Xiaozhong Liu
Beyond Local vs. External: A Game-Theoretic Framework for Trustworthy Knowledge Acquisition
Apr 25, 2026Abstract:Cloud-hosted Large Language Models (LLMs) offer unmatched reasoning capabilities and dynamic knowledge, yet submitting raw queries to these external services risks exposing sensitive user intent. Conversely, relying exclusively on trusted local models preserves privacy but often compromises answer quality due to limited parameter scale and knowledge. To resolve this dilemma, we propose Game-theoretic Trustworthy Knowledge Acquisition (GTKA), a framework that formulates the trade-off between knowledge utility and privacy as a strategic game. GTKA consists of three components: (i) a privacy-aware sub-query generator that decomposes sensitive intent into generalized, low-risk fragments; (ii) an adversarial reconstruction attacker that attempts to infer the original query from these fragments, providing adaptive leakage signals; and (iii) a trusted local integrator that synthesizes external responses within a secure boundary. By training the generator and attacker in an alternating adversarial manner, GTKA optimizes the sub-query generation policy to maximize knowledge acquisition accuracy while minimizing the reconstructability of the original sensitive intent. To validate our approach, we construct two sensitive-domain benchmarks in the biomedical and legal fields. Extensive experiments demonstrate that GTKA significantly reduces intent leakage compared to state-of-the-art baselines while maintaining high-fidelity answer quality.
Can We Trust a Black-box LLM? LLM Untrustworthy Boundary Detection via Bias-Diffusion and Multi-Agent Reinforcement Learning
Apr 07, 2026Abstract:Large Language Models (LLMs) have shown a high capability in answering questions on a diverse range of topics. However, these models sometimes produce biased, ideologized or incorrect responses, limiting their applications if there is no clear understanding of which topics their answers can be trusted. In this research, we introduce a novel algorithm, named as GMRL-BD, designed to identify the untrustworthy boundaries (in terms of topics) of a given LLM, with black-box access to the LLM and under specific query constraints. Based on a general Knowledge Graph (KG) derived from Wikipedia, our algorithm incorporates with multiple reinforcement learning agents to efficiently identify topics (some nodes in KG) where the LLM is likely to generate biased answers. Our experiments demonstrated the efficiency of our algorithm, which can detect the untrustworthy boundary with just limited queries to the LLM. Additionally, we have released a new dataset containing popular LLMs including Llama2, Vicuna, Falcon, Qwen2, Gemma2 and Yi-1.5, along with labels indicating the topics on which each LLM is likely to be biased.
PubMed Reasoner: Dynamic Reasoning-based Retrieval for Evidence-Grounded Biomedical Question Answering
Mar 28, 2026Abstract:Trustworthy biomedical question answering (QA) systems must not only provide accurate answers but also justify them with current, verifiable evidence. Retrieval-augmented approaches partially address this gap but lack mechanisms to iteratively refine poor queries, whereas self-reflection methods kick in only after full retrieval is completed. In this context, we introduce PubMed Reasoner, a biomedical QA agent composed of three stages: self-critic query refinement evaluates MeSH terms for coverage, alignment, and redundancy to enhance PubMed queries based on partial (metadata) retrieval; reflective retrieval processes articles in batches until sufficient evidence is gathered; and evidence-grounded response generation produces answers with explicit citations. PubMed Reasoner with a GPT-4o backbone achieves 78.32% accuracy on PubMedQA, slightly surpassing human experts, and showing consistent gains on MMLU Clinical Knowledge. Moreover, LLM-as-judge evaluations prefer our responses across: reasoning soundness, evidence grounding, clinical relevance, and trustworthiness. By orchestrating retrieval-first reasoning over authoritative sources, our approach provides practical assistance to clinicians and biomedical researchers while controlling compute and token costs.
P2S: Probabilistic Process Supervision for General-Domain Reasoning Question Answering
Jan 28, 2026Abstract:While reinforcement learning with verifiable rewards (RLVR) has advanced LLM reasoning in structured domains like mathematics and programming, its application to general-domain reasoning tasks remains challenging due to the absence of verifiable reward signals. To this end, methods like Reinforcement Learning with Reference Probability Reward (RLPR) have emerged, leveraging the probability of generating the final answer as a reward signal. However, these outcome-focused approaches neglect crucial step-by-step supervision of the reasoning process itself. To address this gap, we introduce Probabilistic Process Supervision (P2S), a novel self-supervision framework that provides fine-grained process rewards without requiring a separate reward model or human-annotated reasoning steps. During reinforcement learning, P2S synthesizes and filters a high-quality reference reasoning chain (gold-CoT). The core of our method is to calculate a Path Faithfulness Reward (PFR) for each reasoning step, which is derived from the conditional probability of generating the gold-CoT's suffix, given the model's current reasoning prefix. Crucially, this PFR can be flexibly integrated with any outcome-based reward, directly tackling the reward sparsity problem by providing dense guidance. Extensive experiments on reading comprehension and medical Question Answering benchmarks show that P2S significantly outperforms strong baselines.
VitalDiagnosis: AI-Driven Ecosystem for 24/7 Vital Monitoring and Chronic Disease Management
Jan 22, 2026Abstract:Chronic diseases have become the leading cause of death worldwide, a challenge intensified by strained medical resources and an aging population. Individually, patients often struggle to interpret early signs of deterioration or maintain adherence to care plans. In this paper, we introduce VitalDiagnosis, an LLM-driven ecosystem designed to shift chronic disease management from passive monitoring to proactive, interactive engagement. By integrating continuous data from wearable devices with the reasoning capabilities of LLMs, the system addresses both acute health anomalies and routine adherence. It analyzes triggers through context-aware inquiries, produces provisional insights within a collaborative patient-clinician workflow, and offers personalized guidance. This approach aims to promote a more proactive and cooperative care paradigm, with the potential to enhance patient self-management and reduce avoidable clinical workload.
Human-in-the-Loop Interactive Report Generation for Chronic Disease Adherence
Jan 10, 2026Abstract:Chronic disease management requires regular adherence feedback to prevent avoidable hospitalizations, yet clinicians lack time to produce personalized patient communications. Manual authoring preserves clinical accuracy but does not scale; AI generation scales but can undermine trust in patient-facing contexts. We present a clinician-in-the-loop interface that constrains AI to data organization and preserves physician oversight through recognition-based review. A single-page editor pairs AI-generated section drafts with time-aligned visualizations, enabling inline editing with visual evidence for each claim. This division of labor (AI organizes, clinician decides) targets both efficiency and accountability. In a pilot with three physicians reviewing 24 cases, AI successfully generated clinically personalized drafts matching physicians' manual authoring practice (overall mean 4.86/10 vs. 5.0/10 baseline), requiring minimal physician editing (mean 8.3\% content modification) with zero safety-critical issues, demonstrating effective automation of content generation. However, review time remained comparable to manual practice, revealing an accountability paradox: in high-stakes clinical contexts, professional responsibility requires complete verification regardless of AI accuracy. We contribute three interaction patterns for clinical AI collaboration: bounded generation with recognition-based review via chart-text pairing, automated urgency flagging that analyzes vital trends and adherence patterns with fail-safe escalation for missed critical monitoring tasks, and progressive disclosure controls that reduce cognitive load while maintaining oversight. These patterns indicate that clinical AI efficiency requires not only accurate models, but also mechanisms for selective verification that preserve accountability.
Teaching According to Students' Aptitude: Personalized Mathematics Tutoring via Persona-, Memory-, and Forgetting-Aware LLMs
Nov 19, 2025Abstract:Large Language Models (LLMs) are increasingly integrated into intelligent tutoring systems to provide human-like and adaptive instruction. However, most existing approaches fail to capture how students' knowledge evolves dynamically across their proficiencies, conceptual gaps, and forgetting patterns. This challenge is particularly acute in mathematics tutoring, where effective instruction requires fine-grained scaffolding precisely calibrated to each student's mastery level and cognitive retention. To address this issue, we propose TASA (Teaching According to Students' Aptitude), a student-aware tutoring framework that integrates persona, memory, and forgetting dynamics for personalized mathematics learning. Specifically, TASA maintains a structured student persona capturing proficiency profiles and an event memory recording prior learning interactions. By incorporating a continuous forgetting curve with knowledge tracing, TASA dynamically updates each student's mastery state and generates contextually appropriate, difficulty-calibrated questions and explanations. Empirical results demonstrate that TASA achieves superior learning outcomes and more adaptive tutoring behavior compared to representative baselines, underscoring the importance of modeling temporal forgetting and learner profiles in LLM-based tutoring systems.
Active Domain Knowledge Acquisition with \$100 Budget: Enhancing LLMs via Cost-Efficient, Expert-Involved Interaction in Sensitive Domains
Aug 24, 2025Abstract:Large Language Models (LLMs) have demonstrated an impressive level of general knowledge. However, they often struggle in highly specialized and cost-sensitive domains such as drug discovery and rare disease research due to the lack of expert knowledge. In this paper, we propose a novel framework (PU-ADKA) designed to efficiently enhance domain-specific LLMs by actively engaging domain experts within a fixed budget. Unlike traditional fine-tuning approaches, PU-ADKA selectively identifies and queries the most appropriate expert from a team, taking into account each expert's availability, knowledge boundaries, and consultation costs. We train PU-ADKA using simulations on PubMed data and validate it through both controlled expert interactions and real-world deployment with a drug development team, demonstrating its effectiveness in enhancing LLM performance in specialized domains under strict budget constraints. In addition to outlining our methodological innovations and experimental results, we introduce a new benchmark dataset, CKAD, for cost-effective LLM domain knowledge acquisition to foster further research in this challenging area.
LeCoDe: A Benchmark Dataset for Interactive Legal Consultation Dialogue Evaluation
May 26, 2025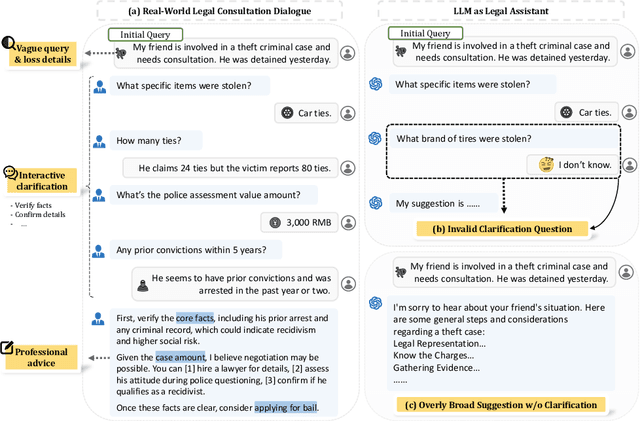
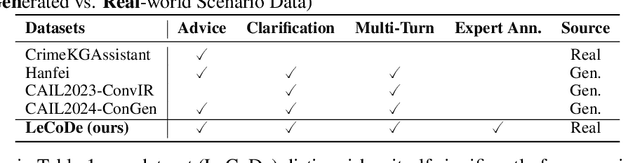


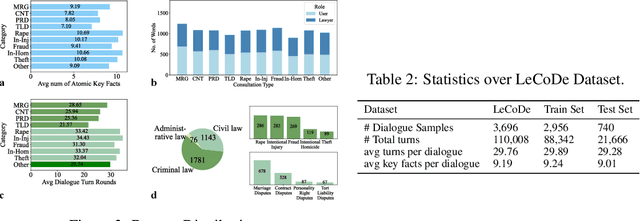
Abstract:Legal consultation is essential for safeguarding individual rights and ensuring access to justice, yet remains costly and inaccessible to many individuals due to the shortage of professionals. While recent advances in Large Language Models (LLMs) offer a promising path toward scalable, low-cost legal assistance, current systems fall short in handling the interactive and knowledge-intensive nature of real-world consultations. To address these challenges, we introduce LeCoDe, a real-world multi-turn benchmark dataset comprising 3,696 legal consultation dialogues with 110,008 dialogue turns, designed to evaluate and improve LLMs' legal consultation capability. With LeCoDe, we innovatively collect live-streamed consultations from short-video platforms, providing authentic multi-turn legal consultation dialogues. The rigorous annotation by legal experts further enhances the dataset with professional insights and expertise. Furthermore, we propose a comprehensive evaluation framework that assesses LLMs' consultation capabilities in terms of (1) clarification capability and (2) professional advice quality. This unified framework incorporates 12 metrics across two dimensions. Through extensive experiments on various general and domain-specific LLMs, our results reveal significant challenges in this task, with even state-of-the-art models like GPT-4 achieving only 39.8% recall for clarification and 59% overall score for advice quality, highlighting the complexity of professional consultation scenarios. Based on these findings, we further explore several strategies to enhance LLMs' legal consultation abilities. Our benchmark contributes to advancing research in legal domain dialogue systems, particularly in simulating more real-world user-expert interactions.
CLaDMoP: Learning Transferrable Models from Successful Clinical Trials via LLMs
May 24, 2025Abstract:Many existing models for clinical trial outcome prediction are optimized using task-specific loss functions on trial phase-specific data. While this scheme may boost prediction for common diseases and drugs, it can hinder learning of generalizable representations, leading to more false positives/negatives. To address this limitation, we introduce CLaDMoP, a new pre-training approach for clinical trial outcome prediction, alongside the Successful Clinical Trials dataset(SCT), specifically designed for this task. CLaDMoP leverages a Large Language Model-to encode trials' eligibility criteria-linked to a lightweight Drug-Molecule branch through a novel multi-level fusion technique. To efficiently fuse long embeddings across levels, we incorporate a grouping block, drastically reducing computational overhead. CLaDMoP avoids reliance on task-specific objectives by pre-training on a "pair matching" proxy task. Compared to established zero-shot and few-shot baselines, our method significantly improves both PR-AUC and ROC-AUC, especially for phase I and phase II trials. We further evaluate and perform ablation on CLaDMoP after Parameter-Efficient Fine-Tuning, comparing it to state-of-the-art supervised baselines, including MEXA-CTP, on the Trial Outcome Prediction(TOP) benchmark. CLaDMoP achieves up to 10.5% improvement in PR-AUC and 3.6% in ROC-AUC, while attaining comparable F1 score to MEXA-CTP, highlighting its potential for clinical trial outcome prediction. Code and SCT dataset can be downloaded from https://github.com/murai-lab/CLaDMoP.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge