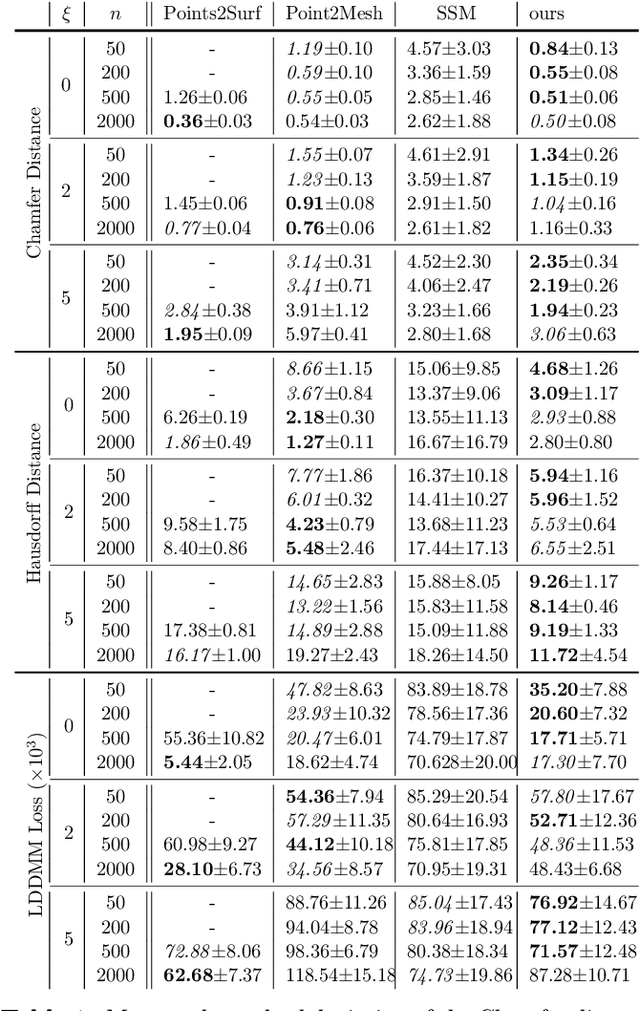
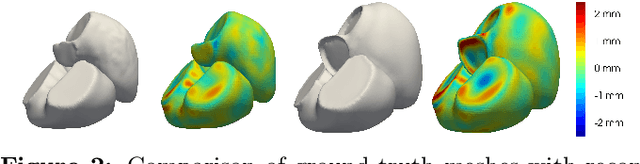
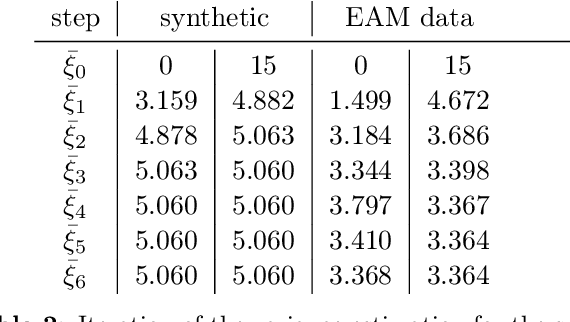
The efficient construction of an anatomical model is one of the major challenges of patient-specific in-silico models of the human heart. Current methods frequently rely on linear statistical models, allowing no advanced topological changes, or requiring medical image segmentation followed by a meshing pipeline, which strongly depends on image resolution, quality, and modality. These approaches are therefore limited in their transferability to other imaging domains. In this work, the cardiac shape is reconstructed by means of three-dimensional deep signed distance functions with Lipschitz regularity. For this purpose, the shapes of cardiac MRI reconstructions are learned from public databases to model the spatial relation of multiple chambers in Cartesian space. We demonstrate that this approach is also capable of reconstructing anatomical models from partial data, such as point clouds from a single ventricle, or modalities different from the trained MRI, such as electroanatomical mapping, and in addition, allows us to generate new anatomical shapes by randomly sampling latent vectors.
Automated COVID-19 CT Image Classification using Multi-head Channel Attention in Deep CNN
Jul 31, 2023
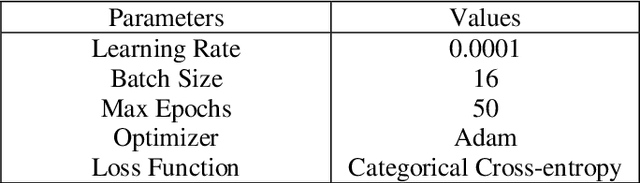

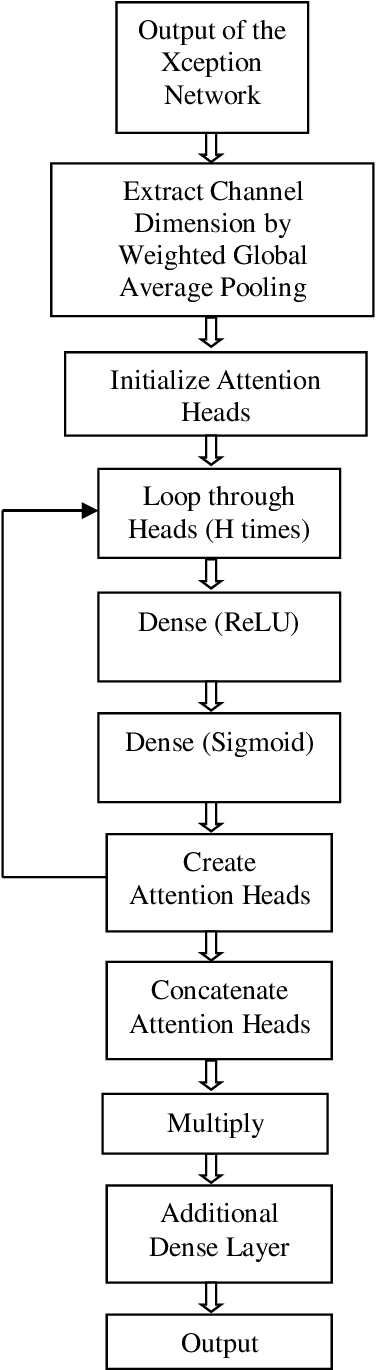

The rapid spread of COVID-19 has necessitated efficient and accurate diagnostic methods. Computed Tomography (CT) scan images have emerged as a valuable tool for detecting the disease. In this article, we present a novel deep learning approach for automated COVID-19 CT scan classification where a modified Xception model is proposed which incorporates a newly designed channel attention mechanism and weighted global average pooling to enhance feature extraction thereby improving classification accuracy. The channel attention module selectively focuses on informative regions within each channel, enabling the model to learn discriminative features for COVID-19 detection. Experiments on a widely used COVID-19 CT scan dataset demonstrate a very good accuracy of 96.99% and show its superiority to other state-of-the-art techniques. This research can contribute to the ongoing efforts in using artificial intelligence to combat current and future pandemics and can offer promising and timely solutions for efficient medical image analysis tasks.
Solving Quadratic Systems with Full-Rank Matrices Using Sparse or Generative Priors
Sep 16, 2023
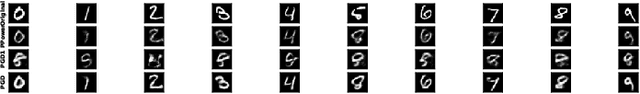
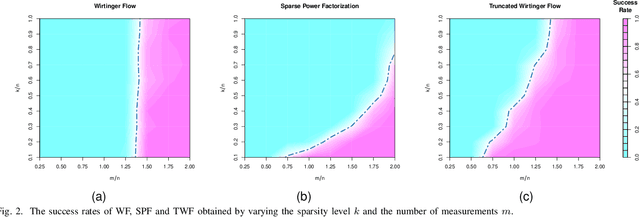

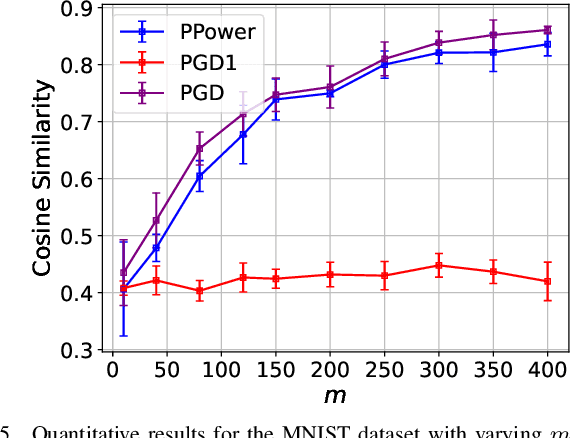
The problem of recovering a signal $\boldsymbol{x} \in \mathbb{R}^n$ from a quadratic system $\{y_i=\boldsymbol{x}^\top\boldsymbol{A}_i\boldsymbol{x},\ i=1,\ldots,m\}$ with full-rank matrices $\boldsymbol{A}_i$ frequently arises in applications such as unassigned distance geometry and sub-wavelength imaging. With i.i.d. standard Gaussian matrices $\boldsymbol{A}_i$, this paper addresses the high-dimensional case where $m\ll n$ by incorporating prior knowledge of $\boldsymbol{x}$. First, we consider a $k$-sparse $\boldsymbol{x}$ and introduce the thresholded Wirtinger flow (TWF) algorithm that does not require the sparsity level $k$. TWF comprises two steps: the spectral initialization that identifies a point sufficiently close to $\boldsymbol{x}$ (up to a sign flip) when $m=O(k^2\log n)$, and the thresholded gradient descent (with a good initialization) that produces a sequence linearly converging to $\boldsymbol{x}$ with $m=O(k\log n)$ measurements. Second, we explore the generative prior, assuming that $\boldsymbol{x}$ lies in the range of an $L$-Lipschitz continuous generative model with $k$-dimensional inputs in an $\ell_2$-ball of radius $r$. We develop the projected gradient descent (PGD) algorithm that also comprises two steps: the projected power method that provides an initial vector with $O\big(\sqrt{\frac{k \log L}{m}}\big)$ $\ell_2$-error given $m=O(k\log(Lnr))$ measurements, and the projected gradient descent that refines the $\ell_2$-error to $O(\delta)$ at a geometric rate when $m=O(k\log\frac{Lrn}{\delta^2})$. Experimental results corroborate our theoretical findings and show that: (i) our approach for the sparse case notably outperforms the existing provable algorithm sparse power factorization; (ii) leveraging the generative prior allows for precise image recovery in the MNIST dataset from a small number of quadratic measurements.
PRE: Vision-Language Prompt Learning with Reparameterization Encoder
Sep 14, 2023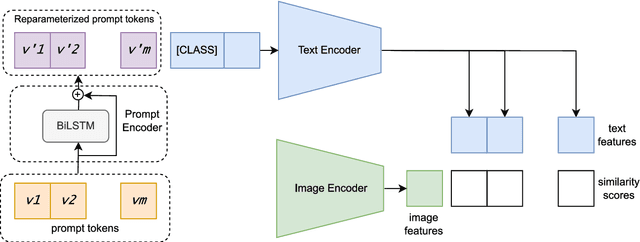
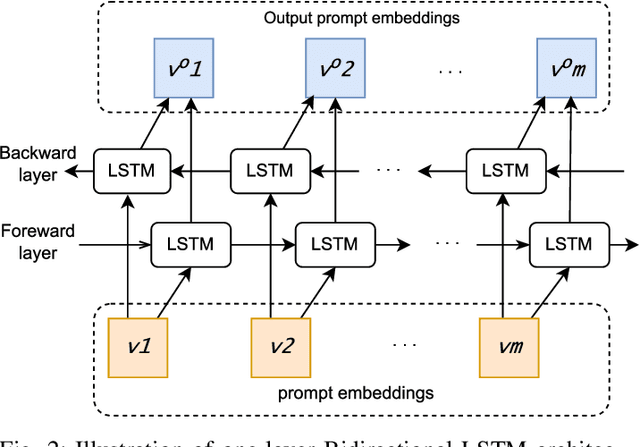
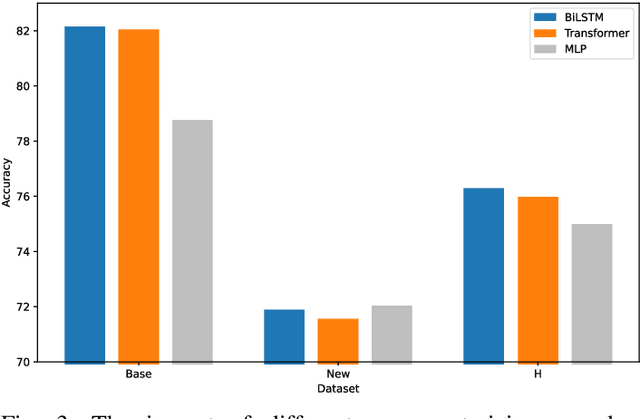
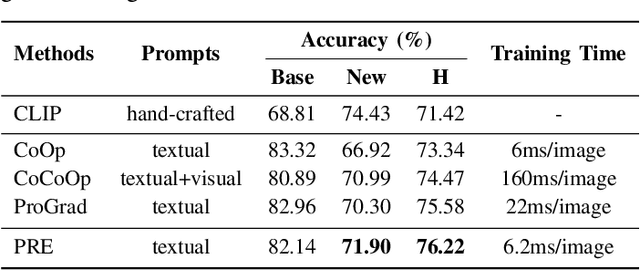
Large pre-trained vision-language models such as CLIP have demonstrated great potential in zero-shot transferability to downstream tasks. However, to attain optimal performance, the manual selection of prompts is necessary to improve alignment between the downstream image distribution and the textual class descriptions. This manual prompt engineering is the major challenge for deploying such models in practice since it requires domain expertise and is extremely time-consuming. To avoid non-trivial prompt engineering, recent work Context Optimization (CoOp) introduced the concept of prompt learning to the vision domain using learnable textual tokens. While CoOp can achieve substantial improvements over manual prompts, its learned context is worse generalizable to wider unseen classes within the same dataset. In this work, we present Prompt Learning with Reparameterization Encoder (PRE) - a simple and efficient method that enhances the generalization ability of the learnable prompt to unseen classes while maintaining the capacity to learn Base classes. Instead of directly optimizing the prompts, PRE employs a prompt encoder to reparameterize the input prompt embeddings, enhancing the exploration of task-specific knowledge from few-shot samples. Experiments and extensive ablation studies on 8 benchmarks demonstrate that our approach is an efficient method for prompt learning. Specifically, PRE achieves a notable enhancement of 5.60% in average accuracy on New classes and 3% in Harmonic mean compared to CoOp in the 16-shot setting, all achieved within a good training time.
Consistency-guided Meta-Learning for Bootstrapping Semi-Supervised Medical Image Segmentation
Jul 21, 2023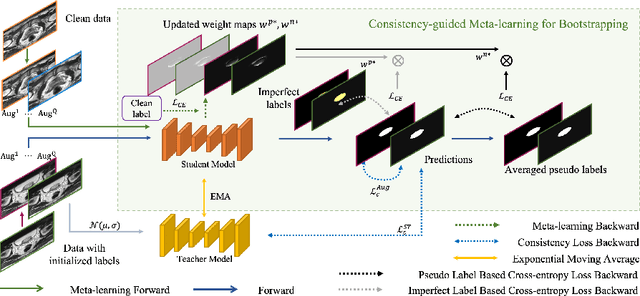
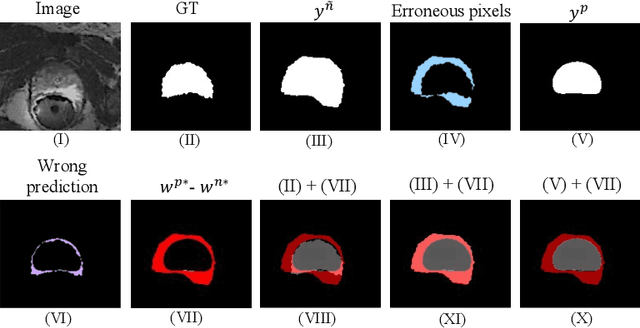
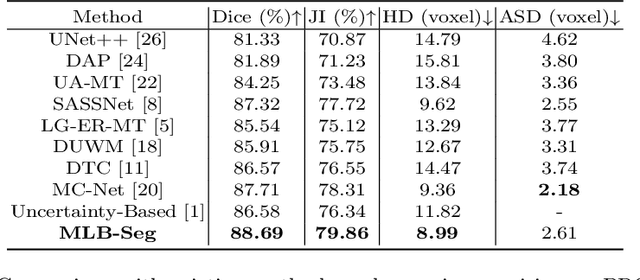
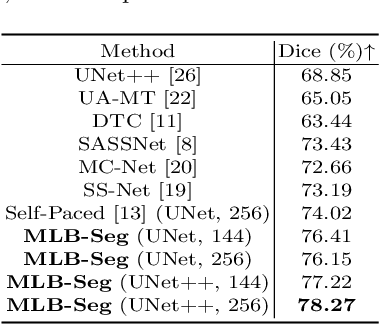

Medical imaging has witnessed remarkable progress but usually requires a large amount of high-quality annotated data which is time-consuming and costly to obtain. To alleviate this burden, semi-supervised learning has garnered attention as a potential solution. In this paper, we present Meta-Learning for Bootstrapping Medical Image Segmentation (MLB-Seg), a novel method for tackling the challenge of semi-supervised medical image segmentation. Specifically, our approach first involves training a segmentation model on a small set of clean labeled images to generate initial labels for unlabeled data. To further optimize this bootstrapping process, we introduce a per-pixel weight mapping system that dynamically assigns weights to both the initialized labels and the model's own predictions. These weights are determined using a meta-process that prioritizes pixels with loss gradient directions closer to those of clean data, which is based on a small set of precisely annotated images. To facilitate the meta-learning process, we additionally introduce a consistency-based Pseudo Label Enhancement (PLE) scheme that improves the quality of the model's own predictions by ensembling predictions from various augmented versions of the same input. In order to improve the quality of the weight maps obtained through multiple augmentations of a single input, we introduce a mean teacher into the PLE scheme. This method helps to reduce noise in the weight maps and stabilize its generation process. Our extensive experimental results on public atrial and prostate segmentation datasets demonstrate that our proposed method achieves state-of-the-art results under semi-supervision. Our code is available at https://github.com/aijinrjinr/MLB-Seg.
MOFI: Learning Image Representations from Noisy Entity Annotated Images
Jun 24, 2023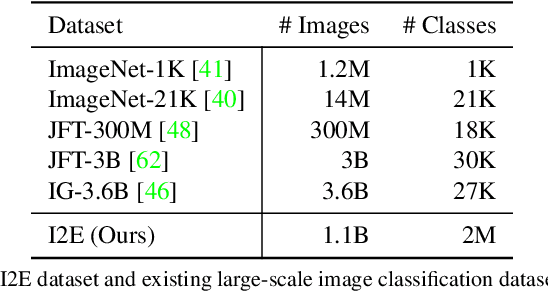

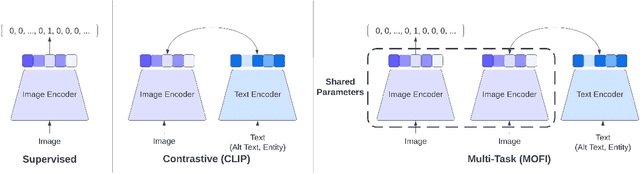
We present MOFI, a new vision foundation model designed to learn image representations from noisy entity annotated images. MOFI differs from previous work in two key aspects: ($i$) pre-training data, and ($ii$) training recipe. Regarding data, we introduce a new approach to automatically assign entity labels to images from noisy image-text pairs. Our approach involves employing a named entity recognition model to extract entities from the alt-text, and then using a CLIP model to select the correct entities as labels of the paired image. The approach is simple, does not require costly human annotation, and can be readily scaled up to billions of image-text pairs mined from the web. Through this method, we have created Image-to-Entities (I2E), a new large-scale dataset with 1 billion images and 2 million distinct entities, covering rich visual concepts in the wild. Building upon the I2E dataset, we study different training recipes, including supervised pre-training, contrastive pre-training, and multi-task learning. For constrastive pre-training, we treat entity names as free-form text, and further enrich them with entity descriptions. Experiments show that supervised pre-training with large-scale fine-grained entity labels is highly effective for image retrieval tasks, and multi-task training further improves the performance. The final MOFI model achieves 86.66% mAP on the challenging GPR1200 dataset, surpassing the previous state-of-the-art performance of 72.19% from OpenAI's CLIP model. Further experiments on zero-shot and linear probe image classification also show that MOFI outperforms a CLIP model trained on the original image-text data, demonstrating the effectiveness of the I2E dataset in learning strong image representations.
MMHQA-ICL: Multimodal In-context Learning for Hybrid Question Answering over Text, Tables and Images
Sep 09, 2023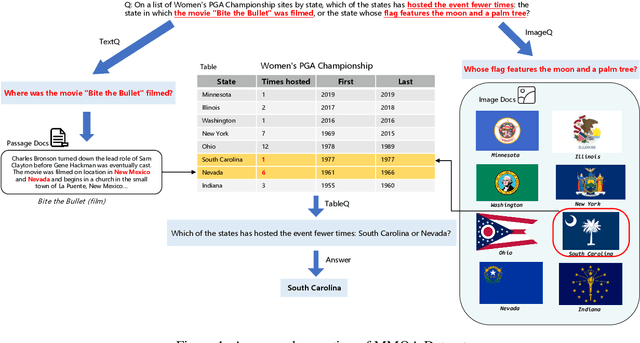
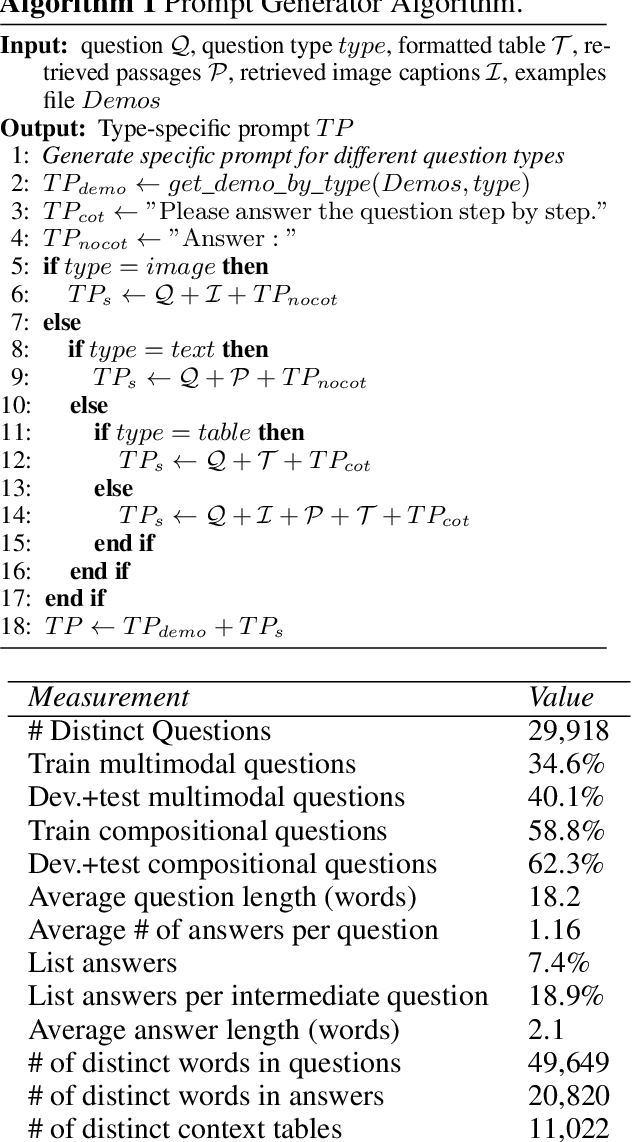
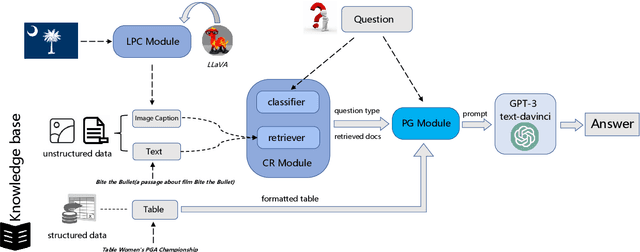

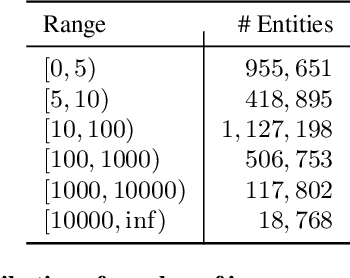
In the real world, knowledge often exists in a multimodal and heterogeneous form. Addressing the task of question answering with hybrid data types, including text, tables, and images, is a challenging task (MMHQA). Recently, with the rise of large language models (LLM), in-context learning (ICL) has become the most popular way to solve QA problems. We propose MMHQA-ICL framework for addressing this problems, which includes stronger heterogeneous data retriever and an image caption module. Most importantly, we propose a Type-specific In-context Learning Strategy for MMHQA, enabling LLMs to leverage their powerful performance in this task. We are the first to use end-to-end LLM prompting method for this task. Experimental results demonstrate that our framework outperforms all baselines and methods trained on the full dataset, achieving state-of-the-art results under the few-shot setting on the MultimodalQA dataset.
HAct: Out-of-Distribution Detection with Neural Net Activation Histograms
Sep 09, 2023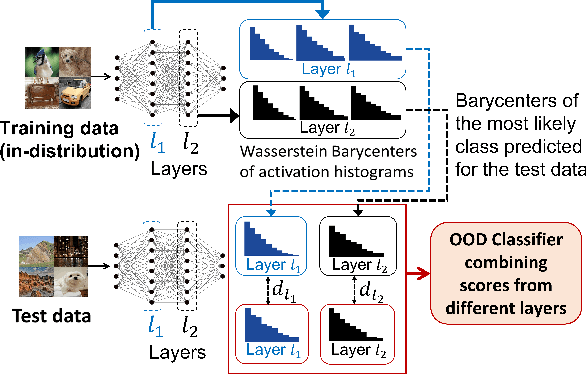
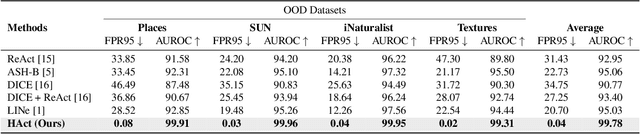
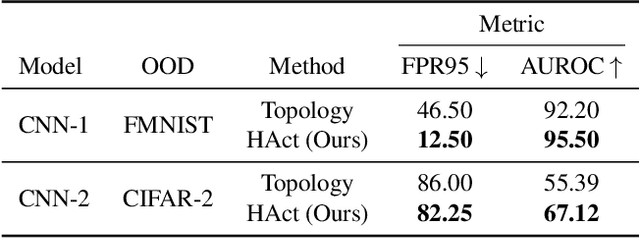
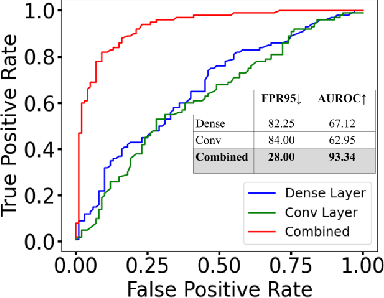
We propose a simple, efficient, and accurate method for detecting out-of-distribution (OOD) data for trained neural networks, a potential first step in methods for OOD generalization. We propose a novel descriptor, HAct - activation histograms, for OOD detection, that is, probability distributions (approximated by histograms) of output values of neural network layers under the influence of incoming data. We demonstrate that HAct is significantly more accurate than state-of-the-art on multiple OOD image classification benchmarks. For instance, our approach achieves a true positive rate (TPR) of 95% with only 0.05% false-positives using Resnet-50 on standard OOD benchmarks, outperforming previous state-of-the-art by 20.66% in the false positive rate (at the same TPR of 95%). The low computational complexity and the ease of implementation make HAct suitable for online implementation in monitoring deployed neural networks in practice at scale.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge