Xiaoyang Chen
Rank4Gen: RAG-Preference-Aligned Document Set Selection and Ranking
Jan 16, 2026Abstract:In the RAG paradigm, the information retrieval module provides context for generators by retrieving and ranking multiple documents to support the aggregation of evidence. However, existing ranking models are primarily optimized for query--document relevance, which often misaligns with generators' preferences for evidence selection and citation, limiting their impact on response quality. Moreover, most approaches do not account for preference differences across generators, resulting in unstable cross-generator performance. We propose \textbf{Rank4Gen}, a generator-aware ranker for RAG that targets the goal of \emph{Ranking for Generators}. Rank4Gen introduces two key preference modeling strategies: (1) \textbf{From Ranking Relevance to Response Quality}, which optimizes ranking with respect to downstream response quality rather than query--document relevance; and (2) \textbf{Generator-Specific Preference Modeling}, which conditions a single ranker on different generators to capture their distinct ranking preferences. To enable such modeling, we construct \textbf{PRISM}, a dataset built from multiple open-source corpora and diverse downstream generators. Experiments on five challenging and recent RAG benchmarks demonstrate that RRank4Gen achieves strong and competitive performance for complex evidence composition in RAG.
Accelerating Multi-modal LLM Gaming Performance via Input Prediction and Mishit Correction
Dec 19, 2025Abstract:Real-time sequential control agents are often bottlenecked by inference latency. Even modest per-step planning delays can destabilize control and degrade overall performance. We propose a speculation-and-correction framework that adapts the predict-then-verify philosophy of speculative execution to model-based control with TD-MPC2. At each step, a pretrained world model and latent-space MPC planner generate a short-horizon action queue together with predicted latent rollouts, allowing the agent to execute multiple planned actions without immediate replanning. When a new observation arrives, the system measures the mismatch between the encoded real latent state and the queued predicted latent. For small to moderate mismatch, a lightweight learned corrector applies a residual update to the speculative action, distilled offline from a replanning teacher. For large mismatch, the agent safely falls back to full replanning and clears stale action queues. We study both a gated two-tower MLP corrector and a temporal Transformer corrector to address local errors and systematic drift. Experiments on the DMC Humanoid-Walk task show that our method reduces the number of planning inferences from 500 to 282, improves end-to-end step latency by 25 percent, and maintains strong control performance with only a 7.1 percent return reduction. Ablation results demonstrate that speculative execution without correction is unreliable over longer horizons, highlighting the necessity of mismatch-aware correction for robust latency reduction.
Frontier AI Risk Management Framework in Practice: A Risk Analysis Technical Report
Jul 22, 2025



Abstract:To understand and identify the unprecedented risks posed by rapidly advancing artificial intelligence (AI) models, this report presents a comprehensive assessment of their frontier risks. Drawing on the E-T-C analysis (deployment environment, threat source, enabling capability) from the Frontier AI Risk Management Framework (v1.0) (SafeWork-F1-Framework), we identify critical risks in seven areas: cyber offense, biological and chemical risks, persuasion and manipulation, uncontrolled autonomous AI R\&D, strategic deception and scheming, self-replication, and collusion. Guided by the "AI-$45^\circ$ Law," we evaluate these risks using "red lines" (intolerable thresholds) and "yellow lines" (early warning indicators) to define risk zones: green (manageable risk for routine deployment and continuous monitoring), yellow (requiring strengthened mitigations and controlled deployment), and red (necessitating suspension of development and/or deployment). Experimental results show that all recent frontier AI models reside in green and yellow zones, without crossing red lines. Specifically, no evaluated models cross the yellow line for cyber offense or uncontrolled AI R\&D risks. For self-replication, and strategic deception and scheming, most models remain in the green zone, except for certain reasoning models in the yellow zone. In persuasion and manipulation, most models are in the yellow zone due to their effective influence on humans. For biological and chemical risks, we are unable to rule out the possibility of most models residing in the yellow zone, although detailed threat modeling and in-depth assessment are required to make further claims. This work reflects our current understanding of AI frontier risks and urges collective action to mitigate these challenges.
DeepMath-Creative: A Benchmark for Evaluating Mathematical Creativity of Large Language Models
May 13, 2025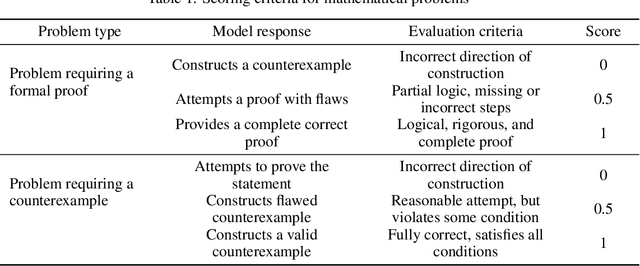
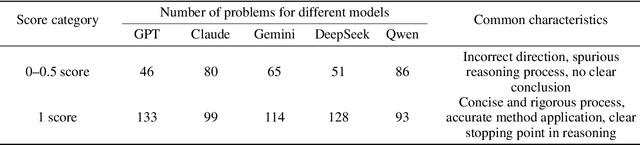
Abstract:To advance the mathematical proficiency of large language models (LLMs), the DeepMath team has launched an open-source initiative aimed at developing an open mathematical LLM and systematically evaluating its mathematical creativity. This paper represents the initial contribution of this initiative. While recent developments in mathematical LLMs have predominantly emphasized reasoning skills, as evidenced by benchmarks on elementary to undergraduate-level mathematical tasks, the creative capabilities of these models have received comparatively little attention, and evaluation datasets remain scarce. To address this gap, we propose an evaluation criteria for mathematical creativity and introduce DeepMath-Creative, a novel, high-quality benchmark comprising constructive problems across algebra, geometry, analysis, and other domains. We conduct a systematic evaluation of mainstream LLMs' creative problem-solving abilities using this dataset. Experimental results show that even under lenient scoring criteria -- emphasizing core solution components and disregarding minor inaccuracies, such as small logical gaps, incomplete justifications, or redundant explanations -- the best-performing model, O3 Mini, achieves merely 70% accuracy, primarily on basic undergraduate-level constructive tasks. Performance declines sharply on more complex problems, with models failing to provide substantive strategies for open problems. These findings suggest that, although current LLMs display a degree of constructive proficiency on familiar and lower-difficulty problems, such performance is likely attributable to the recombination of memorized patterns rather than authentic creative insight or novel synthesis.
3D-AffordanceLLM: Harnessing Large Language Models for Open-Vocabulary Affordance Detection in 3D Worlds
Feb 27, 2025Abstract:3D Affordance detection is a challenging problem with broad applications on various robotic tasks. Existing methods typically formulate the detection paradigm as a label-based semantic segmentation task. This paradigm relies on predefined labels and lacks the ability to comprehend complex natural language, resulting in limited generalization in open-world scene. To address these limitations, we reformulate the traditional affordance detection paradigm into \textit{Instruction Reasoning Affordance Segmentation} (IRAS) task. This task is designed to output a affordance mask region given a query reasoning text, which avoids fixed categories of input labels. We accordingly propose the \textit{3D-AffordanceLLM} (3D-ADLLM), a framework designed for reasoning affordance detection in 3D open-scene. Specifically, 3D-ADLLM introduces large language models (LLMs) to 3D affordance perception with a custom-designed decoder for generating affordance masks, thus achieving open-world reasoning affordance detection. In addition, given the scarcity of 3D affordance datasets for training large models, we seek to extract knowledge from general segmentation data and transfer it to affordance detection. Thus, we propose a multi-stage training strategy that begins with a novel pre-training task, i.e., \textit{Referring Object Part Segmentation}~(ROPS). This stage is designed to equip the model with general recognition and segmentation capabilities at the object-part level. Then followed by fine-tuning with the IRAS task, 3D-ADLLM obtains the reasoning ability for affordance detection. In summary, 3D-ADLLM leverages the rich world knowledge and human-object interaction reasoning ability of LLMs, achieving approximately an 8\% improvement in mIoU on open-vocabulary affordance detection tasks.
Match, Compare, or Select? An Investigation of Large Language Models for Entity Matching
May 27, 2024Abstract:Entity matching (EM) is a critical step in entity resolution. Recently, entity matching based on large language models (LLMs) has shown great promise. However, current LLM-based entity matching approaches typically follow a binary matching paradigm that ignores the global consistency between different records. In this paper, we investigate various methodologies for LLM-based entity matching that incorporate record interactions from different perspectives. Specifically, we comprehensively compare three representative strategies: matching, comparing, and selecting, and analyze their respective advantages and challenges in diverse scenarios. Based on our findings, we further design a compositional entity matching (ComEM) framework that leverages the composition of multiple strategies and LLMs. In this way, ComEM can benefit from the advantages of different sides and achieve improvements in both effectiveness and efficiency. Experimental results show that ComEM not only achieves significant performance gains on various datasets but also reduces the cost of LLM-based entity matching in real-world application.
A Classifier-Free Incremental Learning Framework for Scalable Medical Image Segmentation
May 25, 2024



Abstract:Current methods for developing foundation models in medical image segmentation rely on two primary assumptions: a fixed set of classes and the immediate availability of a substantial and diverse training dataset. However, this can be impractical due to the evolving nature of imaging technology and patient demographics, as well as labor-intensive data curation, limiting their practical applicability and scalability. To address these challenges, we introduce a novel segmentation paradigm enabling the segmentation of a variable number of classes within a single classifier-free network, featuring an architecture independent of class number. This network is trained using contrastive learning and produces discriminative feature representations that facilitate straightforward interpretation. Additionally, we integrate this strategy into a knowledge distillation-based incremental learning framework, facilitating the gradual assimilation of new information from non-stationary data streams while avoiding catastrophic forgetting. Our approach provides a unified solution for tackling both class- and domain-incremental learning scenarios. We demonstrate the flexibility of our method in handling varying class numbers within a unified network and its capacity for incremental learning. Experimental results on an incompletely annotated, multi-modal, multi-source dataset for medical image segmentation underscore its superiority over state-of-the-art alternative approaches.
Towards Universal Dense Blocking for Entity Resolution
Apr 25, 2024Abstract:Blocking is a critical step in entity resolution, and the emergence of neural network-based representation models has led to the development of dense blocking as a promising approach for exploring deep semantics in blocking. However, previous advanced self-supervised dense blocking approaches require domain-specific training on the target domain, which limits the benefits and rapid adaptation of these methods. To address this issue, we propose UniBlocker, a dense blocker that is pre-trained on a domain-independent, easily-obtainable tabular corpus using self-supervised contrastive learning. By conducting domain-independent pre-training, UniBlocker can be adapted to various downstream blocking scenarios without requiring domain-specific fine-tuning. To evaluate the universality of our entity blocker, we also construct a new benchmark covering a wide range of blocking tasks from multiple domains and scenarios. Our experiments show that the proposed UniBlocker, without any domain-specific learning, significantly outperforms previous self- and unsupervised dense blocking methods and is comparable and complementary to the state-of-the-art sparse blocking methods.
Spiral of Silences: How is Large Language Model Killing Information Retrieval? -- A Case Study on Open Domain Question Answering
Apr 18, 2024



Abstract:The practice of Retrieval-Augmented Generation (RAG), which integrates Large Language Models (LLMs) with retrieval systems, has become increasingly prevalent. However, the repercussions of LLM-derived content infiltrating the web and influencing the retrieval-generation feedback loop are largely uncharted territories. In this study, we construct and iteratively run a simulation pipeline to deeply investigate the short-term and long-term effects of LLM text on RAG systems. Taking the trending Open Domain Question Answering (ODQA) task as a point of entry, our findings reveal a potential digital "Spiral of Silence" effect, with LLM-generated text consistently outperforming human-authored content in search rankings, thereby diminishing the presence and impact of human contributions online. This trend risks creating an imbalanced information ecosystem, where the unchecked proliferation of erroneous LLM-generated content may result in the marginalization of accurate information. We urge the academic community to take heed of this potential issue, ensuring a diverse and authentic digital information landscape.
Reconstruction of Cortical Surfaces with Spherical Topology from Infant Brain MRI via Recurrent Deformation Learning
Dec 10, 2023



Abstract:Cortical surface reconstruction (CSR) from MRI is key to investigating brain structure and function. While recent deep learning approaches have significantly improved the speed of CSR, a substantial amount of runtime is still needed to map the cortex to a topologically-correct spherical manifold to facilitate downstream geometric analyses. Moreover, this mapping is possible only if the topology of the surface mesh is homotopic to a sphere. Here, we present a method for simultaneous CSR and spherical mapping efficiently within seconds. Our approach seamlessly connects two sub-networks for white and pial surface generation. Residual diffeomorphic deformations are learned iteratively to gradually warp a spherical template mesh to the white and pial surfaces while preserving mesh topology and uniformity. The one-to-one vertex correspondence between the template sphere and the cortical surfaces allows easy and direct mapping of geometric features like convexity and curvature to the sphere for visualization and downstream processing. We demonstrate the efficacy of our approach on infant brain MRI, which poses significant challenges to CSR due to tissue contrast changes associated with rapid brain development during the first postnatal year. Performance evaluation based on a dataset of infants from 0 to 12 months demonstrates that our method substantially enhances mesh regularity and reduces geometric errors, outperforming state-of-the-art deep learning approaches, all while maintaining high computational efficiency.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge