"Image": models, code, and papers
Generalizing deep learning models for medical image classification
Mar 21, 2024
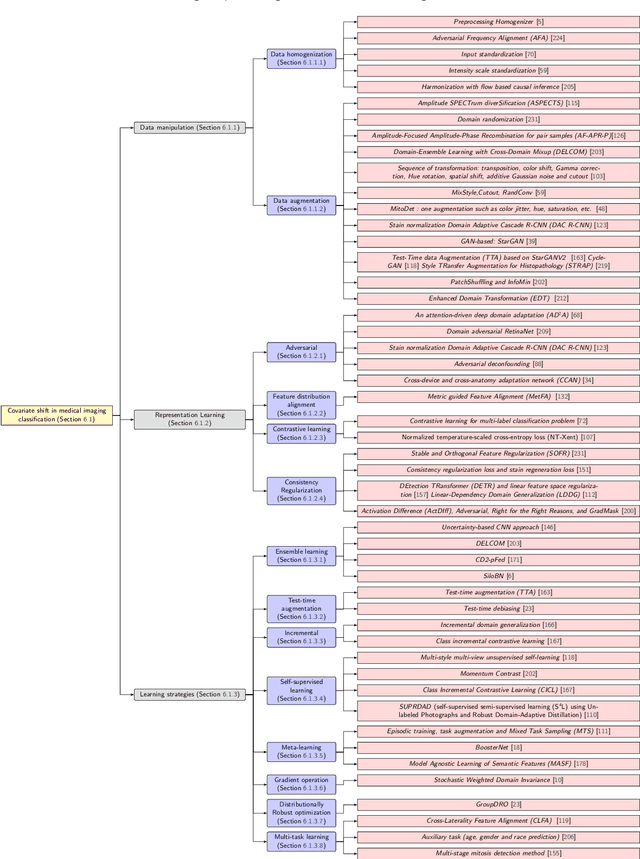
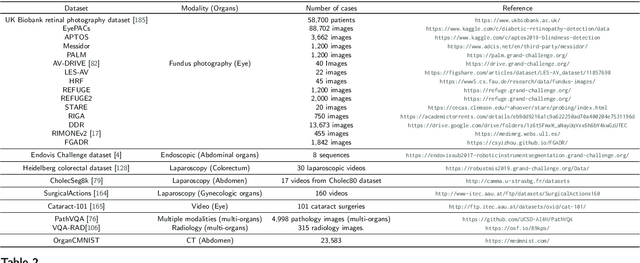
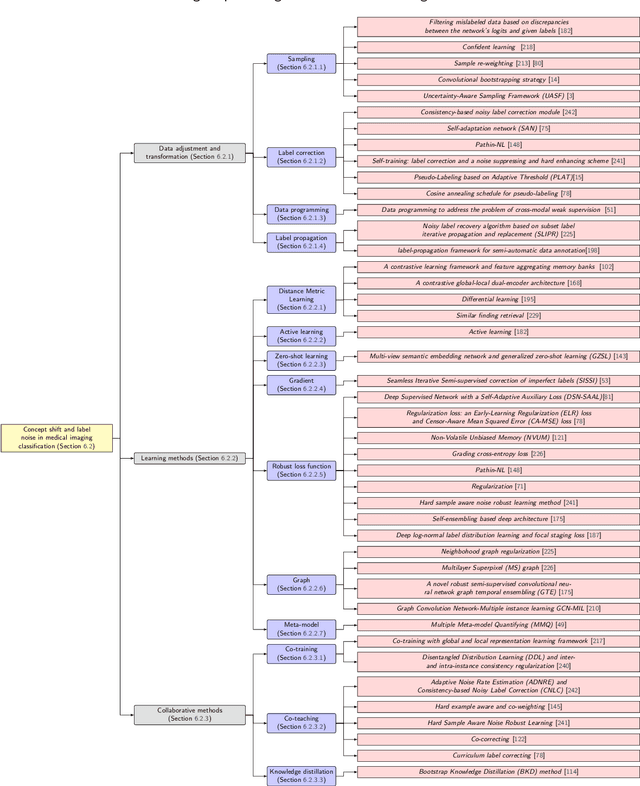
Numerous Deep Learning (DL) models have been developed for a large spectrum of medical image analysis applications, which promises to reshape various facets of medical practice. Despite early advances in DL model validation and implementation, which encourage healthcare institutions to adopt them, some fundamental questions remain: are the DL models capable of generalizing? What causes a drop in DL model performances? How to overcome the DL model performance drop? Medical data are dynamic and prone to domain shift, due to multiple factors such as updates to medical equipment, new imaging workflow, and shifts in patient demographics or populations can induce this drift over time. In this paper, we review recent developments in generalization methods for DL-based classification models. We also discuss future challenges, including the need for improved evaluation protocols and benchmarks, and envisioned future developments to achieve robust, generalized models for medical image classification.
Smooth Deep Saliency
Apr 04, 2024In this work, we investigate methods to reduce the noise in deep saliency maps coming from convolutional downsampling, with the purpose of explaining how a deep learning model detects tumors in scanned histological tissue samples. Those methods make the investigated models more interpretable for gradient-based saliency maps, computed in hidden layers. We test our approach on different models trained for image classification on ImageNet1K, and models trained for tumor detection on Camelyon16 and in-house real-world digital pathology scans of stained tissue samples. Our results show that the checkerboard noise in the gradient gets reduced, resulting in smoother and therefore easier to interpret saliency maps.
Imaging transformer for MRI denoising with the SNR unit training: enabling generalization across field-strengths, imaging contrasts, and anatomy
Apr 03, 2024The ability to recover MRI signal from noise is key to achieve fast acquisition, accurate quantification, and high image quality. Past work has shown convolutional neural networks can be used with abundant and paired low and high-SNR images for training. However, for applications where high-SNR data is difficult to produce at scale (e.g. with aggressive acceleration, high resolution, or low field strength), training a new denoising network using a large quantity of high-SNR images can be infeasible. In this study, we overcome this limitation by improving the generalization of denoising models, enabling application to many settings beyond what appears in the training data. Specifically, we a) develop a training scheme that uses complex MRIs reconstructed in the SNR units (i.e., the images have a fixed noise level, SNR unit training) and augments images with realistic noise based on coil g-factor, and b) develop a novel imaging transformer (imformer) to handle 2D, 2D+T, and 3D MRIs in one model architecture. Through empirical evaluation, we show this combination improves performance compared to CNN models and improves generalization, enabling a denoising model to be used across field-strengths, image contrasts, and anatomy.
MMCert: Provable Defense against Adversarial Attacks to Multi-modal Models
Apr 02, 2024Different from a unimodal model whose input is from a single modality, the input (called multi-modal input) of a multi-modal model is from multiple modalities such as image, 3D points, audio, text, etc. Similar to unimodal models, many existing studies show that a multi-modal model is also vulnerable to adversarial perturbation, where an attacker could add small perturbation to all modalities of a multi-modal input such that the multi-modal model makes incorrect predictions for it. Existing certified defenses are mostly designed for unimodal models, which achieve sub-optimal certified robustness guarantees when extended to multi-modal models as shown in our experimental results. In our work, we propose MMCert, the first certified defense against adversarial attacks to a multi-modal model. We derive a lower bound on the performance of our MMCert under arbitrary adversarial attacks with bounded perturbations to both modalities (e.g., in the context of auto-driving, we bound the number of changed pixels in both RGB image and depth image). We evaluate our MMCert using two benchmark datasets: one for the multi-modal road segmentation task and the other for the multi-modal emotion recognition task. Moreover, we compare our MMCert with a state-of-the-art certified defense extended from unimodal models. Our experimental results show that our MMCert outperforms the baseline.
Effective Lymph Nodes Detection in CT Scans Using Location Debiased Query Selection and Contrastive Query Representation in Transformer
Apr 04, 2024Lymph node (LN) assessment is a critical, indispensable yet very challenging task in the routine clinical workflow of radiology and oncology. Accurate LN analysis is essential for cancer diagnosis, staging, and treatment planning. Finding scatteredly distributed, low-contrast clinically relevant LNs in 3D CT is difficult even for experienced physicians under high inter-observer variations. Previous automatic LN detection works typically yield limited recall and high false positives (FPs) due to adjacent anatomies with similar image intensities, shapes, or textures (vessels, muscles, esophagus, etc). In this work, we propose a new LN DEtection TRansformer, named LN-DETR, to achieve more accurate performance. By enhancing the 2D backbone with a multi-scale 2.5D feature fusion to incorporate 3D context explicitly, more importantly, we make two main contributions to improve the representation quality of LN queries. 1) Considering that LN boundaries are often unclear, an IoU prediction head and a location debiased query selection are proposed to select LN queries of higher localization accuracy as the decoder query's initialization. 2) To reduce FPs, query contrastive learning is employed to explicitly reinforce LN queries towards their best-matched ground-truth queries over unmatched query predictions. Trained and tested on 3D CT scans of 1067 patients (with 10,000+ labeled LNs) via combining seven LN datasets from different body parts (neck, chest, and abdomen) and pathologies/cancers, our method significantly improves the performance of previous leading methods by > 4-5% average recall at the same FP rates in both internal and external testing. We further evaluate on the universal lesion detection task using NIH DeepLesion benchmark, and our method achieves the top performance of 88.46% averaged recall across 0.5 to 4 FPs per image, compared with other leading reported results.
Autonomous Artificial Intelligence Agents for Clinical Decision Making in Oncology
Apr 06, 2024Multimodal artificial intelligence (AI) systems have the potential to enhance clinical decision-making by interpreting various types of medical data. However, the effectiveness of these models across all medical fields is uncertain. Each discipline presents unique challenges that need to be addressed for optimal performance. This complexity is further increased when attempting to integrate different fields into a single model. Here, we introduce an alternative approach to multimodal medical AI that utilizes the generalist capabilities of a large language model (LLM) as a central reasoning engine. This engine autonomously coordinates and deploys a set of specialized medical AI tools. These tools include text, radiology and histopathology image interpretation, genomic data processing, web searches, and document retrieval from medical guidelines. We validate our system across a series of clinical oncology scenarios that closely resemble typical patient care workflows. We show that the system has a high capability in employing appropriate tools (97%), drawing correct conclusions (93.6%), and providing complete (94%), and helpful (89.2%) recommendations for individual patient cases while consistently referencing relevant literature (82.5%) upon instruction. This work provides evidence that LLMs can effectively plan and execute domain-specific models to retrieve or synthesize new information when used as autonomous agents. This enables them to function as specialist, patient-tailored clinical assistants. It also simplifies regulatory compliance by allowing each component tool to be individually validated and approved. We believe, that our work can serve as a proof-of-concept for more advanced LLM-agents in the medical domain.
Segmentation-Guided Knee Radiograph Generation using Conditional Diffusion Models
Apr 04, 2024Deep learning-based medical image processing algorithms require representative data during development. In particular, surgical data might be difficult to obtain, and high-quality public datasets are limited. To overcome this limitation and augment datasets, a widely adopted solution is the generation of synthetic images. In this work, we employ conditional diffusion models to generate knee radiographs from contour and bone segmentations. Remarkably, two distinct strategies are presented by incorporating the segmentation as a condition into the sampling and training process, namely, conditional sampling and conditional training. The results demonstrate that both methods can generate realistic images while adhering to the conditioning segmentation. The conditional training method outperforms the conditional sampling method and the conventional U-Net.
Super-Resolution Analysis for Landfill Waste Classification
Apr 02, 2024Illegal landfills are a critical issue due to their environmental, economic, and public health impacts. This study leverages aerial imagery for environmental crime monitoring. While advances in artificial intelligence and computer vision hold promise, the challenge lies in training models with high-resolution literature datasets and adapting them to open-access low-resolution images. Considering the substantial quality differences and limited annotation, this research explores the adaptability of models across these domains. Motivated by the necessity for a comprehensive evaluation of waste detection algorithms, it advocates cross-domain classification and super-resolution enhancement to analyze the impact of different image resolutions on waste classification as an evaluation to combat the proliferation of illegal landfills. We observed performance improvements by enhancing image quality but noted an influence on model sensitivity, necessitating careful threshold fine-tuning.
GenN2N: Generative NeRF2NeRF Translation
Apr 03, 2024We present GenN2N, a unified NeRF-to-NeRF translation framework for various NeRF translation tasks such as text-driven NeRF editing, colorization, super-resolution, inpainting, etc. Unlike previous methods designed for individual translation tasks with task-specific schemes, GenN2N achieves all these NeRF editing tasks by employing a plug-and-play image-to-image translator to perform editing in the 2D domain and lifting 2D edits into the 3D NeRF space. Since the 3D consistency of 2D edits may not be assured, we propose to model the distribution of the underlying 3D edits through a generative model that can cover all possible edited NeRFs. To model the distribution of 3D edited NeRFs from 2D edited images, we carefully design a VAE-GAN that encodes images while decoding NeRFs. The latent space is trained to align with a Gaussian distribution and the NeRFs are supervised through an adversarial loss on its renderings. To ensure the latent code does not depend on 2D viewpoints but truly reflects the 3D edits, we also regularize the latent code through a contrastive learning scheme. Extensive experiments on various editing tasks show GenN2N, as a universal framework, performs as well or better than task-specific specialists while possessing flexible generative power. More results on our project page: https://xiangyueliu.github.io/GenN2N/
Flickr30K-CFQ: A Compact and Fragmented Query Dataset for Text-image Retrieval
Mar 20, 2024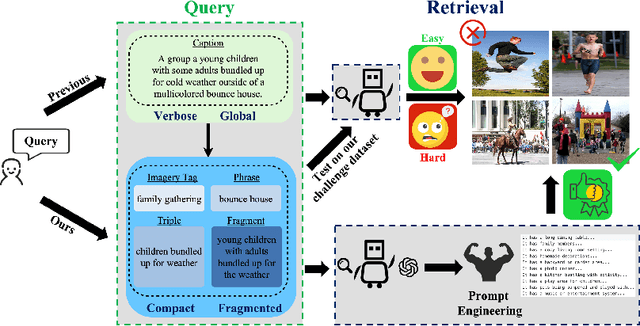

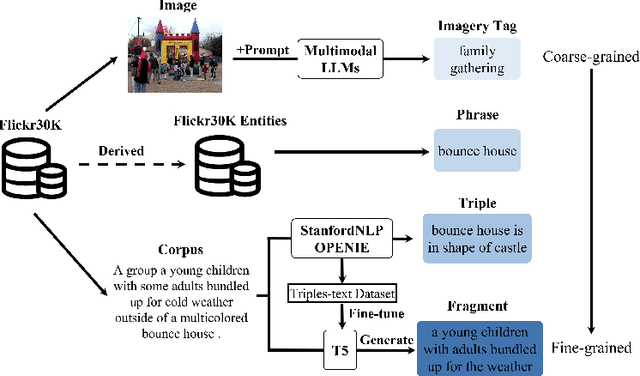

With the explosive growth of multi-modal information on the Internet, unimodal search cannot satisfy the requirement of Internet applications. Text-image retrieval research is needed to realize high-quality and efficient retrieval between different modalities. Existing text-image retrieval research is mostly based on general vision-language datasets (e.g. MS-COCO, Flickr30K), in which the query utterance is rigid and unnatural (i.e. verbosity and formality). To overcome the shortcoming, we construct a new Compact and Fragmented Query challenge dataset (named Flickr30K-CFQ) to model text-image retrieval task considering multiple query content and style, including compact and fine-grained entity-relation corpus. We propose a novel query-enhanced text-image retrieval method using prompt engineering based on LLM. Experiments show that our proposed Flickr30-CFQ reveals the insufficiency of existing vision-language datasets in realistic text-image tasks. Our LLM-based Query-enhanced method applied on different existing text-image retrieval models improves query understanding performance both on public dataset and our challenge set Flickr30-CFQ with over 0.9% and 2.4% respectively. Our project can be available anonymously in https://sites.google.com/view/Flickr30K-cfq.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge