Wuyang Li
EverAnimate: Minute-Scale Human Animation via Latent Flow Restoration
May 14, 2026Abstract:We propose EverAnimate, an efficient post-training method for long-horizon animated video generation that preserves visual quality and character identity. Long-form animation remains challenging because highly dynamic human motion must be synthesized against relatively static environments, making chunk-based generation prone to accumulated drift: (i) low-level quality drift, such as progressive degradation of static backgrounds, and (ii) high-level semantic drift, such as inconsistent character identity and view-dependent attributes. To address this issue, EverAnimate restores drifted flow trajectories by anchoring generation to a persistent latent context memory, consisting of two complementary mechanisms. (i) Persistent Latent Propagation maintains a context memory across chunks to propagate identity and motion in latent space while mitigating temporal forgetting. (ii) Restorative Flow Matching introduces an implicit restoration objective during sampling through velocity adjustment, improving within-chunk fidelity. With only lightweight LoRA tuning, EverAnimate outperforms state-of-the-art long-animation methods in both short- and long-horizon settings: at 10 seconds, it improves PSNR/SSIM by 8%/7% and reduces LPIPS/FID by 22%/11%; at 90 seconds, the gains increase to 15%/15% and 32%/27%, respectively.
Grounded World Model for Semantically Generalizable Planning
Apr 13, 2026Abstract:In Model Predictive Control (MPC), world models predict the future outcomes of various action proposals, which are then scored to guide the selection of the optimal action. For visuomotor MPC, the score function is a distance metric between a predicted image and a goal image, measured in the latent space of a pretrained vision encoder like DINO and JEPA. However, it is challenging to obtain the goal image in advance of the task execution, particularly in new environments. Additionally, conveying the goal through an image offers limited interactivity compared with natural language. In this work, we propose to learn a Grounded World Model (GWM) in a vision-language-aligned latent space. As a result, each proposed action is scored based on how close its future outcome is to the task instruction, reflected by the similarity of embeddings. This approach transforms the visuomotor MPC to a VLA that surpasses VLM-based VLAs in semantic generalization. On the proposed WISER benchmark, GWM-MPC achieves a 87% success rate on the test set comprising 288 tasks that feature unseen visual signals and referring expressions, yet remain solvable with motions demonstrated during training. In contrast, traditional VLAs achieve an average success rate of 22%, even though they overfit the training set with a 90% success rate.
Harnessing Lightweight Transformer with Contextual Synergic Enhancement for Efficient 3D Medical Image Segmentation
Mar 24, 2026Abstract:Transformers have shown remarkable performance in 3D medical image segmentation, but their high computational requirements and need for large amounts of labeled data limit their applicability. To address these challenges, we consider two crucial aspects: model efficiency and data efficiency. Specifically, we propose Light-UNETR, a lightweight transformer designed to achieve model efficiency. Light-UNETR features a Lightweight Dimension Reductive Attention (LIDR) module, which reduces spatial and channel dimensions while capturing both global and local features via multi-branch attention. Additionally, we introduce a Compact Gated Linear Unit (CGLU) to selectively control channel interaction with minimal parameters. Furthermore, we introduce a Contextual Synergic Enhancement (CSE) learning strategy, which aims to boost the data efficiency of Transformers. It first leverages the extrinsic contextual information to support the learning of unlabeled data with Attention-Guided Replacement, then applies Spatial Masking Consistency that utilizes intrinsic contextual information to enhance the spatial context reasoning for unlabeled data. Extensive experiments on various benchmarks demonstrate the superiority of our approach in both performance and efficiency. For example, with only 10% labeled data on the Left Atrial Segmentation dataset, our method surpasses BCP by 1.43% Jaccard while drastically reducing the FLOPs by 90.8% and parameters by 85.8%. Code is released at https://github.com/CUHK-AIM-Group/Light-UNETR.
PreferRec: Learning and Transferring Pareto Preferences for Multi-objective Re-ranking
Mar 23, 2026Abstract:Multi-objective re-ranking has become a critical component of modern multi-stage recommender systems, as it tasked to balance multiple conflicting objectives such as accuracy, diversity, and fairness. Existing multi-objective re-ranking methods typically optimize aggregate objectives at the item level using static or handcrafted preference weights. This design overlooks that users inherently exhibit Pareto-optimal preferences at the intent level, reflecting personalized trade-offs among objectives rather than fixed weight combinations. Moreover, most approaches treat re-ranking task for each user as an isolated problem, and repeatedly learn the preferences from scratch. Such a paradigm not only incurs high computational cost, but also ignores the fact that users often share similar preference trade-off structures across objectives. Inspired by the existence of homogeneous multi-objective optimization spaces where Pareto-optimal patterns are transferable, we propose PreferRec, a novel framework that explicitly models and transfers Pareto preferences across users. Specifically, PreferRec is built upon three tightly coupled components: Preference-Aware Pareto Learning aims to capture user intrinsic trade-offs among multiple conflicting objectives at the intent level. By learning Pareto preference representations from re-ranking populations, this component explicitly models how users prioritize different objectives under diverse contexts. Knowledge-Guided Transfer facilitates efficient cross-user knowledge transfer by distilling shared optimization patterns across homogeneous optimization spaces. The transferred knowledge is then used to guide solution selection and personalized re-ranking, biasing the optimization process toward high-quality regions of the Pareto front while preserving user-specific preference characteristics.
Factorized Video Generation: Decoupling Scene Construction and Temporal Synthesis in Text-to-Video Diffusion Models
Dec 18, 2025



Abstract:State-of-the-art Text-to-Video (T2V) diffusion models can generate visually impressive results, yet they still frequently fail to compose complex scenes or follow logical temporal instructions. In this paper, we argue that many errors, including apparent motion failures, originate from the model's inability to construct a semantically correct or logically consistent initial frame. We introduce Factorized Video Generation (FVG), a pipeline that decouples these tasks by decomposing the Text-to-Video generation into three specialized stages: (1) Reasoning, where a Large Language Model (LLM) rewrites the video prompt to describe only the initial scene, resolving temporal ambiguities; (2) Composition, where a Text-to-Image (T2I) model synthesizes a high-quality, compositionally-correct anchor frame from this new prompt; and (3) Temporal Synthesis, where a video model, finetuned to understand this anchor, focuses its entire capacity on animating the scene and following the prompt. Our decomposed approach sets a new state-of-the-art on the T2V CompBench benchmark and significantly improves all tested models on VBench2. Furthermore, we show that visual anchoring allows us to cut the number of sampling steps by 70% without any loss in performance, leading to a substantial speed-up in sampling. Factorized Video Generation offers a simple yet practical path toward more efficient, robust, and controllable video synthesis
RAP: 3D Rasterization Augmented End-to-End Planning
Oct 05, 2025Abstract:Imitation learning for end-to-end driving trains policies only on expert demonstrations. Once deployed in a closed loop, such policies lack recovery data: small mistakes cannot be corrected and quickly compound into failures. A promising direction is to generate alternative viewpoints and trajectories beyond the logged path. Prior work explores photorealistic digital twins via neural rendering or game engines, but these methods are prohibitively slow and costly, and thus mainly used for evaluation. In this work, we argue that photorealism is unnecessary for training end-to-end planners. What matters is semantic fidelity and scalability: driving depends on geometry and dynamics, not textures or lighting. Motivated by this, we propose 3D Rasterization, which replaces costly rendering with lightweight rasterization of annotated primitives, enabling augmentations such as counterfactual recovery maneuvers and cross-agent view synthesis. To transfer these synthetic views effectively to real-world deployment, we introduce a Raster-to-Real feature-space alignment that bridges the sim-to-real gap. Together, these components form Rasterization Augmented Planning (RAP), a scalable data augmentation pipeline for planning. RAP achieves state-of-the-art closed-loop robustness and long-tail generalization, ranking first on four major benchmarks: NAVSIM v1/v2, Waymo Open Dataset Vision-based E2E Driving, and Bench2Drive. Our results show that lightweight rasterization with feature alignment suffices to scale E2E training, offering a practical alternative to photorealistic rendering. Project page: https://alan-lanfeng.github.io/RAP/.
See&Trek: Training-Free Spatial Prompting for Multimodal Large Language Model
Sep 19, 2025Abstract:We introduce SEE&TREK, the first training-free prompting framework tailored to enhance the spatial understanding of Multimodal Large Language Models (MLLMS) under vision-only constraints. While prior efforts have incorporated modalities like depth or point clouds to improve spatial reasoning, purely visualspatial understanding remains underexplored. SEE&TREK addresses this gap by focusing on two core principles: increasing visual diversity and motion reconstruction. For visual diversity, we conduct Maximum Semantic Richness Sampling, which employs an off-the-shell perception model to extract semantically rich keyframes that capture scene structure. For motion reconstruction, we simulate visual trajectories and encode relative spatial positions into keyframes to preserve both spatial relations and temporal coherence. Our method is training&GPU-free, requiring only a single forward pass, and can be seamlessly integrated into existing MLLM'S. Extensive experiments on the VSI-B ENCH and STI-B ENCH show that S EE &T REK consistently boosts various MLLM S performance across diverse spatial reasoning tasks with the most +3.5% improvement, offering a promising path toward stronger spatial intelligence.
VoxDet: Rethinking 3D Semantic Occupancy Prediction as Dense Object Detection
Jun 05, 2025Abstract:3D semantic occupancy prediction aims to reconstruct the 3D geometry and semantics of the surrounding environment. With dense voxel labels, prior works typically formulate it as a dense segmentation task, independently classifying each voxel. However, this paradigm neglects critical instance-centric discriminability, leading to instance-level incompleteness and adjacent ambiguities. To address this, we highlight a free lunch of occupancy labels: the voxel-level class label implicitly provides insight at the instance level, which is overlooked by the community. Motivated by this observation, we first introduce a training-free Voxel-to-Instance (VoxNT) trick: a simple yet effective method that freely converts voxel-level class labels into instance-level offset labels. Building on this, we further propose VoxDet, an instance-centric framework that reformulates the voxel-level occupancy prediction as dense object detection by decoupling it into two sub-tasks: offset regression and semantic prediction. Specifically, based on the lifted 3D volume, VoxDet first uses (a) Spatially-decoupled Voxel Encoder to generate disentangled feature volumes for the two sub-tasks, which learn task-specific spatial deformation in the densely projected tri-perceptive space. Then, we deploy (b) Task-decoupled Dense Predictor to address this task via dense detection. Here, we first regress a 4D offset field to estimate distances (6 directions) between voxels and object borders in the voxel space. The regressed offsets are then used to guide the instance-level aggregation in the classification branch, achieving instance-aware prediction. Experiments show that VoxDet can be deployed on both camera and LiDAR input, jointly achieving state-of-the-art results on both benchmarks. VoxDet is not only highly efficient, but also achieves 63.0 IoU on the SemanticKITTI test set, ranking 1st on the online leaderboard.
Track Any Anomalous Object: A Granular Video Anomaly Detection Pipeline
Jun 05, 2025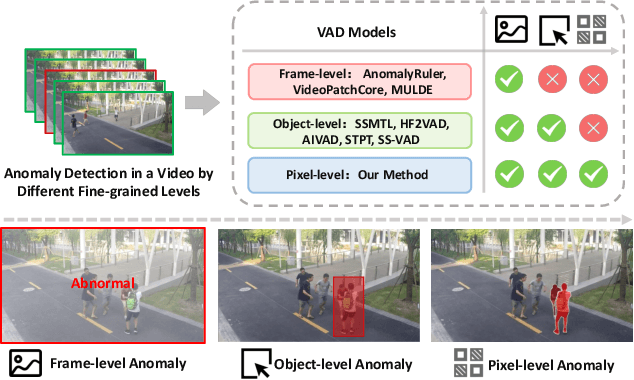
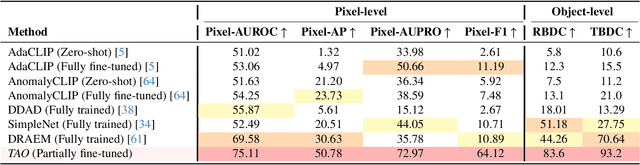
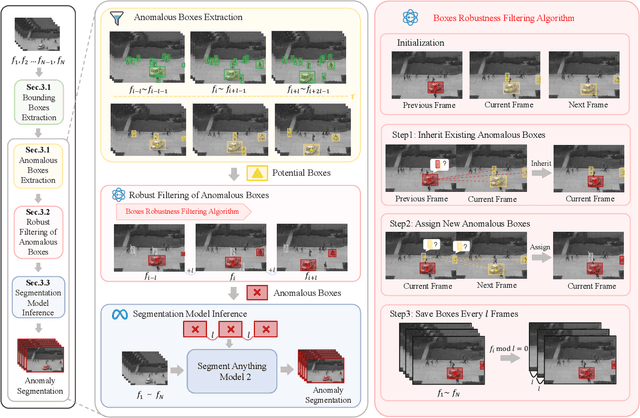
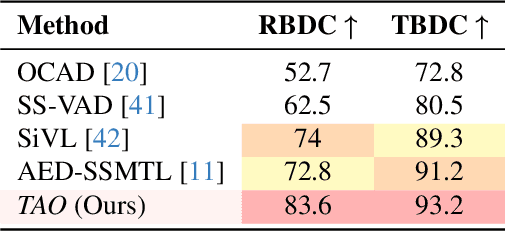
Abstract:Video anomaly detection (VAD) is crucial in scenarios such as surveillance and autonomous driving, where timely detection of unexpected activities is essential. Although existing methods have primarily focused on detecting anomalous objects in videos -- either by identifying anomalous frames or objects -- they often neglect finer-grained analysis, such as anomalous pixels, which limits their ability to capture a broader range of anomalies. To address this challenge, we propose a new framework called Track Any Anomalous Object (TAO), which introduces a granular video anomaly detection pipeline that, for the first time, integrates the detection of multiple fine-grained anomalous objects into a unified framework. Unlike methods that assign anomaly scores to every pixel, our approach transforms the problem into pixel-level tracking of anomalous objects. By linking anomaly scores to downstream tasks such as segmentation and tracking, our method removes the need for threshold tuning and achieves more precise anomaly localization in long and complex video sequences. Experiments demonstrate that TAO sets new benchmarks in accuracy and robustness. Project page available online.
FlexGS: Train Once, Deploy Everywhere with Many-in-One Flexible 3D Gaussian Splatting
Jun 04, 2025Abstract:3D Gaussian splatting (3DGS) has enabled various applications in 3D scene representation and novel view synthesis due to its efficient rendering capabilities. However, 3DGS demands relatively significant GPU memory, limiting its use on devices with restricted computational resources. Previous approaches have focused on pruning less important Gaussians, effectively compressing 3DGS but often requiring a fine-tuning stage and lacking adaptability for the specific memory needs of different devices. In this work, we present an elastic inference method for 3DGS. Given an input for the desired model size, our method selects and transforms a subset of Gaussians, achieving substantial rendering performance without additional fine-tuning. We introduce a tiny learnable module that controls Gaussian selection based on the input percentage, along with a transformation module that adjusts the selected Gaussians to complement the performance of the reduced model. Comprehensive experiments on ZipNeRF, MipNeRF and Tanks\&Temples scenes demonstrate the effectiveness of our approach. Code is available at https://flexgs.github.io.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge