Sebastien Ourselin
King's College London
Artificial Intelligence in the Autonomous Navigation of Endovascular Interventions: A Systematic Review
May 06, 2024



Abstract:Purpose: Autonomous navigation of devices in endovascular interventions can decrease operation times, improve decision-making during surgery, and reduce operator radiation exposure while increasing access to treatment. This systematic review explores recent literature to assess the impact, challenges, and opportunities artificial intelligence (AI) has for the autonomous endovascular intervention navigation. Methods: PubMed and IEEEXplore databases were queried. Eligibility criteria included studies investigating the use of AI in enabling the autonomous navigation of catheters/guidewires in endovascular interventions. Following PRISMA, articles were assessed using QUADAS-2. PROSPERO: CRD42023392259. Results: Among 462 studies, fourteen met inclusion criteria. Reinforcement learning (9/14, 64%) and learning from demonstration (7/14, 50%) were used as data-driven models for autonomous navigation. Studies predominantly utilised physical phantoms (10/14, 71%) and in silico (4/14, 29%) models. Experiments within or around the blood vessels of the heart were reported by the majority of studies (10/14, 71%), while simple non-anatomical vessel platforms were used in three studies (3/14, 21%), and the porcine liver venous system in one study. We observed that risk of bias and poor generalisability were present across studies. No procedures were performed on patients in any of the studies reviewed. Studies lacked patient selection criteria, reference standards, and reproducibility, resulting in low clinical evidence levels. Conclusions: AI's potential in autonomous endovascular navigation is promising, but in an experimental proof-of-concept stage, with a technology readiness level of 3. We highlight that reference standards with well-identified performance metrics are crucial to allow for comparisons of data-driven algorithms proposed in the years to come.
* Abstract shortened for arXiv character limit
A self-supervised text-vision framework for automated brain abnormality detection
May 05, 2024



Abstract:Artificial neural networks trained on large, expert-labelled datasets are considered state-of-the-art for a range of medical image recognition tasks. However, categorically labelled datasets are time-consuming to generate and constrain classification to a pre-defined, fixed set of classes. For neuroradiological applications in particular, this represents a barrier to clinical adoption. To address these challenges, we present a self-supervised text-vision framework that learns to detect clinically relevant abnormalities in brain MRI scans by directly leveraging the rich information contained in accompanying free-text neuroradiology reports. Our training approach consisted of two-steps. First, a dedicated neuroradiological language model - NeuroBERT - was trained to generate fixed-dimensional vector representations of neuroradiology reports (N = 50,523) via domain-specific self-supervised learning tasks. Next, convolutional neural networks (one per MRI sequence) learnt to map individual brain scans to their corresponding text vector representations by optimising a mean square error loss. Once trained, our text-vision framework can be used to detect abnormalities in unreported brain MRI examinations by scoring scans against suitable query sentences (e.g., 'there is an acute stroke', 'there is hydrocephalus' etc.), enabling a range of classification-based applications including automated triage. Potentially, our framework could also serve as a clinical decision support tool, not only by suggesting findings to radiologists and detecting errors in provisional reports, but also by retrieving and displaying examples of pathologies from historical examinations that could be relevant to the current case based on textual descriptors.
Label merge-and-split: A graph-colouring approach for memory-efficient brain parcellation
Apr 16, 2024Abstract:Whole brain parcellation requires inferring hundreds of segmentation labels in large image volumes and thus presents significant practical challenges for deep learning approaches. We introduce label merge-and-split, a method that first greatly reduces the effective number of labels required for learning-based whole brain parcellation and then recovers original labels. Using a greedy graph colouring algorithm, our method automatically groups and merges multiple spatially separate labels prior to model training and inference. The merged labels may be semantically unrelated. A deep learning model is trained to predict merged labels. At inference time, original labels are restored using atlas-based influence regions. In our experiments, the proposed approach reduces the number of labels by up to 68% while achieving segmentation accuracy comparable to the baseline method without label merging and splitting. Moreover, model training and inference times as well as GPU memory requirements were reduced significantly. The proposed method can be applied to all semantic segmentation tasks with a large number of spatially separate classes within an atlas-based prior.
Framework to generate perfusion map from CT and CTA images in patients with acute ischemic stroke: A longitudinal and cross-sectional study
Apr 05, 2024



Abstract:Stroke is a leading cause of disability and death. Effective treatment decisions require early and informative vascular imaging. 4D perfusion imaging is ideal but rarely available within the first hour after stroke, whereas plain CT and CTA usually are. Hence, we propose a framework to extract a predicted perfusion map (PPM) derived from CT and CTA images. In all eighteen patients, we found significantly high spatial similarity (with average Spearman's correlation = 0.7893) between our predicted perfusion map (PPM) and the T-max map derived from 4D-CTP. Voxelwise correlations between the PPM and National Institutes of Health Stroke Scale (NIHSS) subscores for L/R hand motor, gaze, and language on a large cohort of 2,110 subjects reliably mapped symptoms to expected infarct locations. Therefore our PPM could serve as an alternative for 4D perfusion imaging, if the latter is unavailable, to investigate blood perfusion in the first hours after hospital admission.
A Robust Ensemble Algorithm for Ischemic Stroke Lesion Segmentation: Generalizability and Clinical Utility Beyond the ISLES Challenge
Apr 03, 2024



Abstract:Diffusion-weighted MRI (DWI) is essential for stroke diagnosis, treatment decisions, and prognosis. However, image and disease variability hinder the development of generalizable AI algorithms with clinical value. We address this gap by presenting a novel ensemble algorithm derived from the 2022 Ischemic Stroke Lesion Segmentation (ISLES) challenge. ISLES'22 provided 400 patient scans with ischemic stroke from various medical centers, facilitating the development of a wide range of cutting-edge segmentation algorithms by the research community. Through collaboration with leading teams, we combined top-performing algorithms into an ensemble model that overcomes the limitations of individual solutions. Our ensemble model achieved superior ischemic lesion detection and segmentation accuracy on our internal test set compared to individual algorithms. This accuracy generalized well across diverse image and disease variables. Furthermore, the model excelled in extracting clinical biomarkers. Notably, in a Turing-like test, neuroradiologists consistently preferred the algorithm's segmentations over manual expert efforts, highlighting increased comprehensiveness and precision. Validation using a real-world external dataset (N=1686) confirmed the model's generalizability. The algorithm's outputs also demonstrated strong correlations with clinical scores (admission NIHSS and 90-day mRS) on par with or exceeding expert-derived results, underlining its clinical relevance. This study offers two key findings. First, we present an ensemble algorithm (https://github.com/Tabrisrei/ISLES22_Ensemble) that detects and segments ischemic stroke lesions on DWI across diverse scenarios on par with expert (neuro)radiologists. Second, we show the potential for biomedical challenge outputs to extend beyond the challenge's initial objectives, demonstrating their real-world clinical applicability.
MatchSeg: Towards Better Segmentation via Reference Image Matching
Mar 23, 2024



Abstract:Recently, automated medical image segmentation methods based on deep learning have achieved great success. However, they heavily rely on large annotated datasets, which are costly and time-consuming to acquire. Few-shot learning aims to overcome the need for annotated data by using a small labeled dataset, known as a support set, to guide predicting labels for new, unlabeled images, known as the query set. Inspired by this paradigm, we introduce MatchSeg, a novel framework that enhances medical image segmentation through strategic reference image matching. We leverage contrastive language-image pre-training (CLIP) to select highly relevant samples when defining the support set. Additionally, we design a joint attention module to strengthen the interaction between support and query features, facilitating a more effective knowledge transfer between support and query sets. We validated our method across four public datasets. Experimental results demonstrate superior segmentation performance and powerful domain generalization ability of MatchSeg against existing methods for domain-specific and cross-domain segmentation tasks. Our code is made available at https://github.com/keeplearning-again/MatchSeg
RetiGen: A Framework for Generalized Retinal Diagnosis Using Multi-View Fundus Images
Mar 22, 2024



Abstract:This study introduces a novel framework for enhancing domain generalization in medical imaging, specifically focusing on utilizing unlabelled multi-view colour fundus photographs. Unlike traditional approaches that rely on single-view imaging data and face challenges in generalizing across diverse clinical settings, our method leverages the rich information in the unlabelled multi-view imaging data to improve model robustness and accuracy. By incorporating a class balancing method, a test-time adaptation technique and a multi-view optimization strategy, we address the critical issue of domain shift that often hampers the performance of machine learning models in real-world applications. Experiments comparing various state-of-the-art domain generalization and test-time optimization methodologies show that our approach consistently outperforms when combined with existing baseline and state-of-the-art methods. We also show our online method improves all existing techniques. Our framework demonstrates improvements in domain generalization capabilities and offers a practical solution for real-world deployment by facilitating online adaptation to new, unseen datasets. Our code is available at https://github.com/zgy600/RetiGen .
DDSB: An Unsupervised and Training-free Method for Phase Detection in Echocardiography
Mar 19, 2024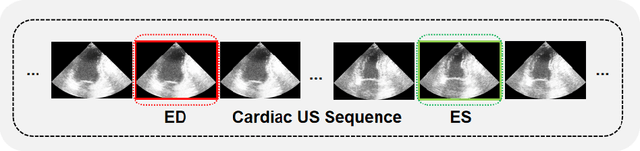
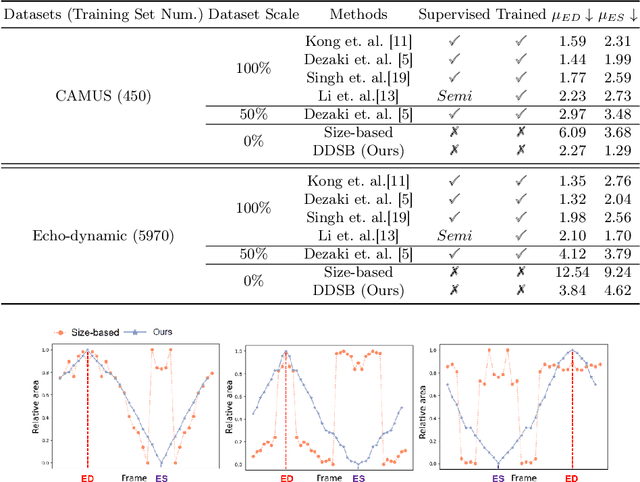
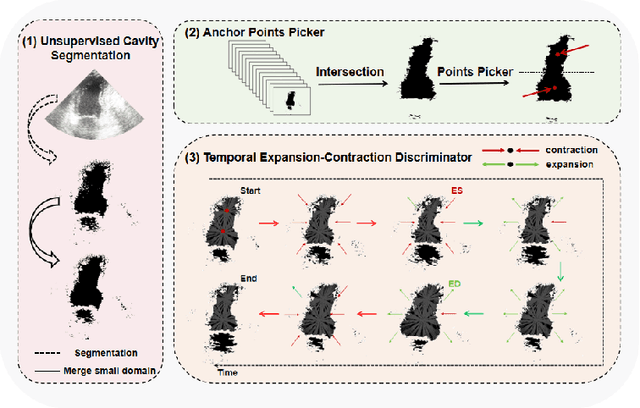
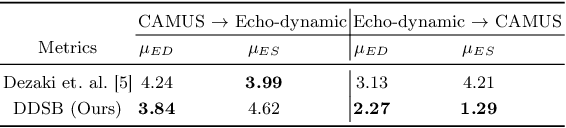
Abstract:Accurate identification of End-Diastolic (ED) and End-Systolic (ES) frames is key for cardiac function assessment through echocardiography. However, traditional methods face several limitations: they require extensive amounts of data, extensive annotations by medical experts, significant training resources, and often lack robustness. Addressing these challenges, we proposed an unsupervised and training-free method, our novel approach leverages unsupervised segmentation to enhance fault tolerance against segmentation inaccuracies. By identifying anchor points and analyzing directional deformation, we effectively reduce dependence on the accuracy of initial segmentation images and enhance fault tolerance, all while improving robustness. Tested on Echo-dynamic and CAMUS datasets, our method achieves comparable accuracy to learning-based models without their associated drawbacks. The code is available at https://github.com/MRUIL/DDSB
DD-VNB: A Depth-based Dual-Loop Framework for Real-time Visually Navigated Bronchoscopy
Mar 15, 2024



Abstract:Real-time 6 DOF localization of bronchoscopes is crucial for enhancing intervention quality. However, current vision-based technologies struggle to balance between generalization to unseen data and computational speed. In this study, we propose a Depth-based Dual-Loop framework for real-time Visually Navigated Bronchoscopy (DD-VNB) that can generalize across patient cases without the need of re-training. The DD-VNB framework integrates two key modules: depth estimation and dual-loop localization. To address the domain gap among patients, we propose a knowledge-embedded depth estimation network that maps endoscope frames to depth, ensuring generalization by eliminating patient-specific textures. The network embeds view synthesis knowledge into a cycle adversarial architecture for scale-constrained monocular depth estimation. For real-time performance, our localization module embeds a fast ego-motion estimation network into the loop of depth registration. The ego-motion inference network estimates the pose change of the bronchoscope in high frequency while depth registration against the pre-operative 3D model provides absolute pose periodically. Specifically, the relative pose changes are fed into the registration process as the initial guess to boost its accuracy and speed. Experiments on phantom and in-vivo data from patients demonstrate the effectiveness of our framework: 1) monocular depth estimation outperforms SOTA, 2) localization achieves an accuracy of Absolute Tracking Error (ATE) of 4.7 $\pm$ 3.17 mm in phantom and 6.49 $\pm$ 3.88 mm in patient data, 3) with a frame-rate approaching video capture speed, 4) without the necessity of case-wise network retraining. The framework's superior speed and accuracy demonstrate its promising clinical potential for real-time bronchoscopic navigation.
Rethinking Low-quality Optical Flow in Unsupervised Surgical Instrument Segmentation
Mar 15, 2024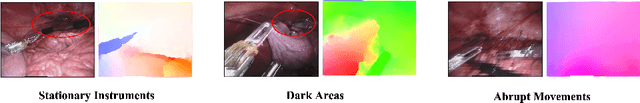
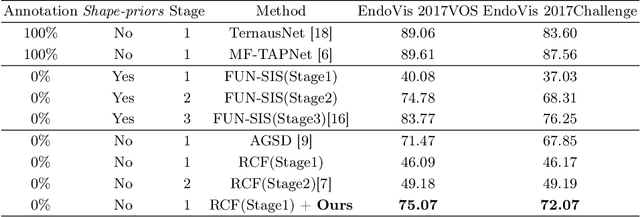
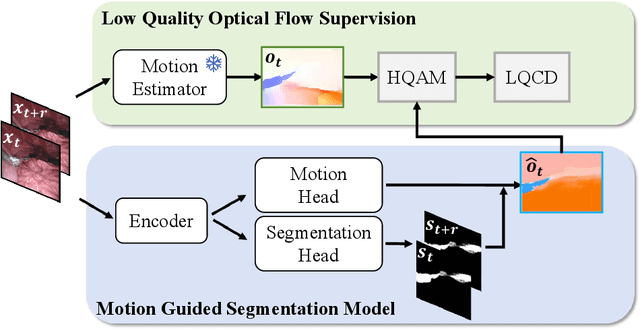
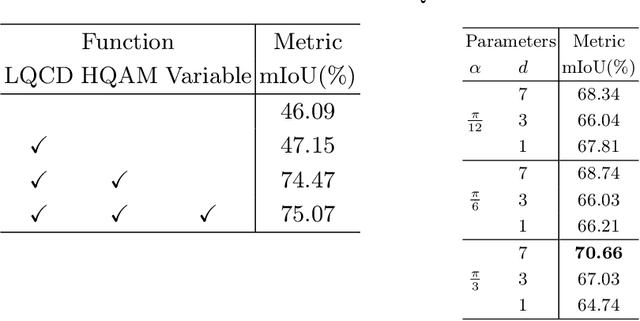
Abstract:Video-based surgical instrument segmentation plays an important role in robot-assisted surgeries. Unlike supervised settings, unsupervised segmentation relies heavily on motion cues, which are challenging to discern due to the typically lower quality of optical flow in surgical footage compared to natural scenes. This presents a considerable burden for the advancement of unsupervised segmentation techniques. In our work, we address the challenge of enhancing model performance despite the inherent limitations of low-quality optical flow. Our methodology employs a three-pronged approach: extracting boundaries directly from the optical flow, selectively discarding frames with inferior flow quality, and employing a fine-tuning process with variable frame rates. We thoroughly evaluate our strategy on the EndoVis2017 VOS dataset and Endovis2017 Challenge dataset, where our model demonstrates promising results, achieving a mean Intersection-over-Union (mIoU) of 0.75 and 0.72, respectively. Our findings suggest that our approach can greatly decrease the need for manual annotations in clinical environments and may facilitate the annotation process for new datasets. The code is available at https://github.com/wpr1018001/Rethinking-Low-quality-Optical-Flow.git
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge