Wenbing Huang
When to Update Your Model: Constrained Model-based Reinforcement Learning
Oct 15, 2022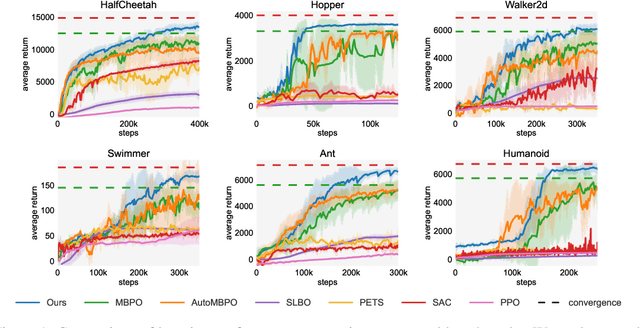
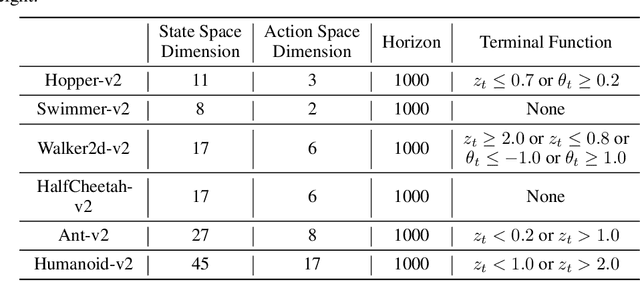

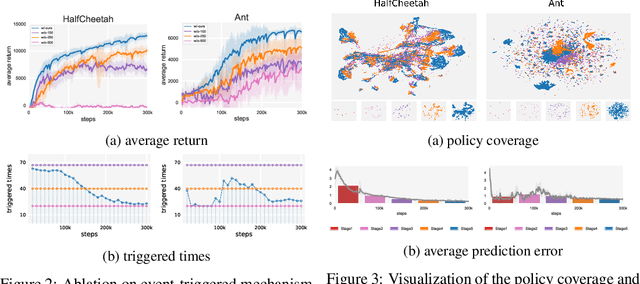
Abstract:Designing and analyzing model-based RL (MBRL) algorithms with guaranteed monotonic improvement has been challenging, mainly due to the interdependence between policy optimization and model learning. Existing discrepancy bounds generally ignore the impacts of model shifts, and their corresponding algorithms are prone to degrade performance by drastic model updating. In this work, we first propose a novel and general theoretical scheme for a non-decreasing performance guarantee of MBRL. Our follow-up derived bounds reveal the relationship between model shifts and performance improvement. These discoveries encourage us to formulate a constrained lower-bound optimization problem to permit the monotonicity of MBRL. A further example demonstrates that learning models from a dynamically-varying number of explorations benefit the eventual returns. Motivated by these analyses, we design a simple but effective algorithm CMLO (Constrained Model-shift Lower-bound Optimization), by introducing an event-triggered mechanism that flexibly determines when to update the model. Experiments show that CMLO surpasses other state-of-the-art methods and produces a boost when various policy optimization methods are employed.
Learning Active Camera for Multi-Object Navigation
Oct 14, 2022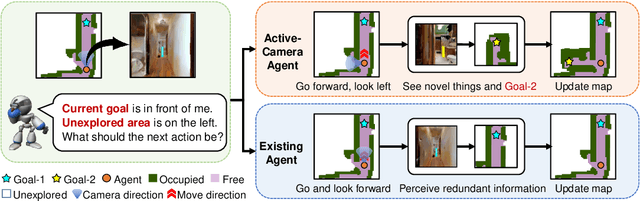
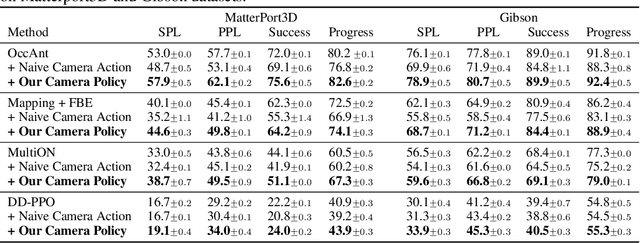
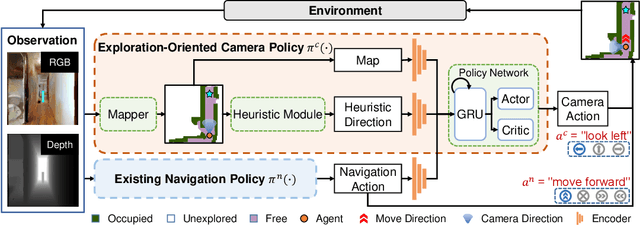
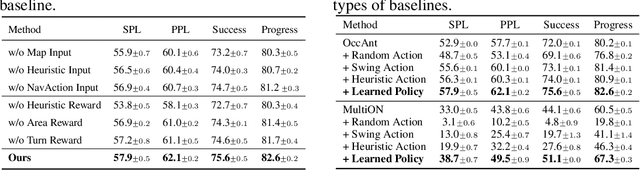
Abstract:Getting robots to navigate to multiple objects autonomously is essential yet difficult in robot applications. One of the key challenges is how to explore environments efficiently with camera sensors only. Existing navigation methods mainly focus on fixed cameras and few attempts have been made to navigate with active cameras. As a result, the agent may take a very long time to perceive the environment due to limited camera scope. In contrast, humans typically gain a larger field of view by looking around for a better perception of the environment. How to make robots perceive the environment as efficiently as humans is a fundamental problem in robotics. In this paper, we consider navigating to multiple objects more efficiently with active cameras. Specifically, we cast moving camera to a Markov Decision Process and reformulate the active camera problem as a reinforcement learning problem. However, we have to address two new challenges: 1) how to learn a good camera policy in complex environments and 2) how to coordinate it with the navigation policy. To address these, we carefully design a reward function to encourage the agent to explore more areas by moving camera actively. Moreover, we exploit human experience to infer a rule-based camera action to guide the learning process. Last, to better coordinate two kinds of policies, the camera policy takes navigation actions into account when making camera moving decisions. Experimental results show our camera policy consistently improves the performance of multi-object navigation over four baselines on two datasets.
Learning Physical Dynamics with Subequivariant Graph Neural Networks
Oct 13, 2022



Abstract:Graph Neural Networks (GNNs) have become a prevailing tool for learning physical dynamics. However, they still encounter several challenges: 1) Physical laws abide by symmetry, which is a vital inductive bias accounting for model generalization and should be incorporated into the model design. Existing simulators either consider insufficient symmetry, or enforce excessive equivariance in practice when symmetry is partially broken by gravity. 2) Objects in the physical world possess diverse shapes, sizes, and properties, which should be appropriately processed by the model. To tackle these difficulties, we propose a novel backbone, Subequivariant Graph Neural Network, which 1) relaxes equivariance to subequivariance by considering external fields like gravity, where the universal approximation ability holds theoretically; 2) introduces a new subequivariant object-aware message passing for learning physical interactions between multiple objects of various shapes in the particle-based representation; 3) operates in a hierarchical fashion, allowing for modeling long-range and complex interactions. Our model achieves on average over 3% enhancement in contact prediction accuracy across 8 scenarios on Physion and 2X lower rollout MSE on RigidFall compared with state-of-the-art GNN simulators, while exhibiting strong generalization and data efficiency.
Planning Assembly Sequence with Graph Transformer
Oct 12, 2022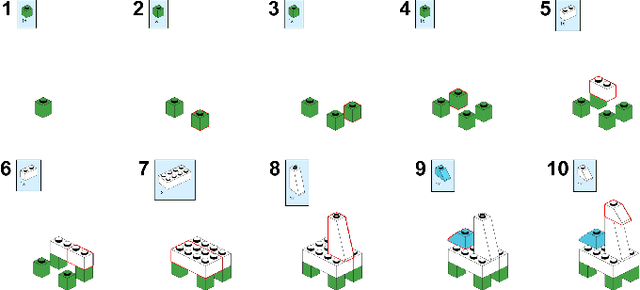
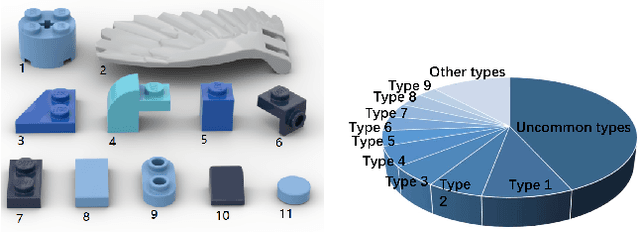
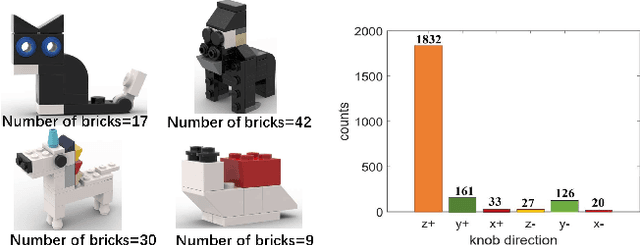

Abstract:Assembly sequence planning (ASP) is the essential process for modern manufacturing, proven to be NP-complete thus its effective and efficient solution has been a challenge for researchers in the field. In this paper, we present a graph-transformer based framework for the ASP problem which is trained and demonstrated on a self-collected ASP database. The ASP database contains a self-collected set of LEGO models. The LEGO model is abstracted to a heterogeneous graph structure after a thorough analysis of the original structure and feature extraction. The ground truth assembly sequence is first generated by brute-force search and then adjusted manually to in line with human rational habits. Based on this self-collected ASP dataset, we propose a heterogeneous graph-transformer framework to learn the latent rules for assembly planning. We evaluated the proposed framework in a series of experiment. The results show that the similarity of the predicted and ground truth sequences can reach 0.44, a medium correlation measured by Kendall's $\tau$. Meanwhile, we compared the different effects of node features and edge features and generated a feasible and reasonable assembly sequence as a benchmark for further research. Our data set and code is available on https://github.com/AIR-DISCOVER/ICRA\_ASP.
Benefits of Permutation-Equivariance in Auction Mechanisms
Oct 11, 2022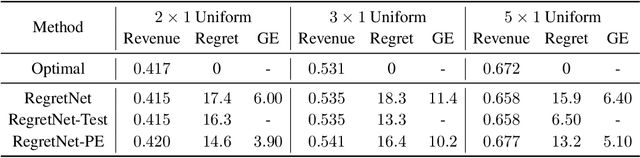
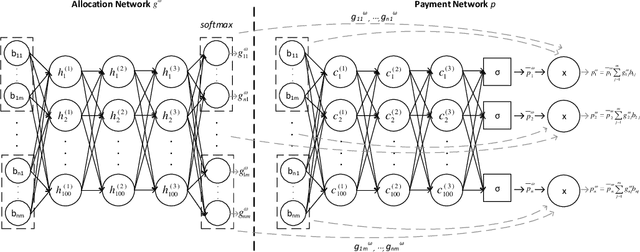
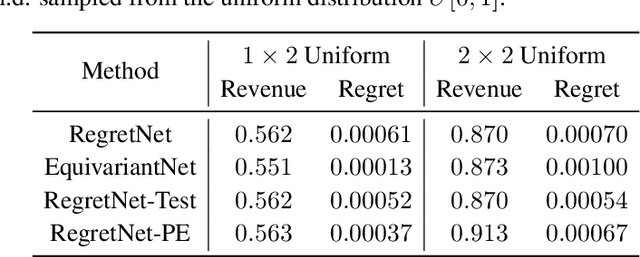
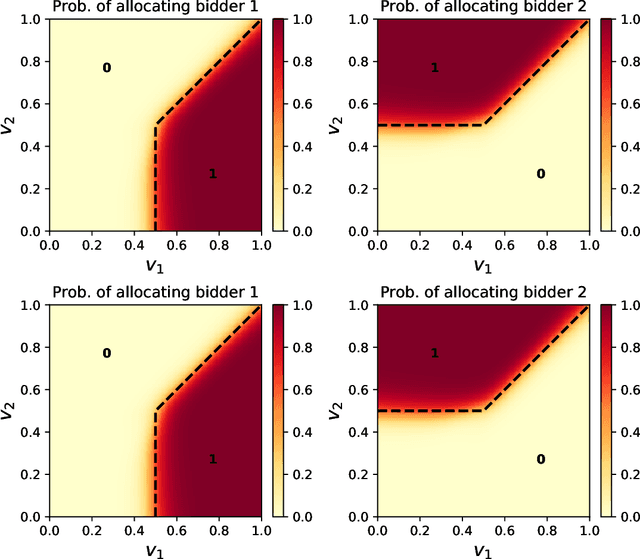
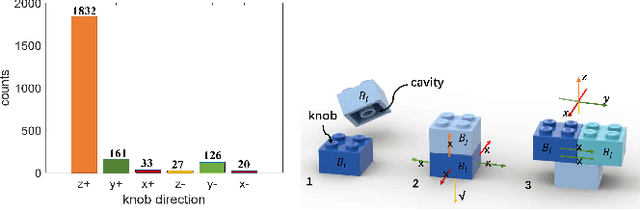
Abstract:Designing an incentive-compatible auction mechanism that maximizes the auctioneer's revenue while minimizes the bidders' ex-post regret is an important yet intricate problem in economics. Remarkable progress has been achieved through learning the optimal auction mechanism by neural networks. In this paper, we consider the popular additive valuation and symmetric valuation setting; i.e., the valuation for a set of items is defined as the sum of all items' valuations in the set, and the valuation distribution is invariant when the bidders and/or the items are permutated. We prove that permutation-equivariant neural networks have significant advantages: the permutation-equivariance decreases the expected ex-post regret, improves the model generalizability, while maintains the expected revenue invariant. This implies that the permutation-equivariance helps approach the theoretically optimal dominant strategy incentive compatible condition, and reduces the required sample complexity for desired generalization. Extensive experiments fully support our theory. To our best knowledge, this is the first work towards understanding the benefits of permutation-equivariance in auction mechanisms.
Bridged Transformer for Vision and Point Cloud 3D Object Detection
Oct 04, 2022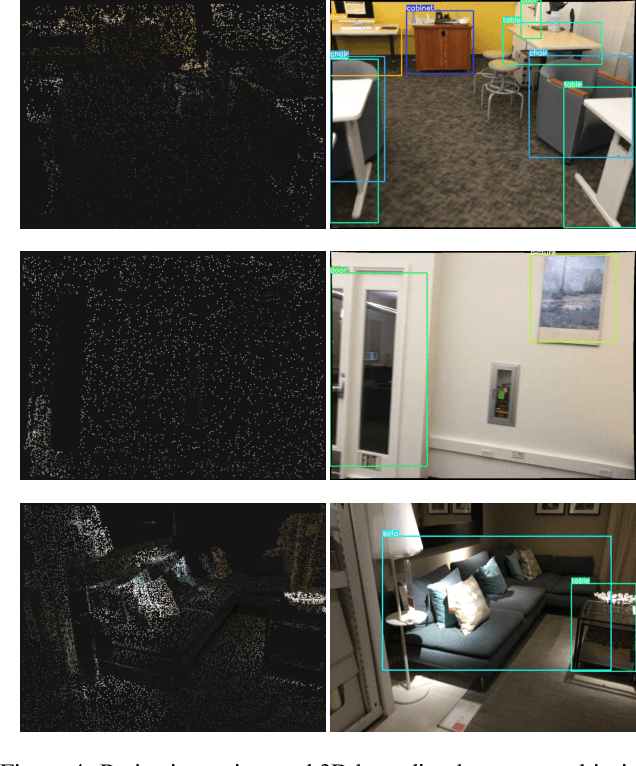
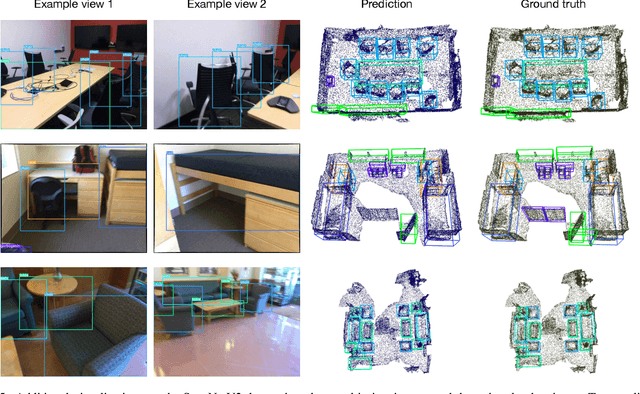

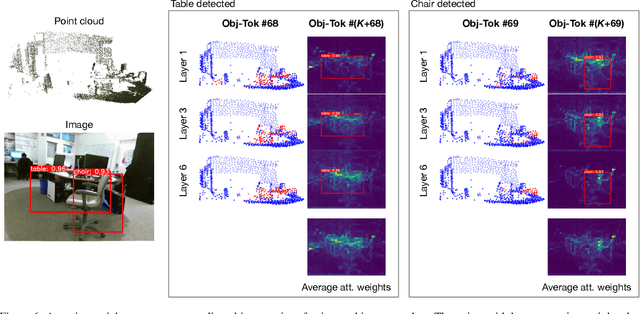
Abstract:3D object detection is a crucial research topic in computer vision, which usually uses 3D point clouds as input in conventional setups. Recently, there is a trend of leveraging multiple sources of input data, such as complementing the 3D point cloud with 2D images that often have richer color and fewer noises. However, due to the heterogeneous geometrics of the 2D and 3D representations, it prevents us from applying off-the-shelf neural networks to achieve multimodal fusion. To that end, we propose Bridged Transformer (BrT), an end-to-end architecture for 3D object detection. BrT is simple and effective, which learns to identify 3D and 2D object bounding boxes from both points and image patches. A key element of BrT lies in the utilization of object queries for bridging 3D and 2D spaces, which unifies different sources of data representations in Transformer. We adopt a form of feature aggregation realized by point-to-patch projections which further strengthen the correlations between images and points. Moreover, BrT works seamlessly for fusing the point cloud with multi-view images. We experimentally show that BrT surpasses state-of-the-art methods on SUN RGB-D and ScanNetV2 datasets.
Conditional Antibody Design as 3D Equivariant Graph Translation
Aug 12, 2022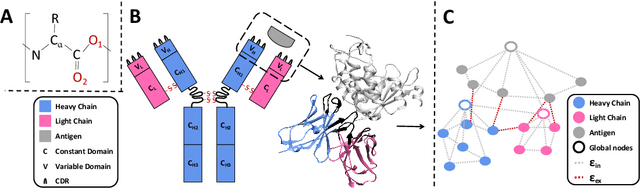
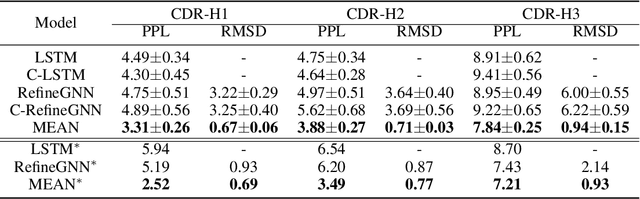
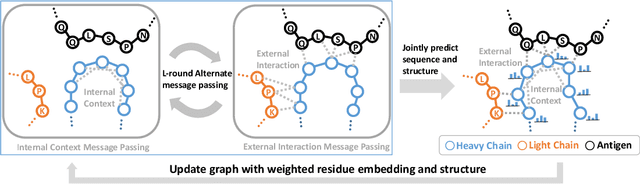
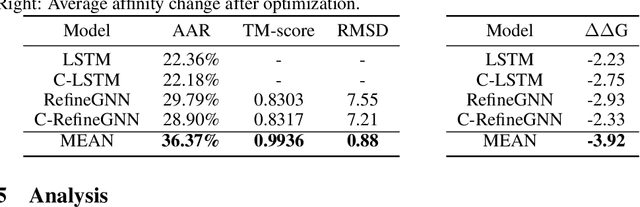
Abstract:Antibody design is valuable for therapeutic usage and biological research. Existing deep-learning-based methods encounter several key issues: 1) incomplete context for Complementarity-Determining Regions (CDRs) generation; 2) incapable of capturing the entire 3D geometry of the input structure; 3) inefficient prediction of the CDR sequences in an autoregressive manner. In this paper, we propose Multi-channel Equivariant Attention Network (MEAN), an end-to-end model that is able to co-design 1D sequences and 3D structures of CDRs. To be specific, MEAN formulates antibody design as a conditional graph translation problem by importing extra components including the target antigen and the light chain of the antibody. Then, MEAN resorts to E(3)-equivariant message passing along with a proposed attention mechanism to better capture the geometrical correlation between different components. Finally, it outputs both the 1D sequences and 3D structure via a multi-round progressive full-shot scheme, which enjoys more efficiency against previous autoregressive approaches. Our method significantly surpasses state-of-the-art models in sequence and structure modeling, antigen-binding antibody design, and binding affinity optimization. Specifically, the relative improvement to baselines is about 22% in antigen-binding CDR design and 34% for affinity optimization.
3D Equivariant Molecular Graph Pretraining
Jul 20, 2022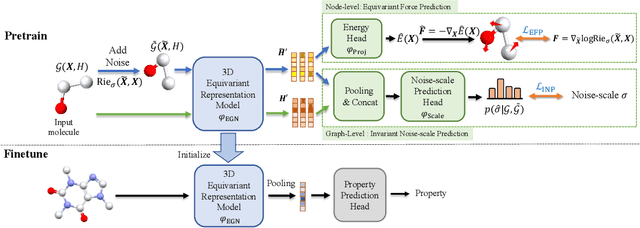
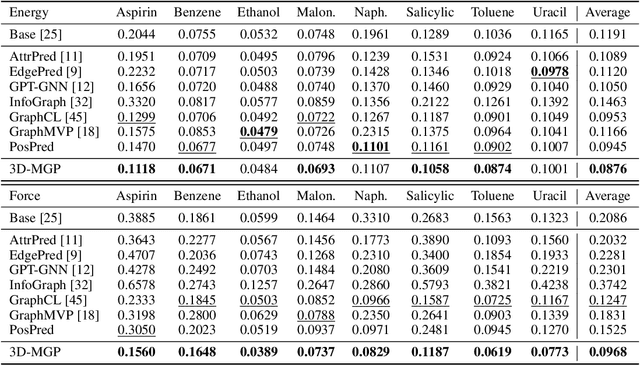
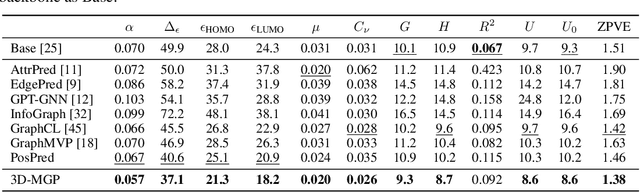
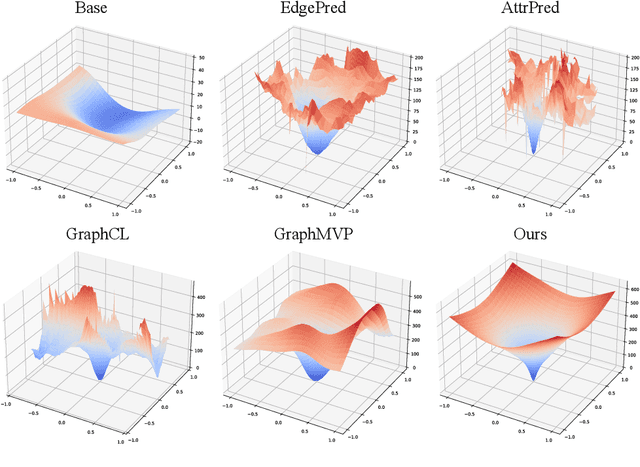
Abstract:Pretraining molecular representation models without labels is fundamental to various applications. Conventional methods mainly process 2D molecular graphs and focus solely on 2D tasks, making their pretrained models incapable of characterizing 3D geometry and thus defective for downstream 3D tasks. In this work, we tackle 3D molecular pretraining in a complete and novel sense. In particular, we first propose to adopt an equivariant energy-based model as the backbone for pretraining, which enjoys the merit of fulfilling the symmetry of 3D space. Then we develop a node-level pretraining loss for force prediction, where we further exploit the Riemann-Gaussian distribution to ensure the loss to be E(3)-invariant, enabling more robustness. Moreover, a graph-level noise scale prediction task is also leveraged to further promote the eventual performance. We evaluate our model pretrained from a large-scale 3D dataset GEOM-QM9 on two challenging 3D benchmarks: MD17 and QM9. The experimental results support the better efficacy of our method against current state-of-the-art pretraining approaches, and verify the validity of our design for each proposed component.
Similarity-aware Positive Instance Sampling for Graph Contrastive Pre-training
Jun 23, 2022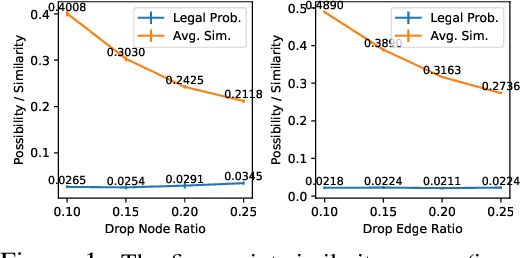
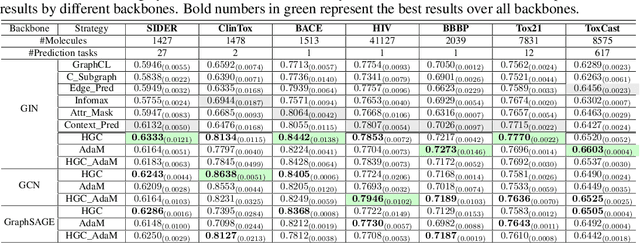
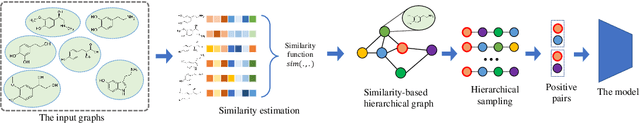
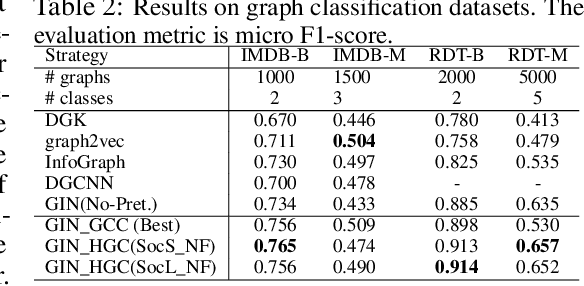
Abstract:Graph instance contrastive learning has been proved as an effective task for Graph Neural Network (GNN) pre-training. However, one key issue may seriously impede the representative power in existing works: Positive instances created by current methods often miss crucial information of graphs or even yield illegal instances (such as non-chemically-aware graphs in molecular generation). To remedy this issue, we propose to select positive graph instances directly from existing graphs in the training set, which ultimately maintains the legality and similarity to the target graphs. Our selection is based on certain domain-specific pair-wise similarity measurements as well as sampling from a hierarchical graph encoding similarity relations among graphs. Besides, we develop an adaptive node-level pre-training method to dynamically mask nodes to distribute them evenly in the graph. We conduct extensive experiments on $13$ graph classification and node classification benchmark datasets from various domains. The results demonstrate that the GNN models pre-trained by our strategies can outperform those trained-from-scratch models as well as the variants obtained by existing methods.
SNAKE: Shape-aware Neural 3D Keypoint Field
Jun 03, 2022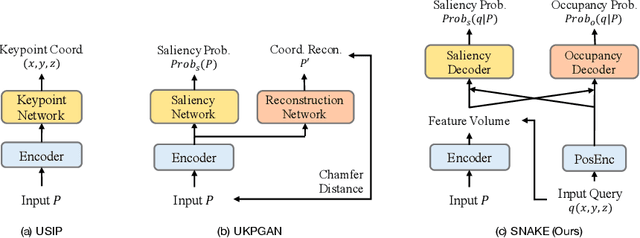
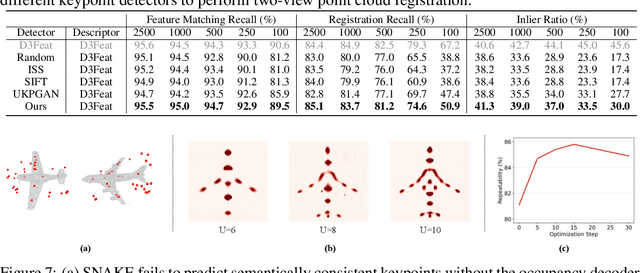
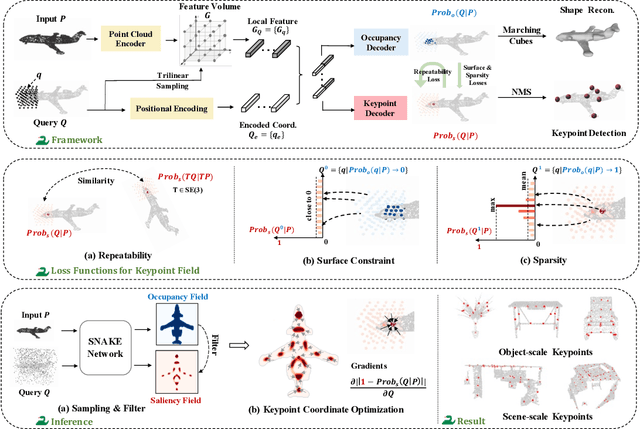
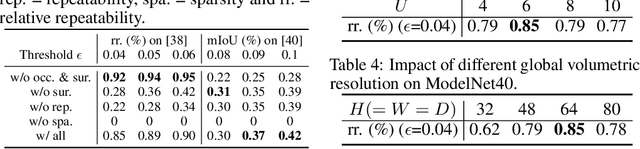
Abstract:Detecting 3D keypoints from point clouds is important for shape reconstruction, while this work investigates the dual question: can shape reconstruction benefit 3D keypoint detection? Existing methods either seek salient features according to statistics of different orders or learn to predict keypoints that are invariant to transformation. Nevertheless, the idea of incorporating shape reconstruction into 3D keypoint detection is under-explored. We argue that this is restricted by former problem formulations. To this end, a novel unsupervised paradigm named SNAKE is proposed, which is short for shape-aware neural 3D keypoint field. Similar to recent coordinate-based radiance or distance field, our network takes 3D coordinates as inputs and predicts implicit shape indicators and keypoint saliency simultaneously, thus naturally entangling 3D keypoint detection and shape reconstruction. We achieve superior performance on various public benchmarks, including standalone object datasets ModelNet40, KeypointNet, SMPL meshes and scene-level datasets 3DMatch and Redwood. Intrinsic shape awareness brings several advantages as follows. (1) SNAKE generates 3D keypoints consistent with human semantic annotation, even without such supervision. (2) SNAKE outperforms counterparts in terms of repeatability, especially when the input point clouds are down-sampled. (3) the generated keypoints allow accurate geometric registration, notably in a zero-shot setting. Codes are available at https://github.com/zhongcl-thu/SNAKE
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge