Alexander Effland
MotionDPS: Motion-Compensated 3D Brain MRI Reconstruction
May 21, 2026Abstract:Magnetic resonance imaging (MRI) is highly susceptible to patient motion due to its relatively long acquisition times and the fact that data are acquired sequentially in k-space. Even small patient movements introduce phase inconsistencies across measurements, leading to severe artifacts such as blurring, ghosting, and geometric distortions that can compromise diagnostic quality. Retrospective motion compensation remains challenging, particularly in accelerated acquisitions, due to the ill-posed nature of the joint reconstruction and motion estimation problem. In this work, we propose a unified Bayesian framework for motion-compensated 3D MRI that jointly estimates the anatomical image, rigid-body motion parameters, and coil sensitivity maps directly from motion-corrupted k-space data. Our approach integrates pretrained 3D complex-valued score-based diffusion models as expressive anatomical image priors within a physics-based forward model. Inference is performed by alternating diffusion posterior image updates with efficient proximal optimization steps for motion and coil sensitivity estimation, enabling fully unsupervised reconstruction without the need for paired motion-free training data. Experiments on simulated and real-motion brain MRI datasets demonstrate that the proposed method achieves improved image quality and motion robustness compared to state-of-the-art classical and learning-based motion correction techniques, particularly in the presence of severe motion and high acceleration.
Semantic Segmentation for Histopathology using Learned Regularization based on Global Proportions
Apr 27, 2026Abstract:In pathology, the spatial distribution and proportions of tissue types are key indicators of disease progression, and are more readily available than fine-grained annotations. However, these assessments are rarely mapped to pixel-wise segmentation. The task is fundamentally underdetermined, as many spatially distinct segmentations can satisfy the same global proportions in the absence of pixel-wise constraints. To address this, we introduce Variational Segmentation from Label Proportions (VSLP), a two-stage framework that infers dense segmentations from global label proportions, without any pixel-level annotations. This framework first leverages a pre-trained transformer model with test-time augmentation to produce a pixel-wise confidence estimate. In the second stage, these estimates are fused by solving a variational optimization problem that incorporates a Wasserstein data fidelity term alongside a learned regularizer. Unlike end-to-end networks, our variational method can visualize the fidelity-regularization energy, resulting in more interpretable segmentation. We validate our approach on two public datasets, achieving superior performance over existing weakly supervised and unsupervised methods. For one of these datasets, proportions have been estimated by an experienced pathologist to provide a realistic benchmark to the community. Furthermore, the method scales to an in-house dataset with noisy pathologist labels, severely outperforming state-of-the-art methods, thereby demonstrating practical applicability. The code and data will be made publicly available upon acceptance at https://github.com/xiaoliangpi/VSLP.
Learned Finite Element-based Regularization of the Inverse Problem in Electrocardiographic Imaging
Feb 07, 2026Abstract:Electrocardiographic imaging (ECGI) seeks to reconstruct cardiac electrical activity from body-surface potentials noninvasively. However, the associated inverse problem is severely ill-posed and requires robust regularization. While classical approaches primarily employ spatial smoothing, the temporal structure of cardiac dynamics remains underexploited despite its physiological relevance. We introduce a space-time regularization framework that couples spatial regularization with a learned temporal Fields-of-Experts (FoE) prior to capture complex spatiotemporal activation patterns. We derive a finite element discretization on unstructured cardiac surface meshes, prove Mosco-convergence, and develop a scalable optimization algorithm capable of handling the FoE term. Numerical experiments on synthetic epicardial data demonstrate improved denoising and inverse reconstructions compared to handcrafted spatiotemporal methods, yielding solutions that are both robust to noise and physiologically plausible.
Finite element-based space-time total variation-type regularization of the inverse problem in electrocardiographic imaging
Aug 21, 2024



Abstract:Reconstructing cardiac electrical activity from body surface electric potential measurements results in the severely ill-posed inverse problem in electrocardiography. Many different regularization approaches have been proposed to improve numerical results and provide unique results. This work presents a novel approach for reconstructing the epicardial potential from body surface potential maps based on a space-time total variation-type regularization using finite elements, where a first-order primal-dual algorithm solves the underlying convex optimization problem. In several numerical experiments, the superior performance of this method and the benefit of space-time regularization for the reconstruction of epicardial potential on two-dimensional torso data and a three-dimensional rabbit heart compared to state-of-the-art methods are demonstrated.
Gadolinium dose reduction for brain MRI using conditional deep learning
Mar 06, 2024



Abstract:Recently, deep learning (DL)-based methods have been proposed for the computational reduction of gadolinium-based contrast agents (GBCAs) to mitigate adverse side effects while preserving diagnostic value. Currently, the two main challenges for these approaches are the accurate prediction of contrast enhancement and the synthesis of realistic images. In this work, we address both challenges by utilizing the contrast signal encoded in the subtraction images of pre-contrast and post-contrast image pairs. To avoid the synthesis of any noise or artifacts and solely focus on contrast signal extraction and enhancement from low-dose subtraction images, we train our DL model using noise-free standard-dose subtraction images as targets. As a result, our model predicts the contrast enhancement signal only; thereby enabling synthesization of images beyond the standard dose. Furthermore, we adapt the embedding idea of recent diffusion-based models to condition our model on physical parameters affecting the contrast enhancement behavior. We demonstrate the effectiveness of our approach on synthetic and real datasets using various scanners, field strengths, and contrast agents.
Shape of my heart: Cardiac models through learned signed distance functions
Sep 05, 2023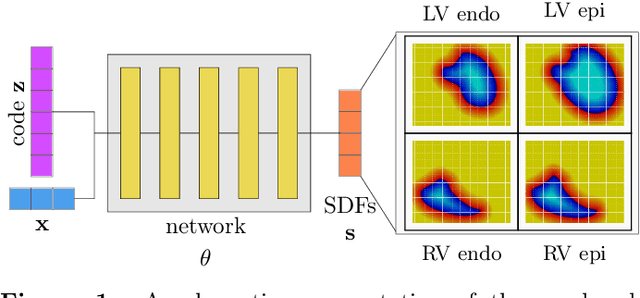
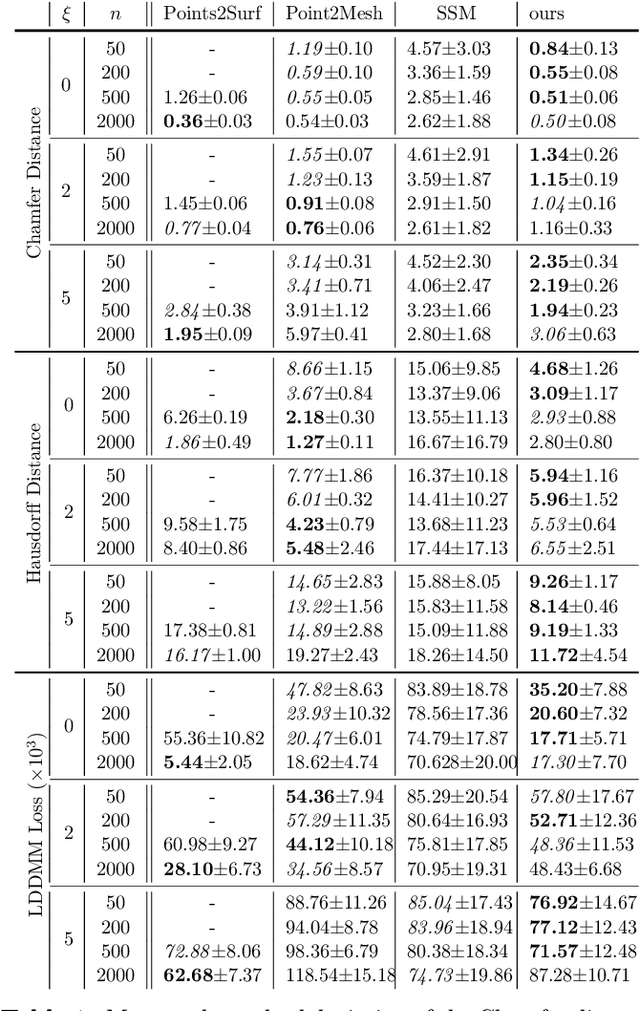
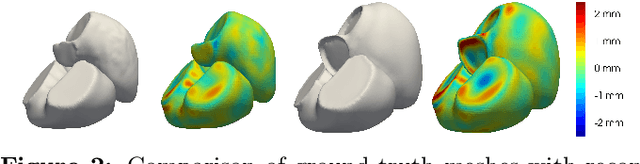
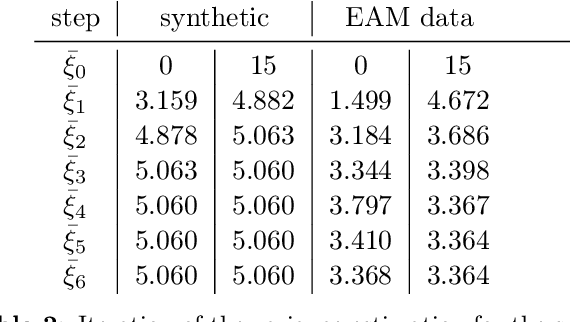
Abstract:The efficient construction of an anatomical model is one of the major challenges of patient-specific in-silico models of the human heart. Current methods frequently rely on linear statistical models, allowing no advanced topological changes, or requiring medical image segmentation followed by a meshing pipeline, which strongly depends on image resolution, quality, and modality. These approaches are therefore limited in their transferability to other imaging domains. In this work, the cardiac shape is reconstructed by means of three-dimensional deep signed distance functions with Lipschitz regularity. For this purpose, the shapes of cardiac MRI reconstructions are learned from public databases to model the spatial relation of multiple chambers in Cartesian space. We demonstrate that this approach is also capable of reconstructing anatomical models from partial data, such as point clouds from a single ventricle, or modalities different from the trained MRI, such as electroanatomical mapping, and in addition, allows us to generate new anatomical shapes by randomly sampling latent vectors.
Digital twinning of cardiac electrophysiology models from the surface ECG: a geodesic backpropagation approach
Aug 16, 2023Abstract:The eikonal equation has become an indispensable tool for modeling cardiac electrical activation accurately and efficiently. In principle, by matching clinically recorded and eikonal-based electrocardiograms (ECGs), it is possible to build patient-specific models of cardiac electrophysiology in a purely non-invasive manner. Nonetheless, the fitting procedure remains a challenging task. The present study introduces a novel method, Geodesic-BP, to solve the inverse eikonal problem. Geodesic-BP is well-suited for GPU-accelerated machine learning frameworks, allowing us to optimize the parameters of the eikonal equation to reproduce a given ECG. We show that Geodesic-BP can reconstruct a simulated cardiac activation with high accuracy in a synthetic test case, even in the presence of modeling inaccuracies. Furthermore, we apply our algorithm to a publicly available dataset of a rabbit model, with very positive results. Given the future shift towards personalized medicine, Geodesic-BP has the potential to help in future functionalizations of cardiac models meeting clinical time constraints while maintaining the physiological accuracy of state-of-the-art cardiac models.
Faithful Synthesis of Low-dose Contrast-enhanced Brain MRI Scans using Noise-preserving Conditional GANs
Jun 26, 2023



Abstract:Today Gadolinium-based contrast agents (GBCA) are indispensable in Magnetic Resonance Imaging (MRI) for diagnosing various diseases. However, GBCAs are expensive and may accumulate in patients with potential side effects, thus dose-reduction is recommended. Still, it is unclear to which extent the GBCA dose can be reduced while preserving the diagnostic value -- especially in pathological regions. To address this issue, we collected brain MRI scans at numerous non-standard GBCA dosages and developed a conditional GAN model for synthesizing corresponding images at fractional dose levels. Along with the adversarial loss, we advocate a novel content loss function based on the Wasserstein distance of locally paired patch statistics for the faithful preservation of noise. Our numerical experiments show that conditional GANs are suitable for generating images at different GBCA dose levels and can be used to augment datasets for virtual contrast models. Moreover, our model can be transferred to openly available datasets such as BraTS, where non-standard GBCA dosage images do not exist.
Bayesian Uncertainty Estimation of Learned Variational MRI Reconstruction
Feb 12, 2021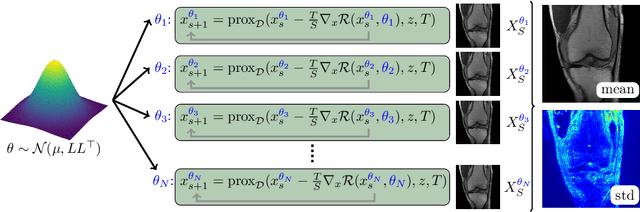
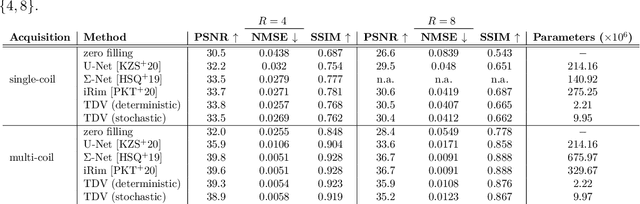
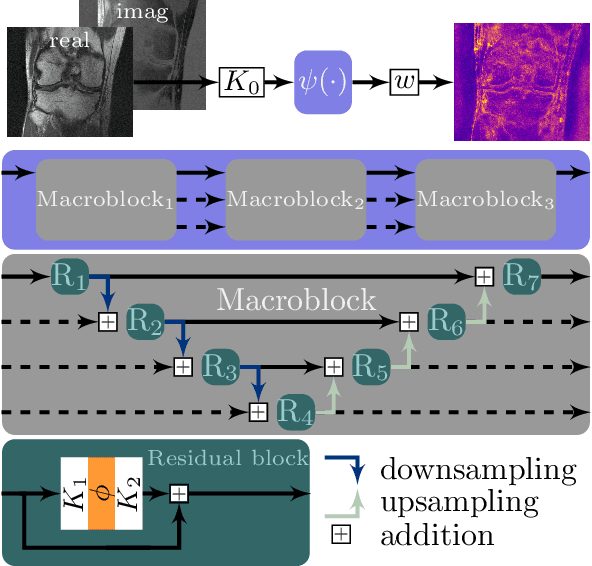
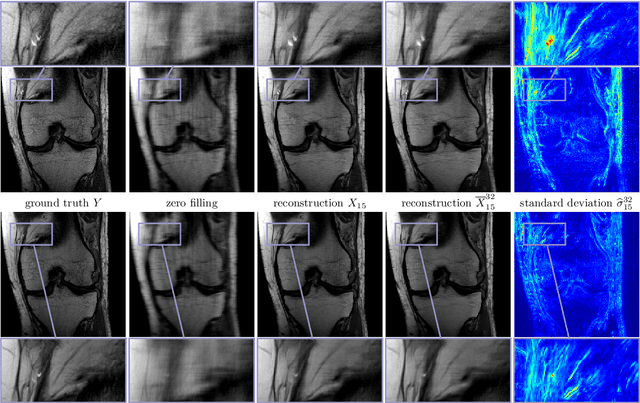
Abstract:Recent deep learning approaches focus on improving quantitative scores of dedicated benchmarks, and therefore only reduce the observation-related (aleatoric) uncertainty. However, the model-immanent (epistemic) uncertainty is less frequently systematically analyzed. In this work, we introduce a Bayesian variational framework to quantify the epistemic uncertainty. To this end, we solve the linear inverse problem of undersampled MRI reconstruction in a variational setting. The associated energy functional is composed of a data fidelity term and the total deep variation (TDV) as a learned parametric regularizer. To estimate the epistemic uncertainty we draw the parameters of the TDV regularizer from a multivariate Gaussian distribution, whose mean and covariance matrix are learned in a stochastic optimal control problem. In several numerical experiments, we demonstrate that our approach yields competitive results for undersampled MRI reconstruction. Moreover, we can accurately quantify the pixelwise epistemic uncertainty, which can serve radiologists as an additional resource to visualize reconstruction reliability.
Shared Prior Learning of Energy-Based Models for Image Reconstruction
Nov 13, 2020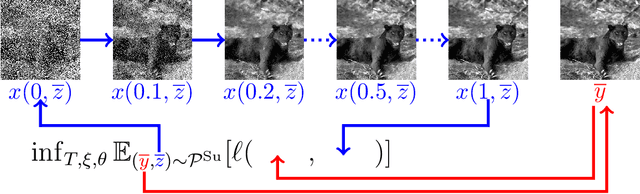
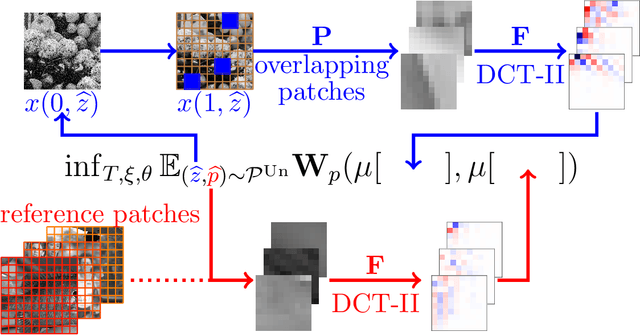
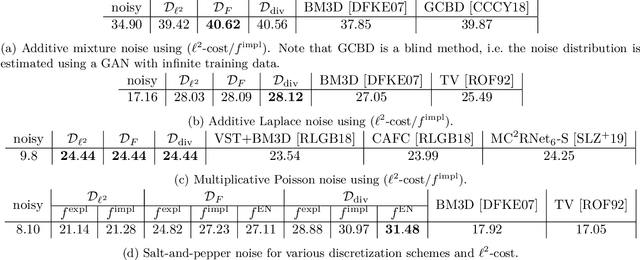
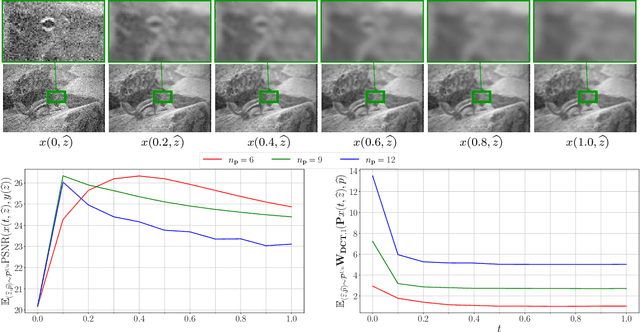
Abstract:We propose a novel learning-based framework for image reconstruction particularly designed for training without ground truth data, which has three major building blocks: energy-based learning, a patch-based Wasserstein loss functional, and shared prior learning. In energy-based learning, the parameters of an energy functional composed of a learned data fidelity term and a data-driven regularizer are computed in a mean-field optimal control problem. In the absence of ground truth data, we change the loss functional to a patch-based Wasserstein functional, in which local statistics of the output images are compared to uncorrupted reference patches. Finally, in shared prior learning, both aforementioned optimal control problems are optimized simultaneously with shared learned parameters of the regularizer to further enhance unsupervised image reconstruction. We derive several time discretization schemes of the gradient flow and verify their consistency in terms of Mosco convergence. In numerous numerical experiments, we demonstrate that the proposed method generates state-of-the-art results for various image reconstruction applications--even if no ground truth images are available for training.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge