Shumin Deng
Continual Multimodal Knowledge Graph Construction
May 15, 2023Abstract:Multimodal Knowledge Graph Construction (MMKC) refers to the process of creating a structured representation of entities and relationships through multiple modalities such as text, images, videos, etc. However, existing MMKC models have limitations in handling the introduction of new entities and relations due to the dynamic nature of the real world. Moreover, most state-of-the-art studies in MMKC only consider entity and relation extraction from text data while neglecting other multi-modal sources. Meanwhile, the current continual setting for knowledge graph construction only consider entity and relation extraction from text data while neglecting other multi-modal sources. Therefore, there arises the need to explore the challenge of continuous multimodal knowledge graph construction to address the phenomenon of catastrophic forgetting and ensure the retention of past knowledge extracted from different forms of data. This research focuses on investigating this complex topic by developing lifelong multimodal benchmark datasets. Based on the empirical findings that several state-of-the-art MMKC models, when trained on multimedia data, might unexpectedly underperform compared to those solely utilizing textual resources in a continual setting, we propose a Lifelong MultiModal Consistent Transformer Framework (LMC) for continuous multimodal knowledge graph construction. By combining the advantages of consistent KGC strategies within the context of continual learning, we achieve greater balance between stability and plasticity. Our experiments demonstrate the superior performance of our method over prevailing continual learning techniques or multimodal approaches in dynamic scenarios. Code and datasets can be found at https://github.com/zjunlp/ContinueMKGC.
Reasoning with Language Model Prompting: A Survey
Dec 19, 2022



Abstract:Reasoning, as an essential ability for complex problem-solving, can provide back-end support for various real-world applications, such as medical diagnosis, negotiation, etc. This paper provides a comprehensive survey of cutting-edge research on reasoning with language model prompting. We introduce research works with comparisons and summaries and provide systematic resources to help beginners. We also discuss the potential reasons for emerging such reasoning abilities and highlight future research directions.
Schema-aware Reference as Prompt Improves Data-Efficient Relational Triple and Event Extraction
Oct 19, 2022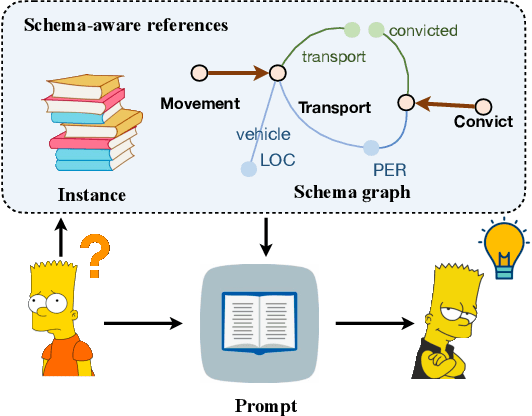
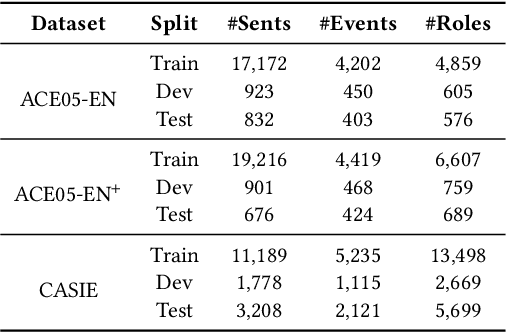
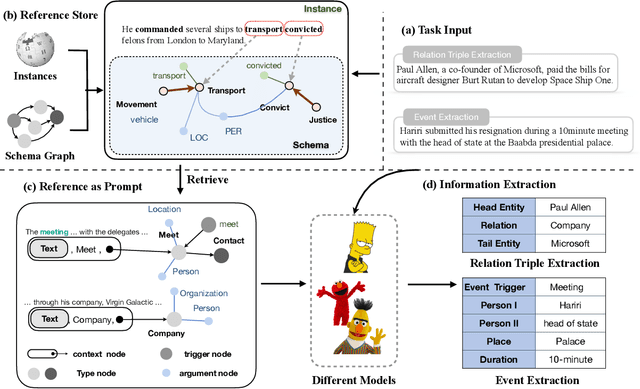
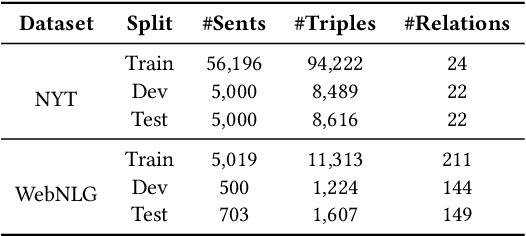
Abstract:Information Extraction, which aims to extract structural relational triple or event from unstructured texts, often suffers from data scarcity issues. With the development of pre-trained language models, many prompt-based approaches to data-efficient information extraction have been proposed and achieved impressive performance. However, existing prompt learning methods for information extraction are still susceptible to several potential limitations: (i) semantic gap between natural language and output structure knowledge with pre-defined schema; (ii) representation learning with locally individual instances limits the performance given the insufficient features. In this paper, we propose a novel approach of schema-aware Reference As Prompt (RAP), which dynamically leverage schema and knowledge inherited from global (few-shot) training data for each sample. Specifically, we propose a schema-aware reference store, which unifies symbolic schema and relevant textual instances. Then, we employ a dynamic reference integration module to retrieve pertinent knowledge from the datastore as prompts during training and inference. Experimental results demonstrate that RAP can be plugged into various existing models and outperforms baselines in low-resource settings on five datasets of relational triple extraction and event extraction. In addition, we provide comprehensive empirical ablations and case analysis regarding different types and scales of knowledge in order to better understand the mechanisms of RAP. Code is available in https://github.com/zjunlp/RAP.
PromptKG: A Prompt Learning Framework for Knowledge Graph Representation Learning and Application
Oct 01, 2022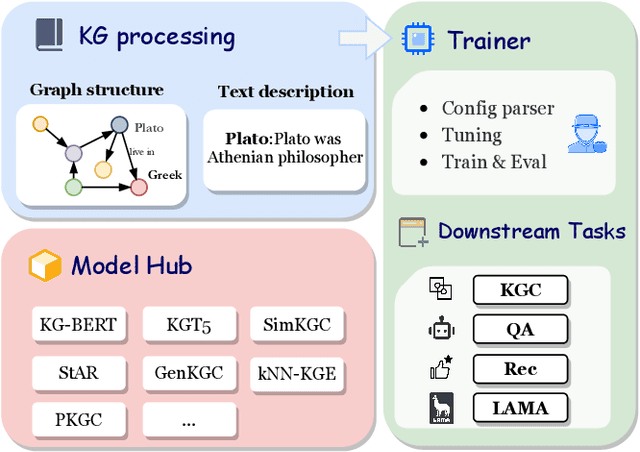
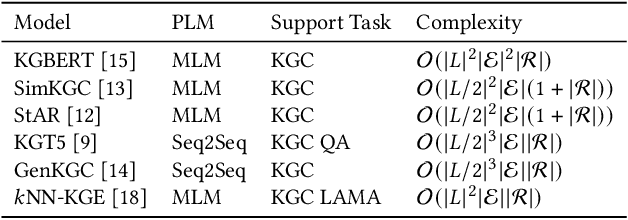
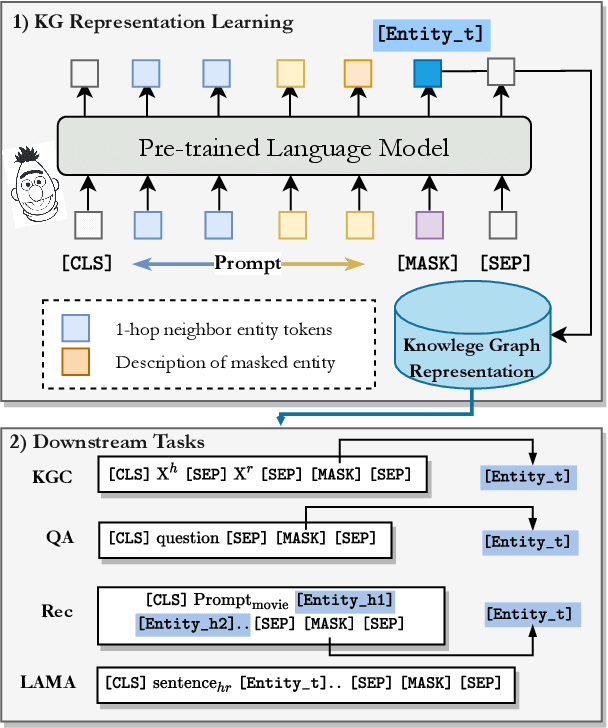
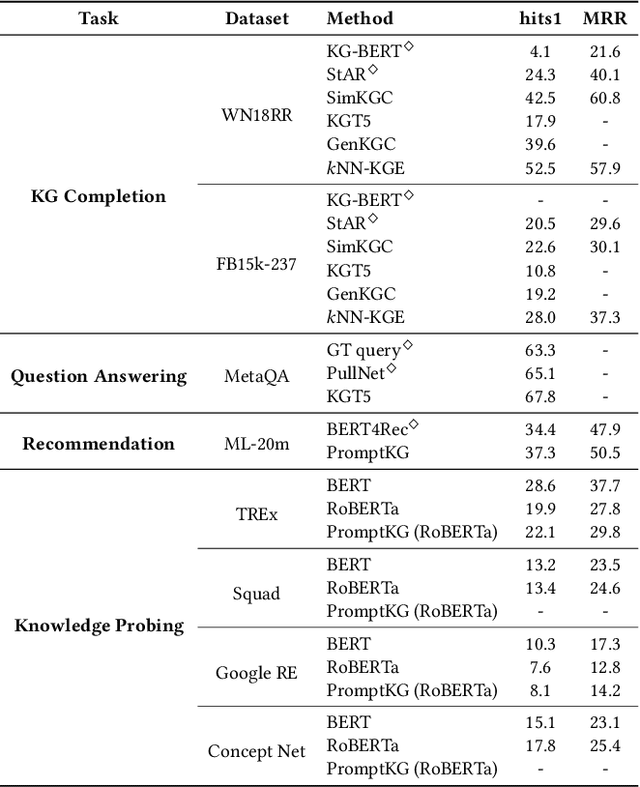
Abstract:Knowledge Graphs (KGs) often have two characteristics: heterogeneous graph structure and text-rich entity/relation information. KG representation models should consider graph structures and text semantics, but no comprehensive open-sourced framework is mainly designed for KG regarding informative text description. In this paper, we present PromptKG, a prompt learning framework for KG representation learning and application that equips the cutting-edge text-based methods, integrates a new prompt learning model and supports various tasks (e.g., knowledge graph completion, question answering, recommendation, and knowledge probing). PromptKG is publicly open-sourced at https://github.com/zjunlp/PromptKG with long-term technical support.
Multimodal Analogical Reasoning over Knowledge Graphs
Oct 01, 2022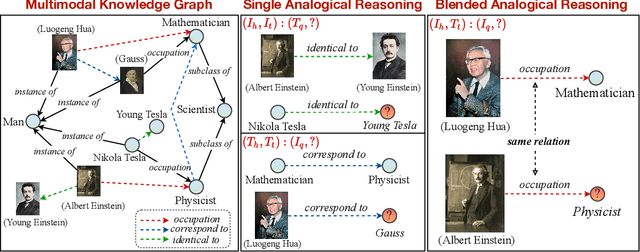
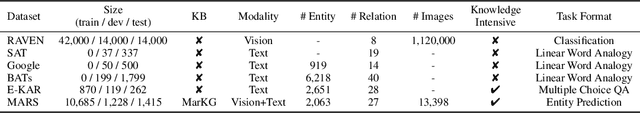

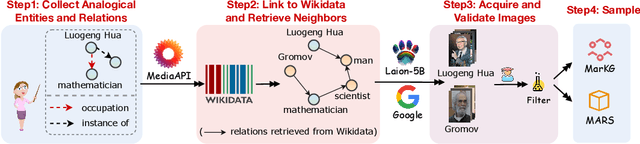
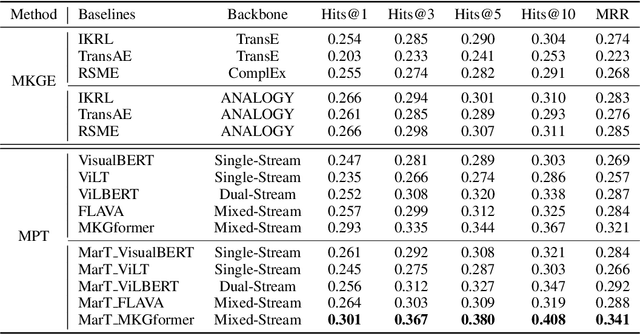
Abstract:Analogical reasoning is fundamental to human cognition and holds an important place in various fields. However, previous studies mainly focus on single-modal analogical reasoning and ignore taking advantage of structure knowledge. Notably, the research in cognitive psychology has demonstrated that information from multimodal sources always brings more powerful cognitive transfer than single modality sources. To this end, we introduce the new task of multimodal analogical reasoning over knowledge graphs, which requires multimodal reasoning ability with the help of background knowledge. Specifically, we construct a Multimodal Analogical Reasoning dataSet (MARS) and a multimodal knowledge graph MarKG. We evaluate with multimodal knowledge graph embedding and pre-trained Transformer baselines, illustrating the potential challenges of the proposed task. We further propose a novel model-agnostic Multimodal analogical reasoning framework with Transformer (MarT) motivated by the structure mapping theory, which can obtain better performance.
Construction and Applications of Open Business Knowledge Graph
Sep 30, 2022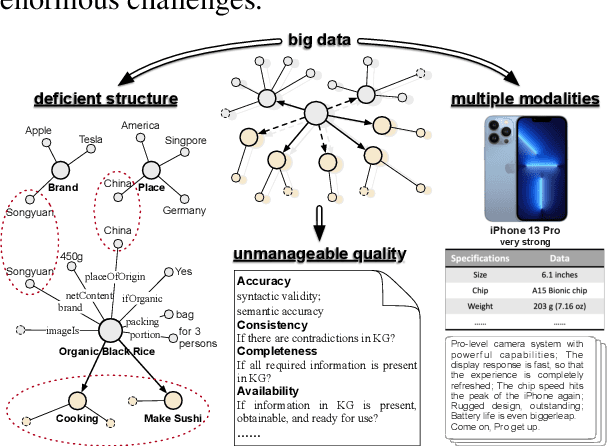
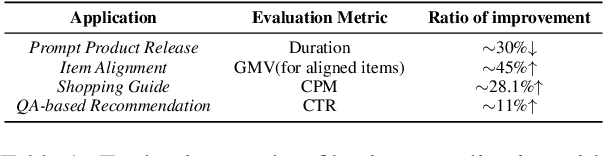

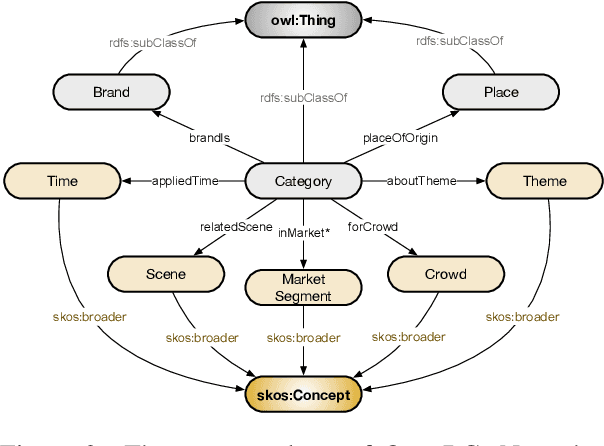
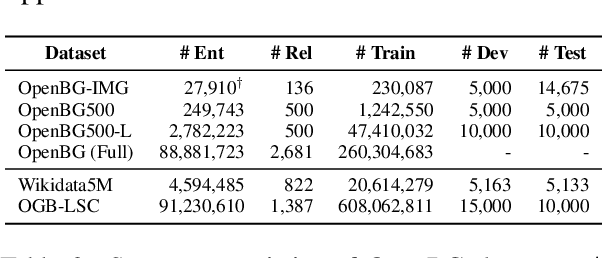
Abstract:Business Knowledge Graph is important to many enterprises today, providing the factual knowledge and structured data that steer many products and make them more intelligent. Despite the welcome outcome, building business KG brings prohibitive issues of deficient structure, multiple modalities and unmanageable quality. In this paper, we advance the practical challenges related to building KG in non-trivial real-world systems. We introduce the process of building an open business knowledge graph (OpenBG) derived from a well-known enterprise. Specifically, we define a core ontology to cover various abstract products and consumption demands, with fine-grained taxonomy and multi-modal facts in deployed applications. OpenBG is ongoing, and the current version contains more than 2.6 billion triples with more than 88 million entities and 2,681 types of relations. We release all the open resources (OpenBG benchmark) derived from it for the community. We also report benchmark results with best learned lessons \url{https://github.com/OpenBGBenchmark/OpenBG}.
Decoupling Knowledge from Memorization: Retrieval-augmented Prompt Learning
May 29, 2022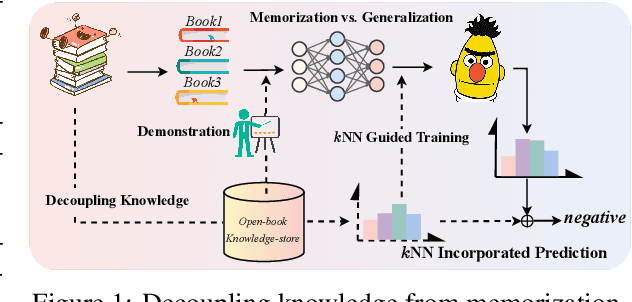
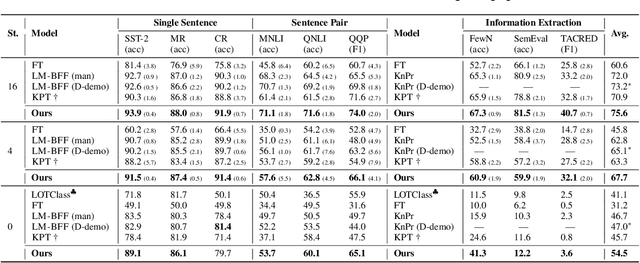
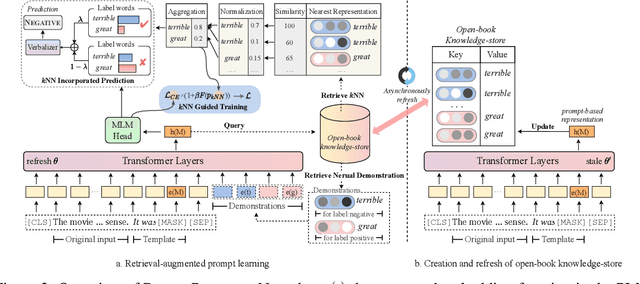
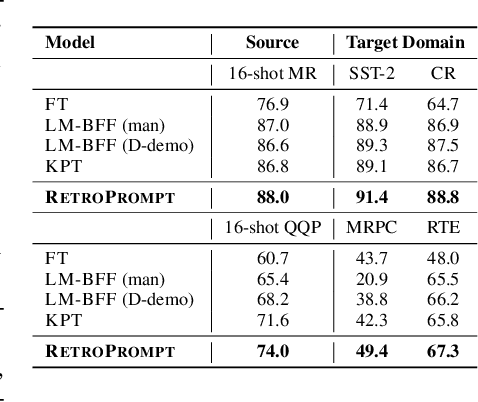
Abstract:Prompt learning approaches have made waves in natural language processing by inducing better few-shot performance while they still follow a parametric-based learning paradigm; the oblivion and rote memorization problems in learning may encounter unstable generalization issues. Specifically, vanilla prompt learning may struggle to utilize atypical instances by rote during fully-supervised training or overfit shallow patterns with low-shot data. To alleviate such limitations, we develop RetroPrompt with the motivation of decoupling knowledge from memorization to help the model strike a balance between generalization and memorization. In contrast with vanilla prompt learning, RetroPrompt constructs an open-book knowledge-store from training instances and implements a retrieval mechanism during the process of input, training and inference, thus equipping the model with the ability to retrieve related contexts from the training corpus as cues for enhancement. Extensive experiments demonstrate that RetroPrompt can obtain better performance in both few-shot and zero-shot settings. Besides, we further illustrate that our proposed RetroPrompt can yield better generalization abilities with new datasets. Detailed analysis of memorization indeed reveals RetroPrompt can reduce the reliance of language models on memorization; thus, improving generalization for downstream tasks.
Good Visual Guidance Makes A Better Extractor: Hierarchical Visual Prefix for Multimodal Entity and Relation Extraction
May 07, 2022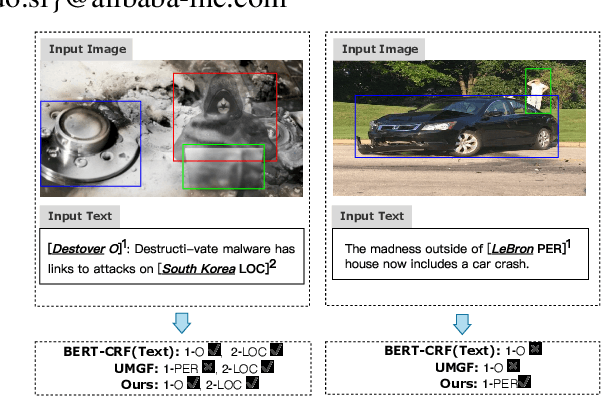
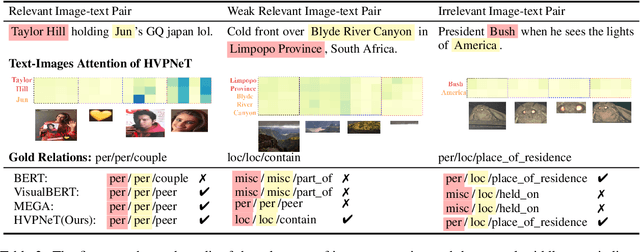
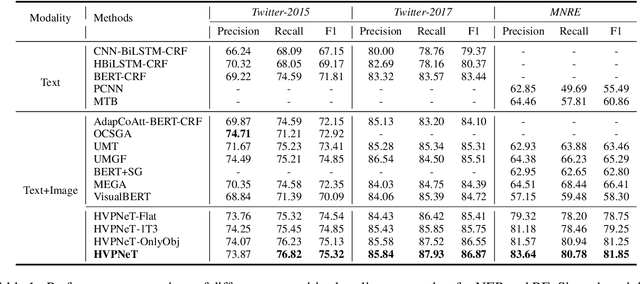
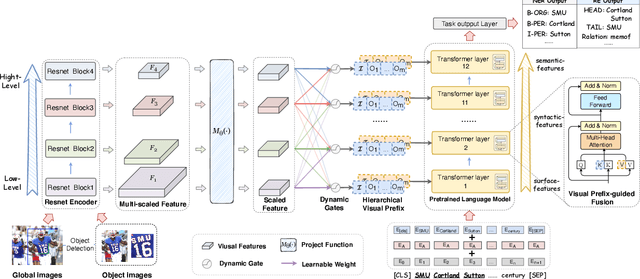

Abstract:Multimodal named entity recognition and relation extraction (MNER and MRE) is a fundamental and crucial branch in information extraction. However, existing approaches for MNER and MRE usually suffer from error sensitivity when irrelevant object images incorporated in texts. To deal with these issues, we propose a novel Hierarchical Visual Prefix fusion NeTwork (HVPNeT) for visual-enhanced entity and relation extraction, aiming to achieve more effective and robust performance. Specifically, we regard visual representation as pluggable visual prefix to guide the textual representation for error insensitive forecasting decision. We further propose a dynamic gated aggregation strategy to achieve hierarchical multi-scaled visual features as visual prefix for fusion. Extensive experiments on three benchmark datasets demonstrate the effectiveness of our method, and achieve state-of-the-art performance. Code is available in https://github.com/zjunlp/HVPNeT.
Hybrid Transformer with Multi-level Fusion for Multimodal Knowledge Graph Completion
May 04, 2022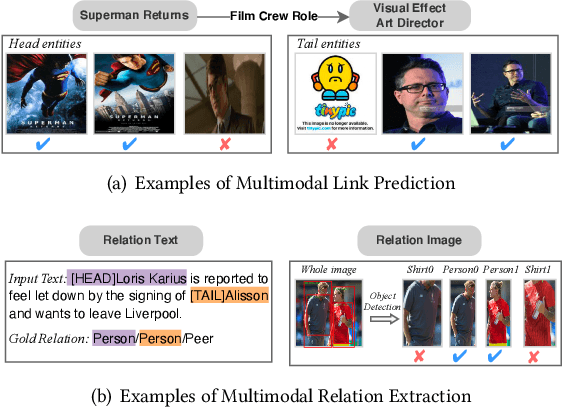
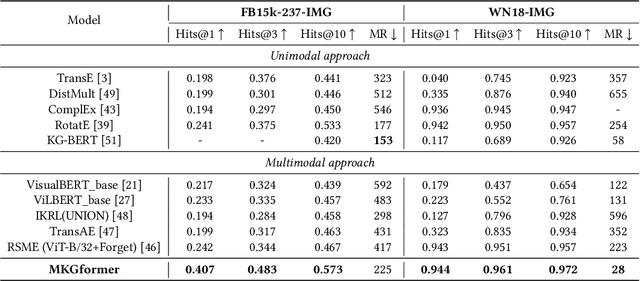
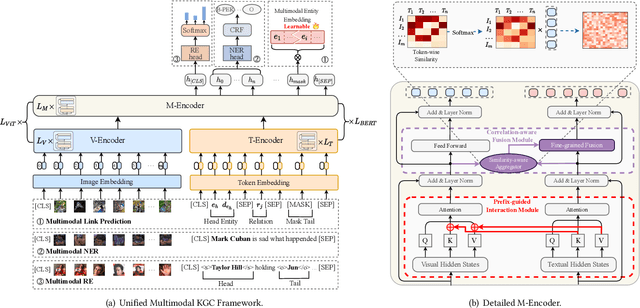
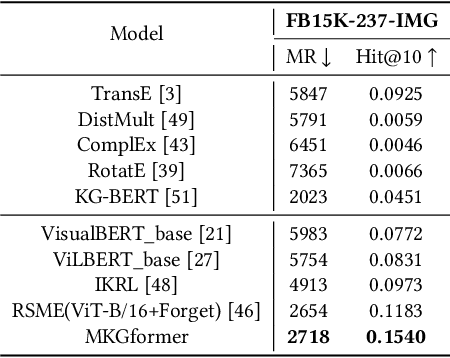
Abstract:Multimodal Knowledge Graphs (MKGs), which organize visual-text factual knowledge, have recently been successfully applied to tasks such as information retrieval, question answering, and recommendation system. Since most MKGs are far from complete, extensive knowledge graph completion studies have been proposed focusing on the multimodal entity, relation extraction and link prediction. However, different tasks and modalities require changes to the model architecture, and not all images/objects are relevant to text input, which hinders the applicability to diverse real-world scenarios. In this paper, we propose a hybrid transformer with multi-level fusion to address those issues. Specifically, we leverage a hybrid transformer architecture with unified input-output for diverse multimodal knowledge graph completion tasks. Moreover, we propose multi-level fusion, which integrates visual and text representation via coarse-grained prefix-guided interaction and fine-grained correlation-aware fusion modules. We conduct extensive experiments to validate that our MKGformer can obtain SOTA performance on four datasets of multimodal link prediction, multimodal RE, and multimodal NER. Code is available in https://github.com/zjunlp/MKGformer.
Knowledge Extraction in Low-Resource Scenarios: Survey and Perspective
Feb 16, 2022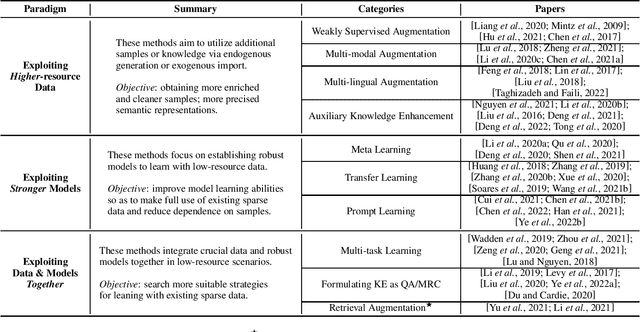
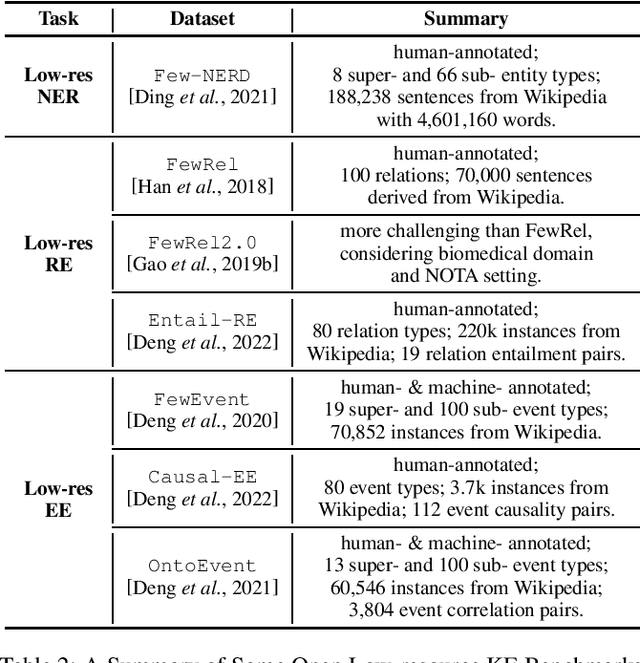
Abstract:Knowledge Extraction (KE) which aims to extract structural information from unstructured texts often suffers from data scarcity and emerging unseen types, i.e., low-resource scenarios. Many neural approaches on low-resource KE have been widely investigated and achieved impressive performance. In this paper, we present a literature review towards KE in low-resource scenarios, and systematically categorize existing works into three paradigms: (1) exploiting higher-resource data, (2) exploiting stronger models, and (3) exploiting data and models together. In addition, we describe promising applications and outline some potential directions for future research. We hope that our survey can help both the academic and industrial community to better understand this field, inspire more ideas and boost broader applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge