Yanwu Xu
South China University of Technology
Learning to screen Glaucoma like the ophthalmologists
Sep 23, 2022
Abstract:GAMMA Challenge is organized to encourage the AI models to screen the glaucoma from a combination of 2D fundus image and 3D optical coherence tomography volume, like the ophthalmologists.
Multi-Scale Multi-Target Domain Adaptation for Angle Closure Classification
Aug 25, 2022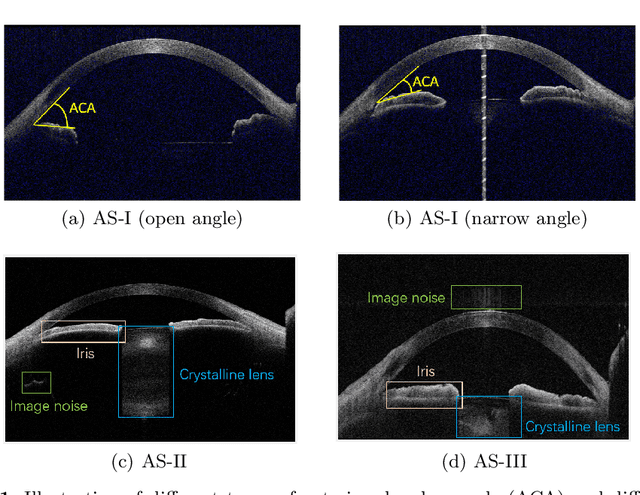
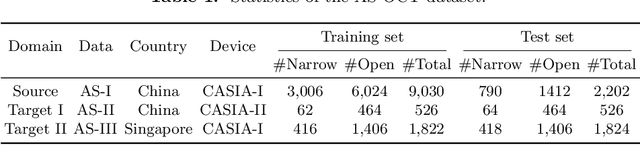
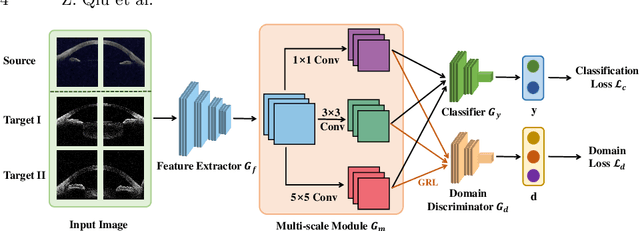
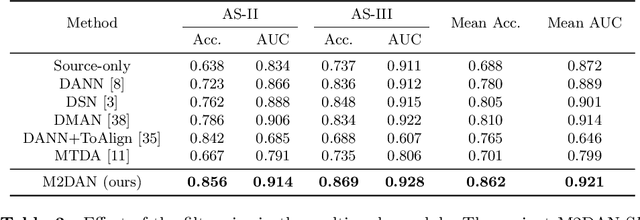
Abstract:Deep learning (DL) has made significant progress in angle closure classification with anterior segment optical coherence tomography (AS-OCT) images. These AS-OCT images are often acquired by different imaging devices/conditions, which results in a vast change of underlying data distributions (called "data domains"). Moreover, due to practical labeling difficulties, some domains (e.g., devices) may not have any data labels. As a result, deep models trained on one specific domain (e.g., a specific device) are difficult to adapt to and thus may perform poorly on other domains (e.g., other devices). To address this issue, we present a multi-target domain adaptation paradigm to transfer a model trained on one labeled source domain to multiple unlabeled target domains. Specifically, we propose a novel Multi-scale Multi-target Domain Adversarial Network (M2DAN) for angle closure classification. M2DAN conducts multi-domain adversarial learning for extracting domain-invariant features and develops a multi-scale module for capturing local and global information of AS-OCT images. Based on these domain-invariant features at different scales, the deep model trained on the source domain is able to classify angle closure on multiple target domains even without any annotations in these domains. Extensive experiments on a real-world AS-OCT dataset demonstrate the effectiveness of the proposed method.
Calibrate the inter-observer segmentation uncertainty via diagnosis-first principle
Aug 05, 2022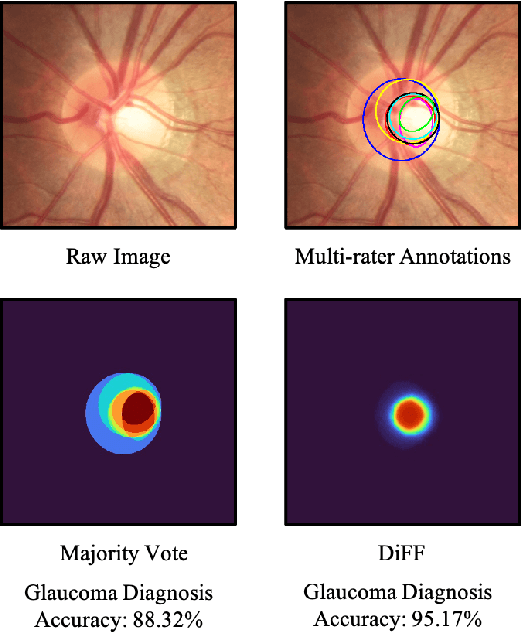
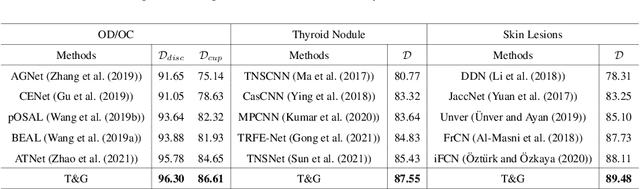
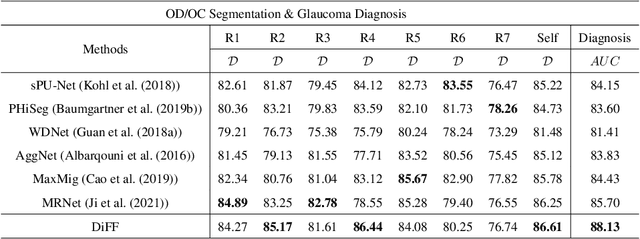
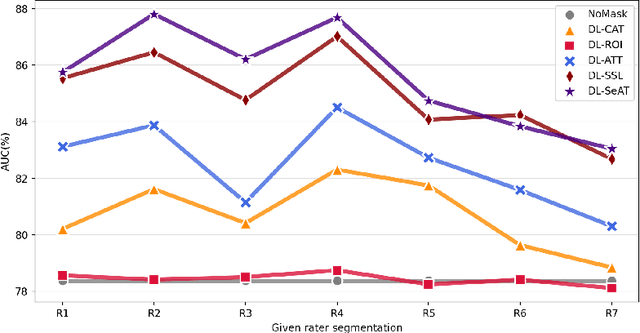
Abstract:On the medical images, many of the tissues/lesions may be ambiguous. That is why the medical segmentation is typically annotated by a group of clinical experts to mitigate the personal bias. However, this clinical routine also brings new challenges to the application of machine learning algorithms. Without a definite ground-truth, it will be difficult to train and evaluate the deep learning models. When the annotations are collected from different graders, a common choice is majority vote. However such a strategy ignores the difference between the grader expertness. In this paper, we consider the task of predicting the segmentation with the calibrated inter-observer uncertainty. We note that in clinical practice, the medical image segmentation is usually used to assist the disease diagnosis. Inspired by this observation, we propose diagnosis-first principle, which is to take disease diagnosis as the criterion to calibrate the inter-observer segmentation uncertainty. Following this idea, a framework named Diagnosis First segmentation Framework (DiFF) is proposed to estimate diagnosis-first segmentation from the raw images.Specifically, DiFF will first learn to fuse the multi-rater segmentation labels to a single ground-truth which could maximize the disease diagnosis performance. We dubbed the fused ground-truth as Diagnosis First Ground-truth (DF-GT).Then, we further propose Take and Give Modelto segment DF-GT from the raw image. We verify the effectiveness of DiFF on three different medical segmentation tasks: OD/OC segmentation on fundus images, thyroid nodule segmentation on ultrasound images, and skin lesion segmentation on dermoscopic images. Experimental results show that the proposed DiFF is able to significantly facilitate the corresponding disease diagnosis, which outperforms previous state-of-the-art multi-rater learning methods.
Dataset and Evaluation algorithm design for GOALS Challenge
Jul 29, 2022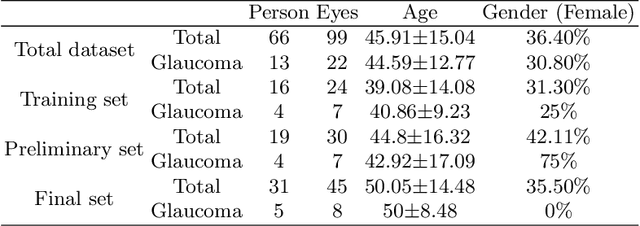
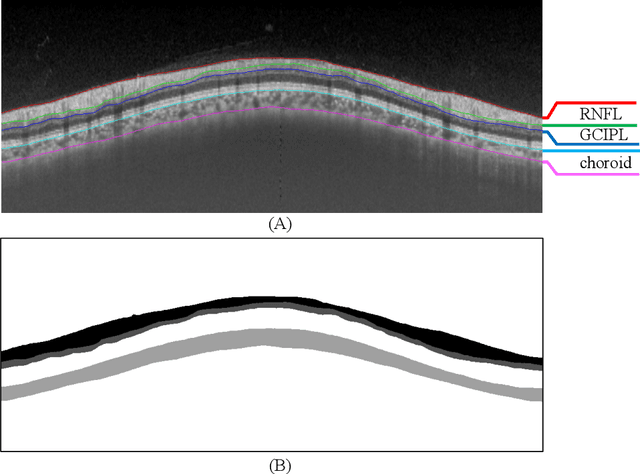
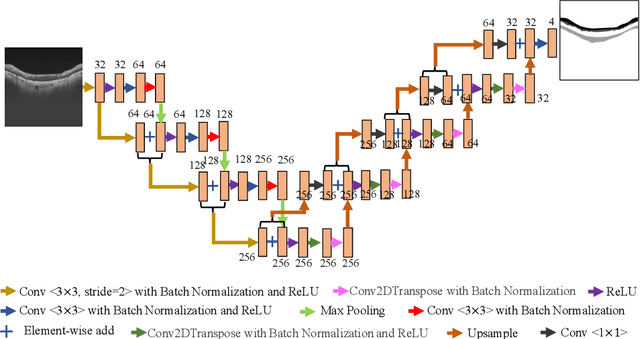
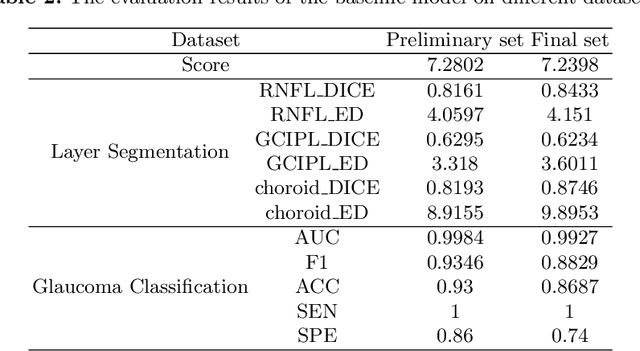
Abstract:Glaucoma causes irreversible vision loss due to damage to the optic nerve, and there is no cure for glaucoma.OCT imaging modality is an essential technique for assessing glaucomatous damage since it aids in quantifying fundus structures. To promote the research of AI technology in the field of OCT-assisted diagnosis of glaucoma, we held a Glaucoma OCT Analysis and Layer Segmentation (GOALS) Challenge in conjunction with the International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI) 2022 to provide data and corresponding annotations for researchers studying layer segmentation from OCT images and the classification of glaucoma. This paper describes the released 300 circumpapillary OCT images, the baselines of the two sub-tasks, and the evaluation methodology. The GOALS Challenge is accessible at https://aistudio.baidu.com/aistudio/competition/detail/230.
Towards Lightweight Super-Resolution with Dual Regression Learning
Jul 21, 2022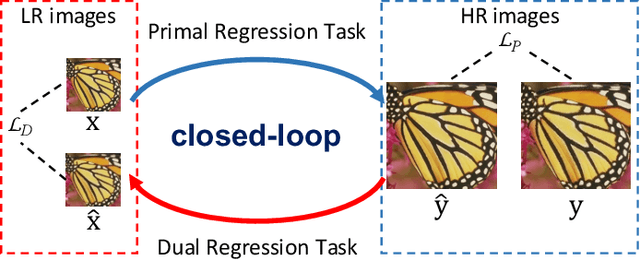
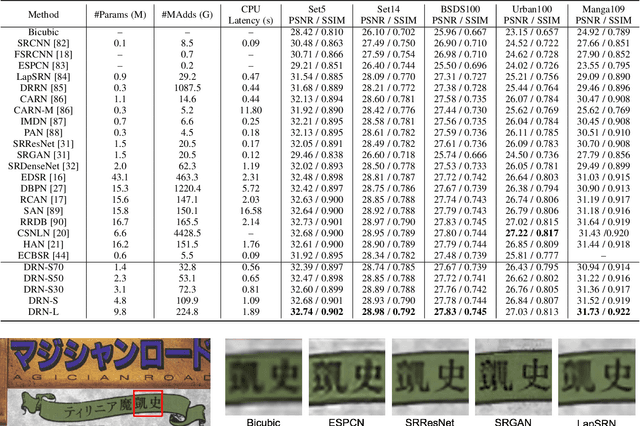
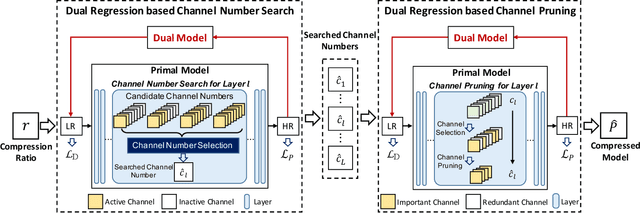
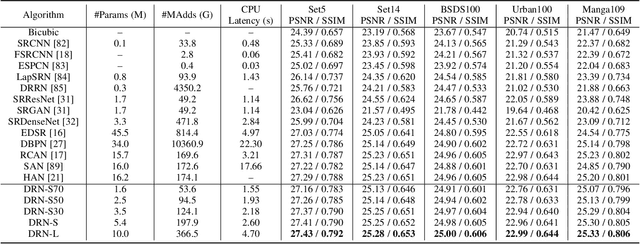
Abstract:Deep neural networks have exhibited remarkable performance in image super-resolution (SR) tasks by learning a mapping from low-resolution (LR) images to high-resolution (HR) images. However, the SR problem is typically an ill-posed problem and existing methods would come with several limitations. First, the possible mapping space of SR can be extremely large since there may exist many different HR images that can be downsampled to the same LR image. As a result, it is hard to directly learn a promising SR mapping from such a large space. Second, it is often inevitable to develop very large models with extremely high computational cost to yield promising SR performance. In practice, one can use model compression techniques to obtain compact models by reducing model redundancy. Nevertheless, it is hard for existing model compression methods to accurately identify the redundant components due to the extremely large SR mapping space. To alleviate the first challenge, we propose a dual regression learning scheme to reduce the space of possible SR mappings. Specifically, in addition to the mapping from LR to HR images, we learn an additional dual regression mapping to estimate the downsampling kernel and reconstruct LR images. In this way, the dual mapping acts as a constraint to reduce the space of possible mappings. To address the second challenge, we propose a lightweight dual regression compression method to reduce model redundancy in both layer-level and channel-level based on channel pruning. Specifically, we first develop a channel number search method that minimizes the dual regression loss to determine the redundancy of each layer. Given the searched channel numbers, we further exploit the dual regression manner to evaluate the importance of channels and prune the redundant ones. Extensive experiments show the effectiveness of our method in obtaining accurate and efficient SR models.
Adversarial Consistency for Single Domain Generalization in Medical Image Segmentation
Jun 29, 2022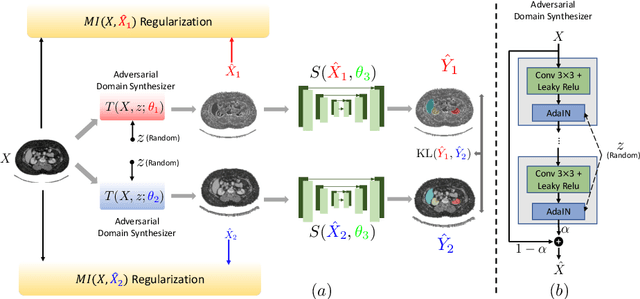
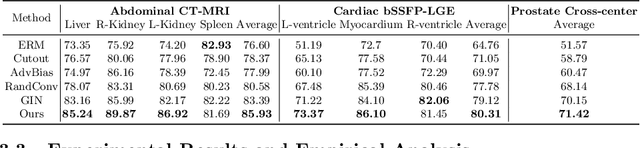
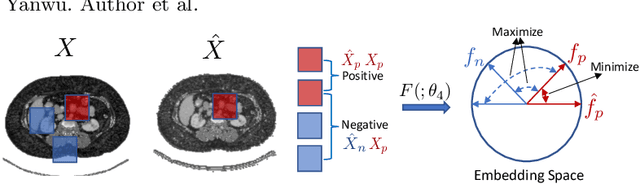
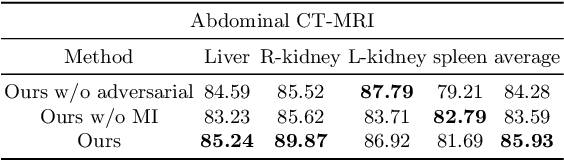
Abstract:An organ segmentation method that can generalize to unseen contrasts and scanner settings can significantly reduce the need for retraining of deep learning models. Domain Generalization (DG) aims to achieve this goal. However, most DG methods for segmentation require training data from multiple domains during training. We propose a novel adversarial domain generalization method for organ segmentation trained on data from a \emph{single} domain. We synthesize the new domains via learning an adversarial domain synthesizer (ADS) and presume that the synthetic domains cover a large enough area of plausible distributions so that unseen domains can be interpolated from synthetic domains. We propose a mutual information regularizer to enforce the semantic consistency between images from the synthetic domains, which can be estimated by patch-level contrastive learning. We evaluate our method for various organ segmentation for unseen modalities, scanning protocols, and scanner sites.
SeATrans: Learning Segmentation-Assisted diagnosis model via Transformer
Jun 22, 2022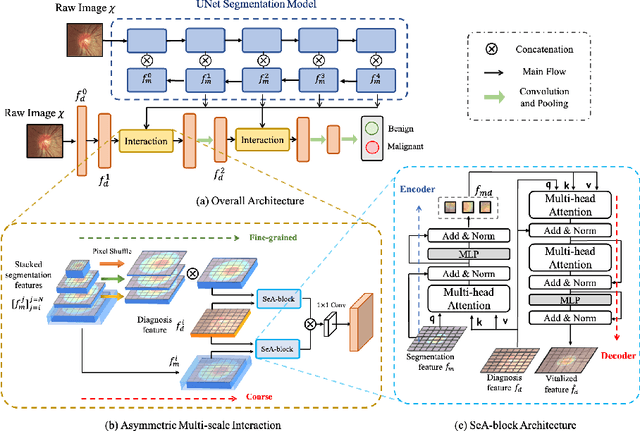
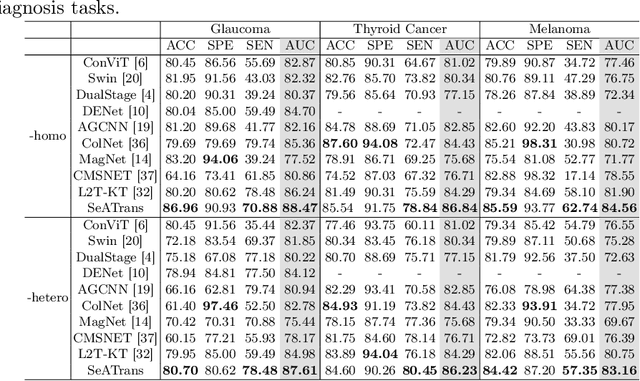
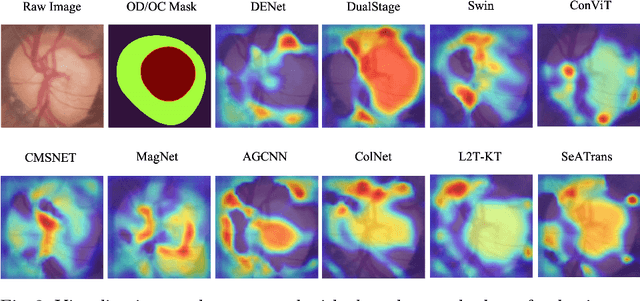
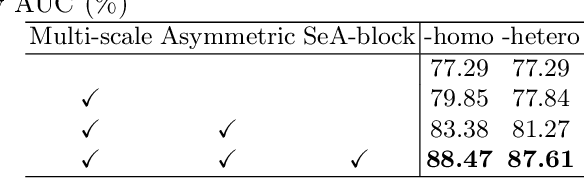
Abstract:Clinically, the accurate annotation of lesions/tissues can significantly facilitate the disease diagnosis. For example, the segmentation of optic disc/cup (OD/OC) on fundus image would facilitate the glaucoma diagnosis, the segmentation of skin lesions on dermoscopic images is helpful to the melanoma diagnosis, etc. With the advancement of deep learning techniques, a wide range of methods proved the lesions/tissues segmentation can also facilitate the automated disease diagnosis models. However, existing methods are limited in the sense that they can only capture static regional correlations in the images. Inspired by the global and dynamic nature of Vision Transformer, in this paper, we propose Segmentation-Assisted diagnosis Transformer (SeATrans) to transfer the segmentation knowledge to the disease diagnosis network. Specifically, we first propose an asymmetric multi-scale interaction strategy to correlate each single low-level diagnosis feature with multi-scale segmentation features. Then, an effective strategy called SeA-block is adopted to vitalize diagnosis feature via correlated segmentation features. To model the segmentation-diagnosis interaction, SeA-block first embeds the diagnosis feature based on the segmentation information via the encoder, and then transfers the embedding back to the diagnosis feature space by a decoder. Experimental results demonstrate that SeATrans surpasses a wide range of state-of-the-art (SOTA) segmentation-assisted diagnosis methods on several disease diagnosis tasks.
Learning self-calibrated optic disc and cup segmentation from multi-rater annotations
Jun 14, 2022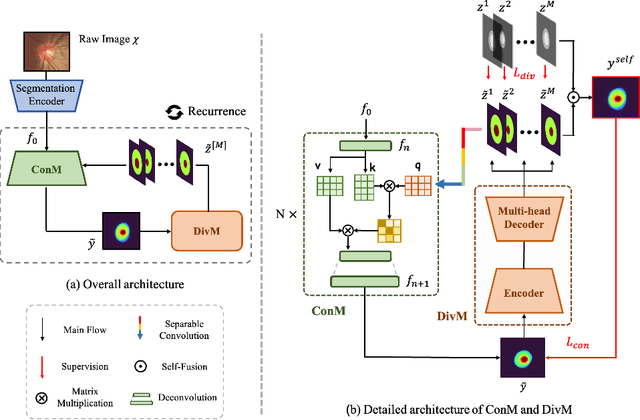
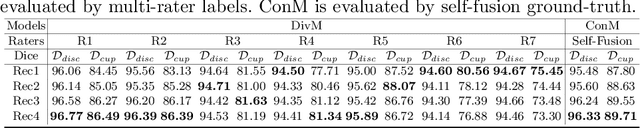
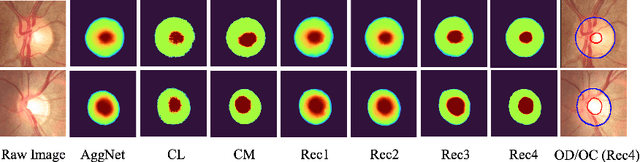
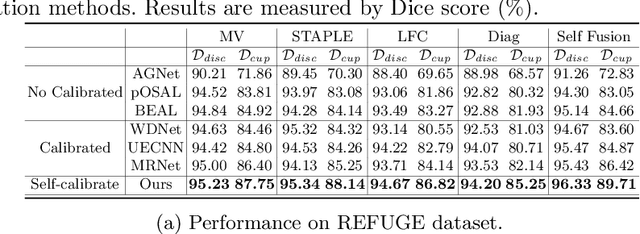
Abstract:The segmentation of optic disc(OD) and optic cup(OC) from fundus images is an important fundamental task for glaucoma diagnosis. In the clinical practice, it is often necessary to collect opinions from multiple experts to obtain the final OD/OC annotation. This clinical routine helps to mitigate the individual bias. But when data is multiply annotated, standard deep learning models will be inapplicable. In this paper, we propose a novel neural network framework to learn OD/OC segmentation from multi-rater annotations. The segmentation results are self-calibrated through the iterative optimization of multi-rater expertness estimation and calibrated OD/OC segmentation. In this way, the proposed method can realize a mutual improvement of both tasks and finally obtain a refined segmentation result. Specifically, we propose Diverging Model(DivM) and Converging Model(ConM) to process the two tasks respectively. ConM segments the raw image based on the multi-rater expertness map provided by DivM. DivM generates multi-rater expertness map from the segmentation mask provided by ConM. The experiment results show that by recurrently running ConM and DivM, the results can be self-calibrated so as to outperform a range of state-of-the-art(SOTA) multi-rater segmentation methods.
One Hyper-Initializer for All Network Architectures in Medical Image Analysis
Jun 08, 2022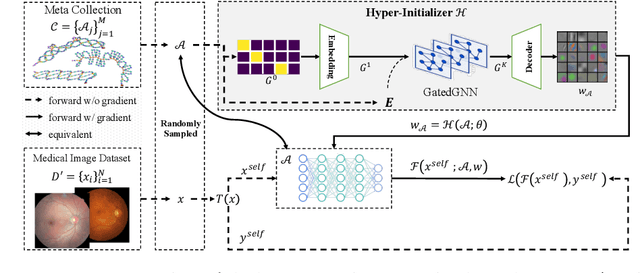
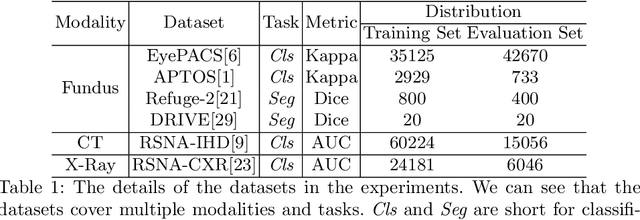
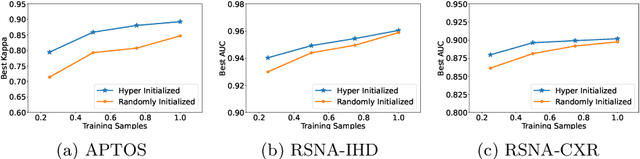
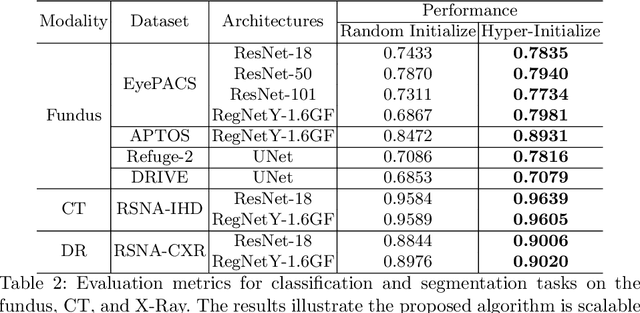
Abstract:Pre-training is essential to deep learning model performance, especially in medical image analysis tasks where limited training data are available. However, existing pre-training methods are inflexible as the pre-trained weights of one model cannot be reused by other network architectures. In this paper, we propose an architecture-irrelevant hyper-initializer, which can initialize any given network architecture well after being pre-trained for only once. The proposed initializer is a hypernetwork which takes a downstream architecture as input graphs and outputs the initialization parameters of the respective architecture. We show the effectiveness and efficiency of the hyper-initializer through extensive experimental results on multiple medical imaging modalities, especially in data-limited fields. Moreover, we prove that the proposed algorithm can be reused as a favorable plug-and-play initializer for any downstream architecture and task (both classification and segmentation) of the same modality.
Contrastive Centroid Supervision Alleviates Domain Shift in Medical Image Classification
May 31, 2022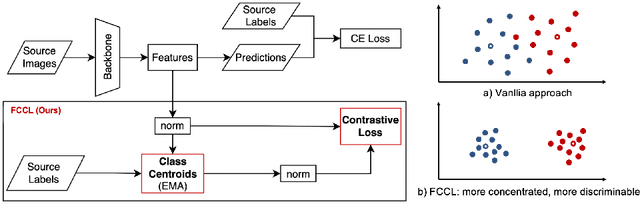
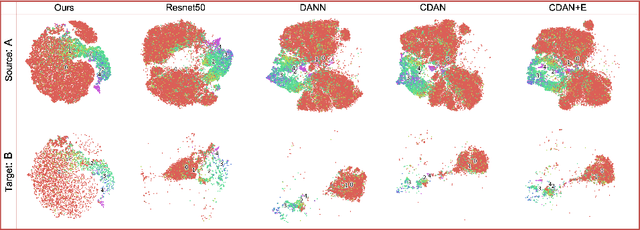
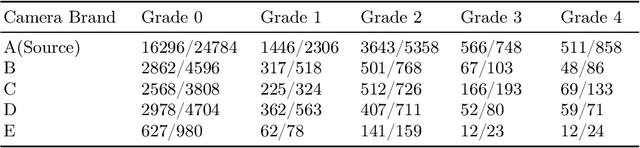
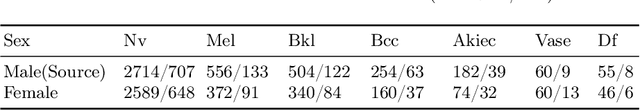

Abstract:Deep learning based medical imaging classification models usually suffer from the domain shift problem, where the classification performance drops when training data and real-world data differ in imaging equipment manufacturer, image acquisition protocol, patient populations, etc. We propose Feature Centroid Contrast Learning (FCCL), which can improve target domain classification performance by extra supervision during training with contrastive loss between instance and class centroid. Compared with current unsupervised domain adaptation and domain generalization methods, FCCL performs better while only requires labeled image data from a single source domain and no target domain. We verify through extensive experiments that FCCL can achieve superior performance on at least three imaging modalities, i.e. fundus photographs, dermatoscopic images, and H & E tissue images.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge