Yunguan Fu
Image quality assessment by overlapping task-specific and task-agnostic measures: application to prostate multiparametric MR images for cancer segmentation
Feb 20, 2022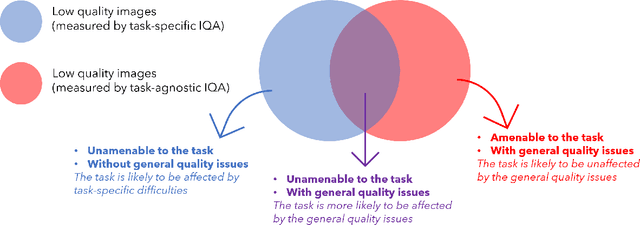
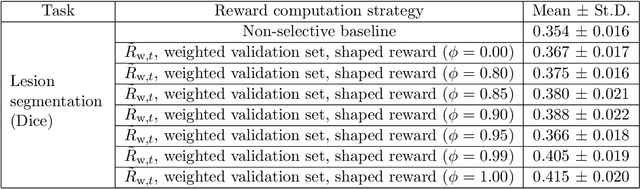

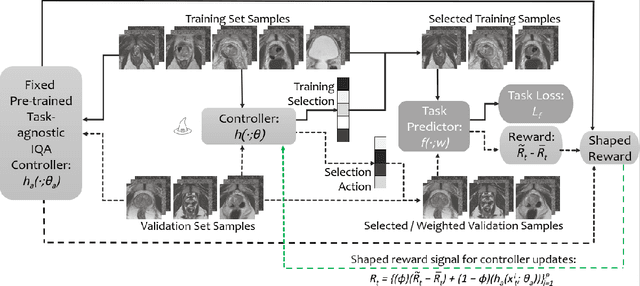
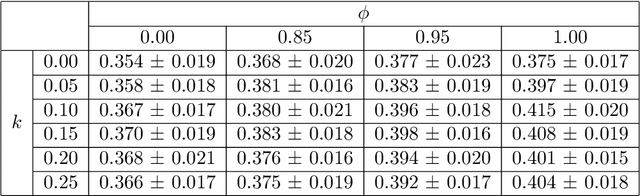
Abstract:Image quality assessment (IQA) in medical imaging can be used to ensure that downstream clinical tasks can be reliably performed. Quantifying the impact of an image on the specific target tasks, also named as task amenability, is needed. A task-specific IQA has recently been proposed to learn an image-amenability-predicting controller simultaneously with a target task predictor. This allows for the trained IQA controller to measure the impact an image has on the target task performance, when this task is performed using the predictor, e.g. segmentation and classification neural networks in modern clinical applications. In this work, we propose an extension to this task-specific IQA approach, by adding a task-agnostic IQA based on auto-encoding as the target task. Analysing the intersection between low-quality images, deemed by both the task-specific and task-agnostic IQA, may help to differentiate the underpinning factors that caused the poor target task performance. For example, common imaging artefacts may not adversely affect the target task, which would lead to a low task-agnostic quality and a high task-specific quality, whilst individual cases considered clinically challenging, which can not be improved by better imaging equipment or protocols, is likely to result in a high task-agnostic quality but a low task-specific quality. We first describe a flexible reward shaping strategy which allows for the adjustment of weighting between task-agnostic and task-specific quality scoring. Furthermore, we evaluate the proposed algorithm using a clinically challenging target task of prostate tumour segmentation on multiparametric magnetic resonance (mpMR) images, from 850 patients. The proposed reward shaping strategy, with appropriately weighted task-specific and task-agnostic qualities, successfully identified samples that need re-acquisition due to defected imaging process.
Few-shot image segmentation for cross-institution male pelvic organs using registration-assisted prototypical learning
Jan 17, 2022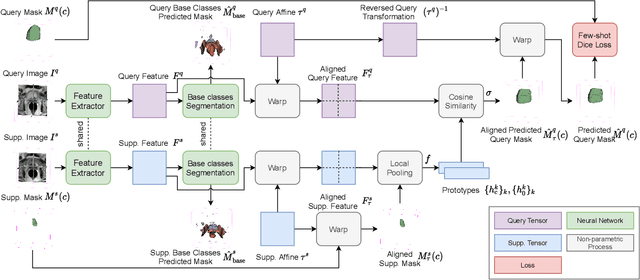
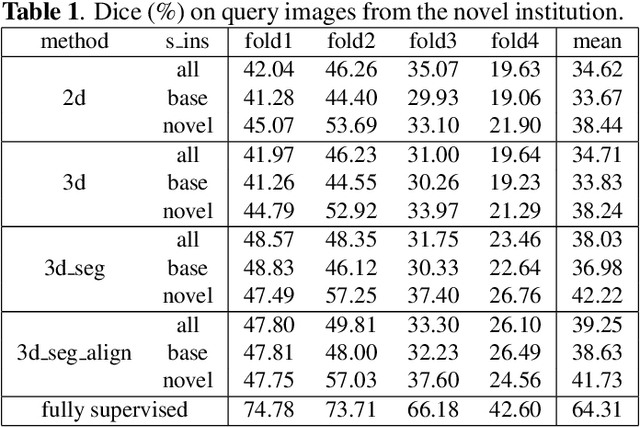
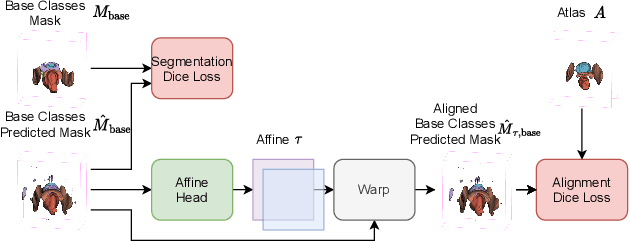
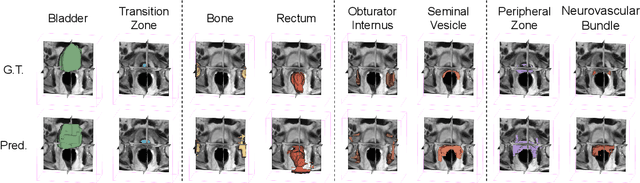
Abstract:The ability to adapt medical image segmentation networks for a novel class such as an unseen anatomical or pathological structure, when only a few labelled examples of this class are available from local healthcare providers, is sought-after. This potentially addresses two widely recognised limitations in deploying modern deep learning models to clinical practice, expertise-and-labour-intensive labelling and cross-institution generalisation. This work presents the first 3D few-shot interclass segmentation network for medical images, using a labelled multi-institution dataset from prostate cancer patients with eight regions of interest. We propose an image alignment module registering the predicted segmentation of both query and support data, in a standard prototypical learning algorithm, to a reference atlas space. The built-in registration mechanism can effectively utilise the prior knowledge of consistent anatomy between subjects, regardless whether they are from the same institution or not. Experimental results demonstrated that the proposed registration-assisted prototypical learning significantly improved segmentation accuracy (p-values<0.01) on query data from a holdout institution, with varying availability of support data from multiple institutions. We also report the additional benefits of the proposed 3D networks with 75% fewer parameters and an arguably simpler implementation, compared with existing 2D few-shot approaches that segment 2D slices of volumetric medical images.
Few-shot Semantic Segmentation with Self-supervision from Pseudo-classes
Oct 22, 2021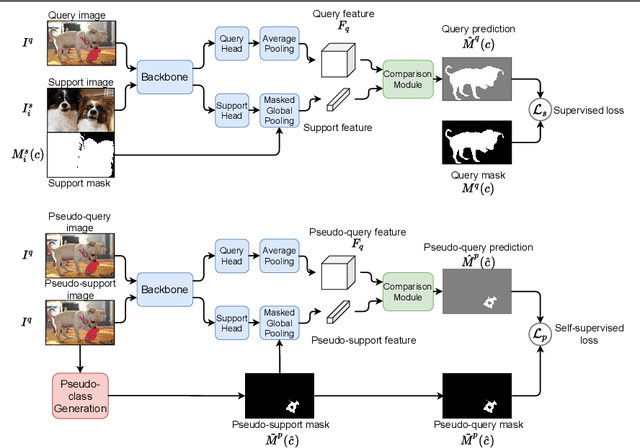
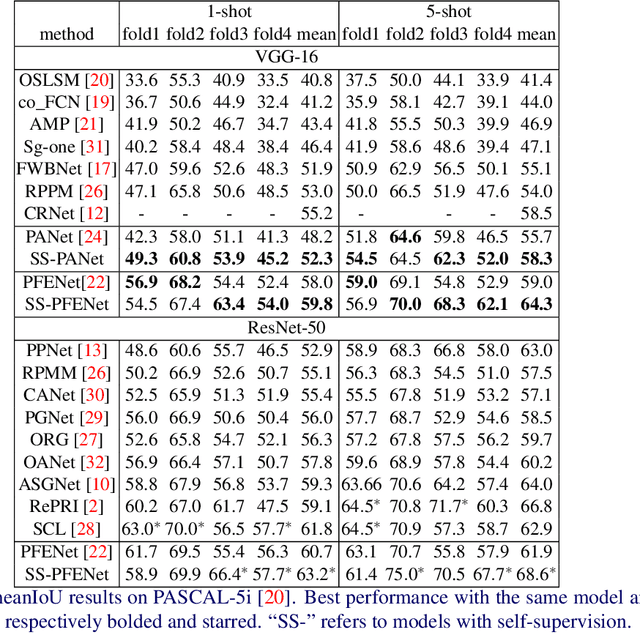

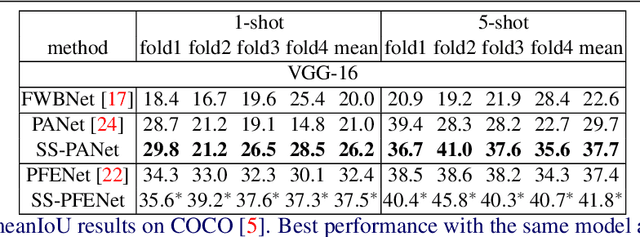
Abstract:Despite the success of deep learning methods for semantic segmentation, few-shot semantic segmentation remains a challenging task due to the limited training data and the generalisation requirement for unseen classes. While recent progress has been particularly encouraging, we discover that existing methods tend to have poor performance in terms of meanIoU when query images contain other semantic classes besides the target class. To address this issue, we propose a novel self-supervised task that generates random pseudo-classes in the background of the query images, providing extra training data that would otherwise be unavailable when predicting individual target classes. To that end, we adopted superpixel segmentation for generating the pseudo-classes. With this extra supervision, we improved the meanIoU performance of the state-of-the-art method by 2.5% and 5.1% on the one-shot tasks, as well as 6.7% and 4.4% on the five-shot tasks, on the PASCAL-5i and COCO benchmarks, respectively.
Adaptable image quality assessment using meta-reinforcement learning of task amenability
Jul 31, 2021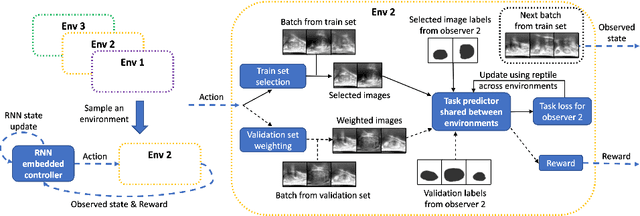
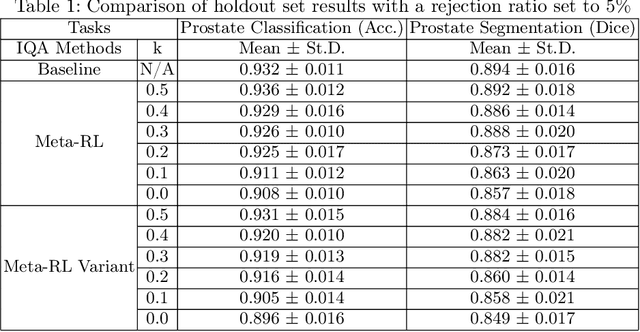
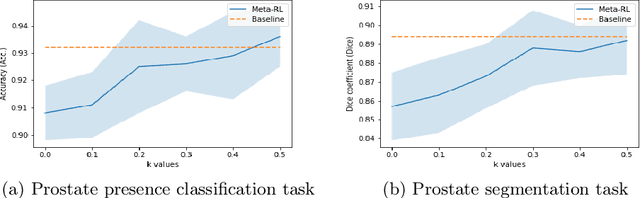
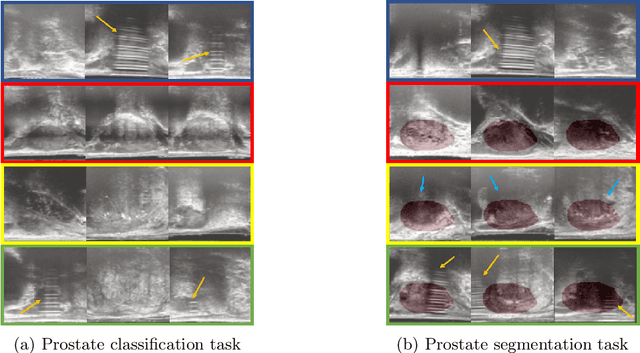
Abstract:The performance of many medical image analysis tasks are strongly associated with image data quality. When developing modern deep learning algorithms, rather than relying on subjective (human-based) image quality assessment (IQA), task amenability potentially provides an objective measure of task-specific image quality. To predict task amenability, an IQA agent is trained using reinforcement learning (RL) with a simultaneously optimised task predictor, such as a classification or segmentation neural network. In this work, we develop transfer learning or adaptation strategies to increase the adaptability of both the IQA agent and the task predictor so that they are less dependent on high-quality, expert-labelled training data. The proposed transfer learning strategy re-formulates the original RL problem for task amenability in a meta-reinforcement learning (meta-RL) framework. The resulting algorithm facilitates efficient adaptation of the agent to different definitions of image quality, each with its own Markov decision process environment including different images, labels and an adaptable task predictor. Our work demonstrates that the IQA agents pre-trained on non-expert task labels can be adapted to predict task amenability as defined by expert task labels, using only a small set of expert labels. Using 6644 clinical ultrasound images from 249 prostate cancer patients, our results for image classification and segmentation tasks show that the proposed IQA method can be adapted using data with as few as respective 19.7% and 29.6% expert-reviewed consensus labels and still achieve comparable IQA and task performance, which would otherwise require a training dataset with 100% expert labels.
Learning image quality assessment by reinforcing task amenable data selection
Feb 15, 2021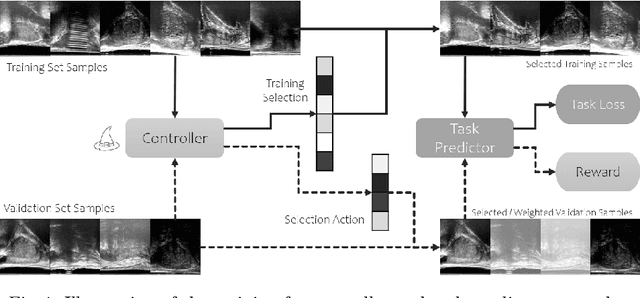
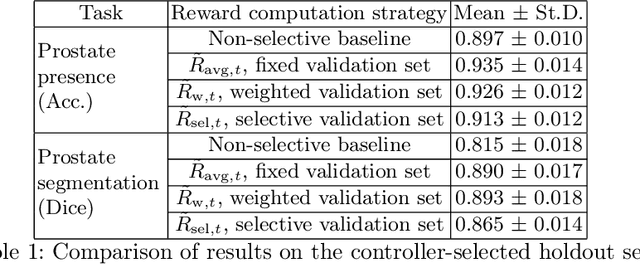

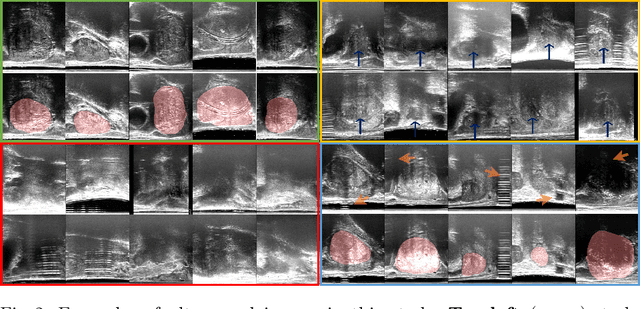
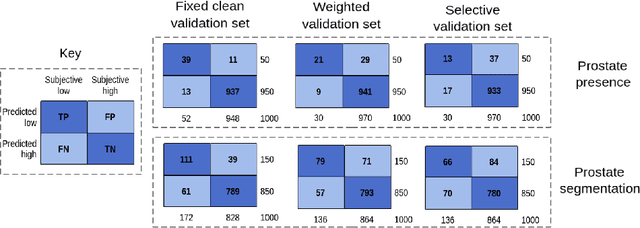
Abstract:In this paper, we consider a type of image quality assessment as a task-specific measurement, which can be used to select images that are more amenable to a given target task, such as image classification or segmentation. We propose to train simultaneously two neural networks for image selection and a target task using reinforcement learning. A controller network learns an image selection policy by maximising an accumulated reward based on the target task performance on the controller-selected validation set, whilst the target task predictor is optimised using the training set. The trained controller is therefore able to reject those images that lead to poor accuracy in the target task. In this work, we show that the controller-predicted image quality can be significantly different from the task-specific image quality labels that are manually defined by humans. Furthermore, we demonstrate that it is possible to learn effective image quality assessment without using a ``clean'' validation set, thereby avoiding the requirement for human labelling of images with respect to their amenability for the task. Using $6712$, labelled and segmented, clinical ultrasound images from $259$ patients, experimental results on holdout data show that the proposed image quality assessment achieved a mean classification accuracy of $0.94\pm0.01$ and a mean segmentation Dice of $0.89\pm0.02$, by discarding $5\%$ and $15\%$ of the acquired images, respectively. The significantly improved performance was observed for both tested tasks, compared with the respective $0.90\pm0.01$ and $0.82\pm0.02$ from networks without considering task amenability. This enables image quality feedback during real-time ultrasound acquisition among many other medical imaging applications.
Morphological Change Forecasting for Prostate Glands using Feature-based Registration and Kernel Density Extrapolation
Jan 16, 2021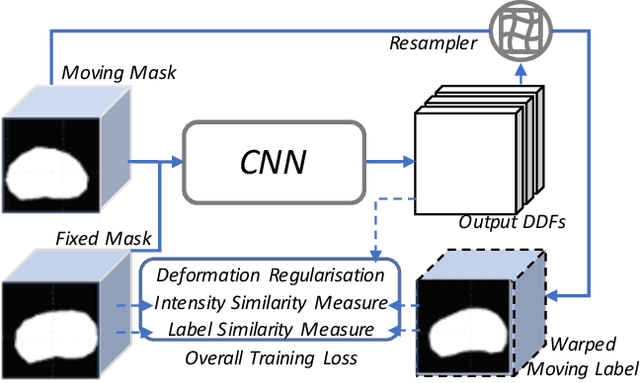
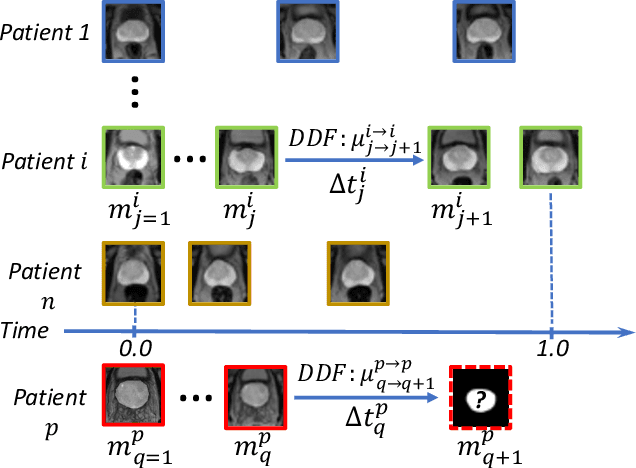
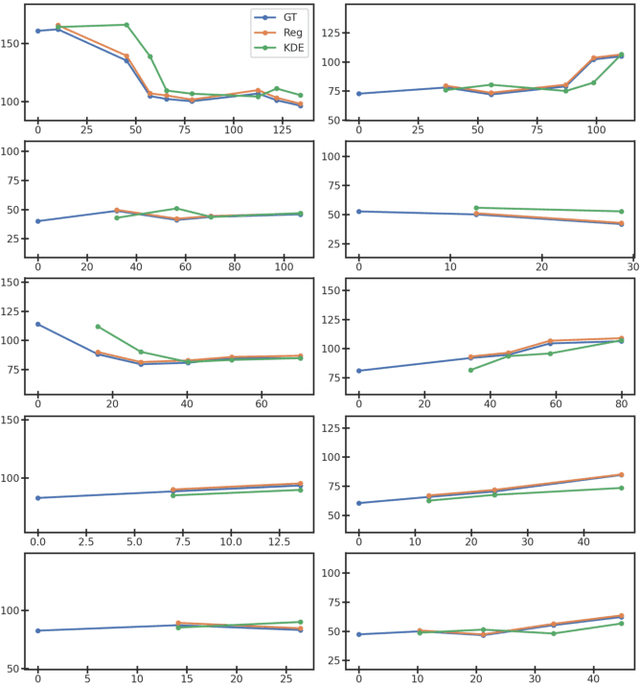
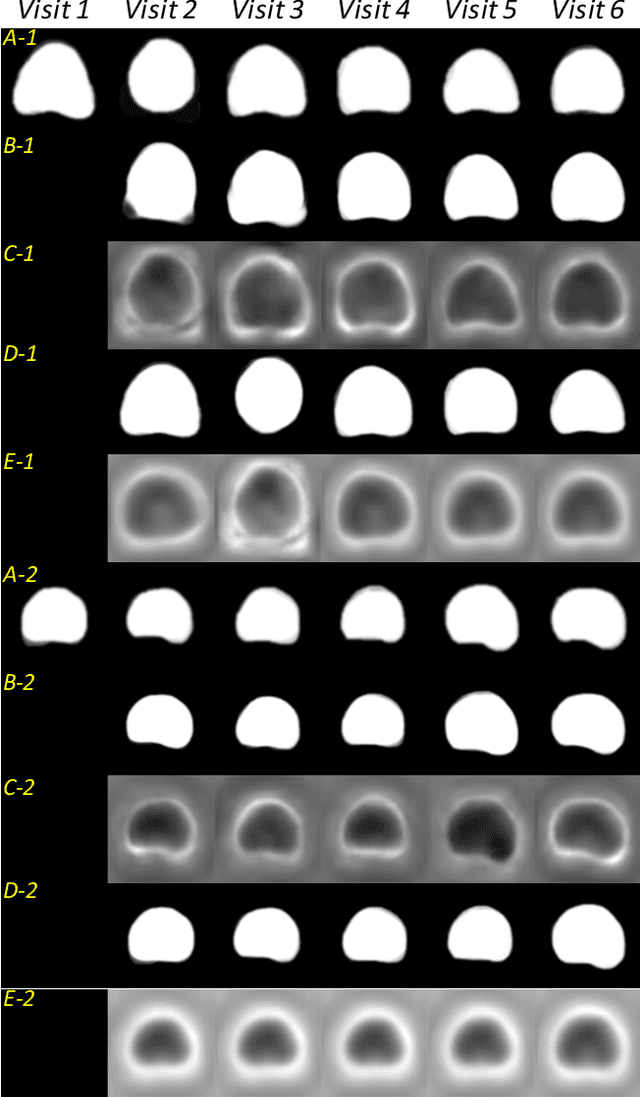
Abstract:Organ morphology is a key indicator for prostate disease diagnosis and prognosis. For instance, In longitudinal study of prostate cancer patients under active surveillance, the volume, boundary smoothness and their changes are closely monitored on time-series MR image data. In this paper, we describe a new framework for forecasting prostate morphological changes, as the ability to detect such changes earlier than what is currently possible may enable timely treatment or avoiding unnecessary confirmatory biopsies. In this work, an efficient feature-based MR image registration is first developed to align delineated prostate gland capsules to quantify the morphological changes using the inferred dense displacement fields (DDFs). We then propose to use kernel density estimation (KDE) of the probability density of the DDF-represented \textit{future morphology changes}, between current and future time points, before the future data become available. The KDE utilises a novel distance function that takes into account morphology, stage-of-progression and duration-of-change, which are considered factors in such subject-specific forecasting. We validate the proposed approach on image masks unseen to registration network training, without using any data acquired at the future target time points. The experiment results are presented on a longitudinal data set with 331 images from 73 patients, yielding an average Dice score of 0.865 on a holdout set, between the ground-truth and the image masks warped by the KDE-predicted-DDFs.
DeepReg: a deep learning toolkit for medical image registration
Nov 04, 2020Abstract:DeepReg (https://github.com/DeepRegNet/DeepReg) is a community-supported open-source toolkit for research and education in medical image registration using deep learning.
Longitudinal Image Registration with Temporal-order and Subject-specificity Discrimination
Aug 29, 2020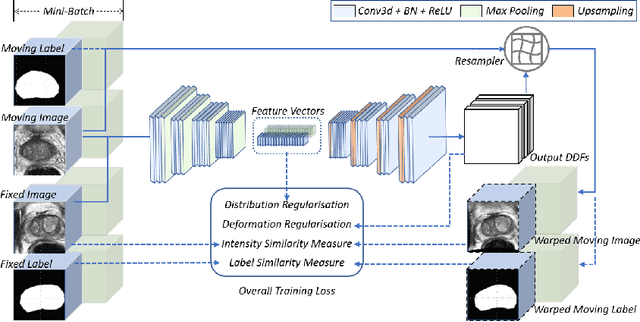

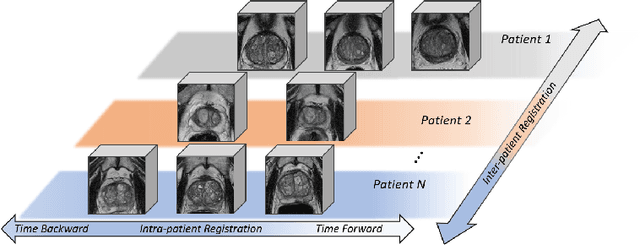
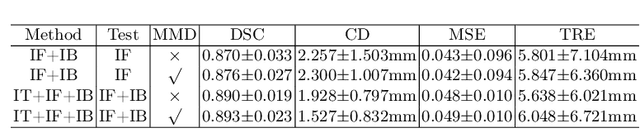
Abstract:Morphological analysis of longitudinal MR images plays a key role in monitoring disease progression for prostate cancer patients, who are placed under an active surveillance program. In this paper, we describe a learning-based image registration algorithm to quantify changes on regions of interest between a pair of images from the same patient, acquired at two different time points. Combining intensity-based similarity and gland segmentation as weak supervision, the population-data-trained registration networks significantly lowered the target registration errors (TREs) on holdout patient data, compared with those before registration and those from an iterative registration algorithm. Furthermore, this work provides a quantitative analysis on several longitudinal-data-sampling strategies and, in turn, we propose a novel regularisation method based on maximum mean discrepancy, between differently-sampled training image pairs. Based on 216 3D MR images from 86 patients, we report a mean TRE of 5.6 mm and show statistically significant differences between the different training data sampling strategies.
More unlabelled data or label more data? A study on semi-supervised laparoscopic image segmentation
Aug 20, 2019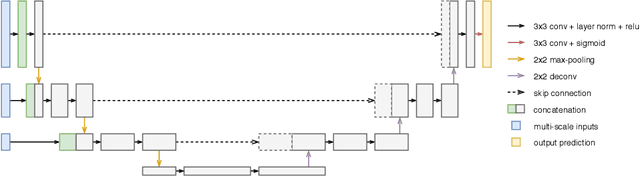
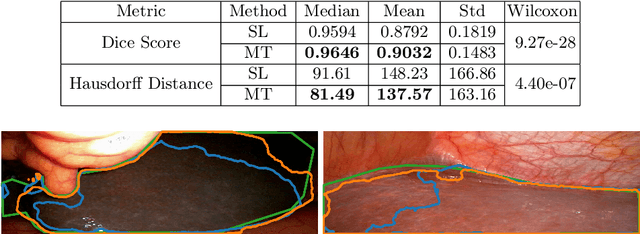
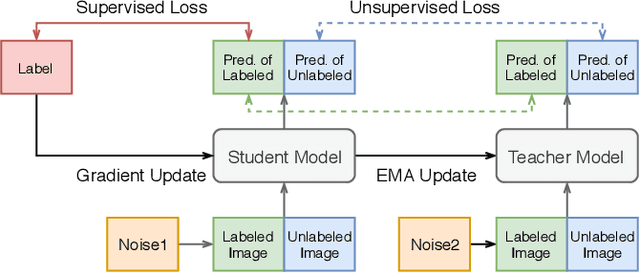
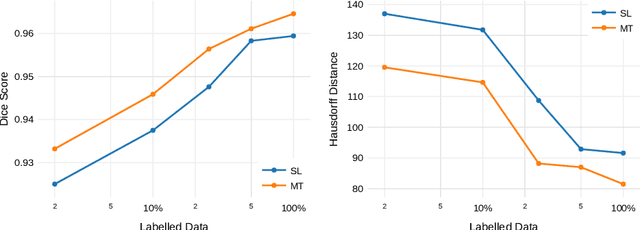
Abstract:Improving a semi-supervised image segmentation task has the option of adding more unlabelled images, labelling the unlabelled images or combining both, as neither image acquisition nor expert labelling can be considered trivial in most clinical applications. With a laparoscopic liver image segmentation application, we investigate the performance impact by altering the quantities of labelled and unlabelled training data, using a semi-supervised segmentation algorithm based on the mean teacher learning paradigm. We first report a significantly higher segmentation accuracy, compared with supervised learning. Interestingly, this comparison reveals that the training strategy adopted in the semi-supervised algorithm is also responsible for this observed improvement, in addition to the added unlabelled data. We then compare different combinations of labelled and unlabelled data set sizes for training semi-supervised segmentation networks, to provide a quantitative example of the practically useful trade-off between the two data planning strategies in this surgical guidance application.
Ranked Reward: Enabling Self-Play Reinforcement Learning for Combinatorial Optimization
Jul 06, 2018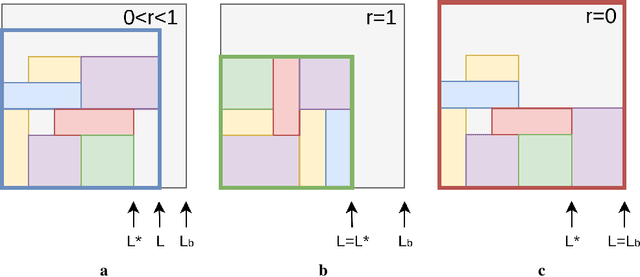
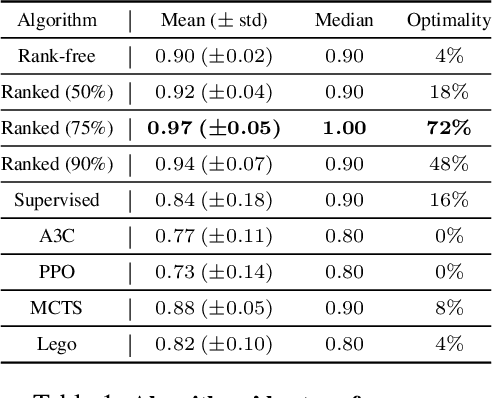
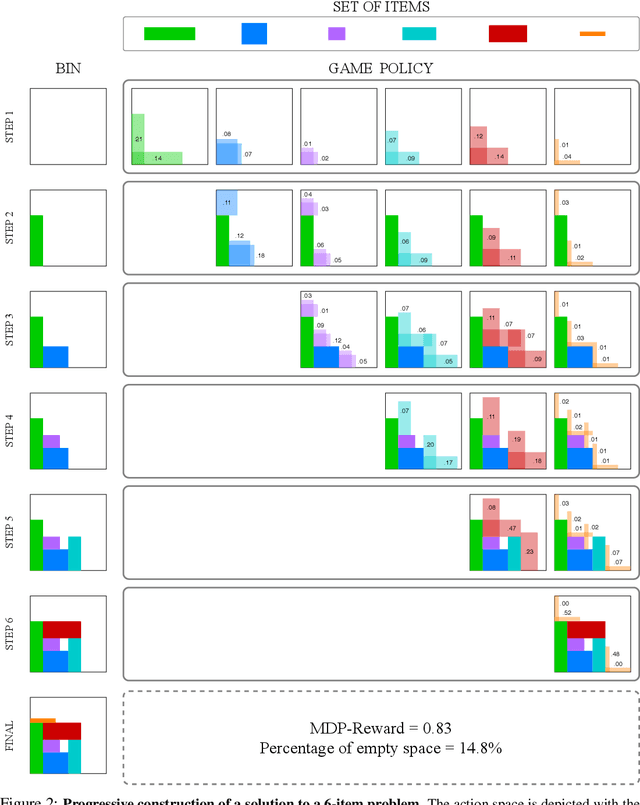
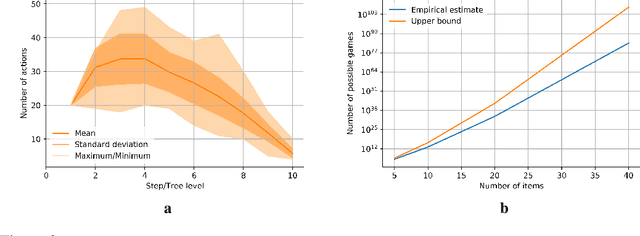
Abstract:Adversarial self-play in two-player games has delivered impressive results when used with reinforcement learning algorithms that combine deep neural networks and tree search. Algorithms like AlphaZero and Expert Iteration learn tabula-rasa, producing highly informative training data on the fly. However, the self-play training strategy is not directly applicable to single-player games. Recently, several practically important combinatorial optimization problems, such as the traveling salesman problem and the bin packing problem, have been reformulated as reinforcement learning problems, increasing the importance of enabling the benefits of self-play beyond two-player games. We present the Ranked Reward (R2) algorithm which accomplishes this by ranking the rewards obtained by a single agent over multiple games to create a relative performance metric. Results from applying the R2 algorithm to instances of a two-dimensional bin packing problem show that it outperforms generic Monte Carlo tree search, heuristic algorithms and reinforcement learning algorithms not using ranked rewards.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge