Satoshi Kondo
Surgical tool classification and localization: results and methods from the MICCAI 2022 SurgToolLoc challenge
May 11, 2023



Abstract:The ability to automatically detect and track surgical instruments in endoscopic videos can enable transformational interventions. Assessing surgical performance and efficiency, identifying skilled tool use and choreography, and planning operational and logistical aspects of OR resources are just a few of the applications that could benefit. Unfortunately, obtaining the annotations needed to train machine learning models to identify and localize surgical tools is a difficult task. Annotating bounding boxes frame-by-frame is tedious and time-consuming, yet large amounts of data with a wide variety of surgical tools and surgeries must be captured for robust training. Moreover, ongoing annotator training is needed to stay up to date with surgical instrument innovation. In robotic-assisted surgery, however, potentially informative data like timestamps of instrument installation and removal can be programmatically harvested. The ability to rely on tool installation data alone would significantly reduce the workload to train robust tool-tracking models. With this motivation in mind we invited the surgical data science community to participate in the challenge, SurgToolLoc 2022. The goal was to leverage tool presence data as weak labels for machine learning models trained to detect tools and localize them in video frames with bounding boxes. We present the results of this challenge along with many of the team's efforts. We conclude by discussing these results in the broader context of machine learning and surgical data science. The training data used for this challenge consisting of 24,695 video clips with tool presence labels is also being released publicly and can be accessed at https://console.cloud.google.com/storage/browser/isi-surgtoolloc-2022.
Why is the winner the best?
Mar 30, 2023



Abstract:International benchmarking competitions have become fundamental for the comparative performance assessment of image analysis methods. However, little attention has been given to investigating what can be learnt from these competitions. Do they really generate scientific progress? What are common and successful participation strategies? What makes a solution superior to a competing method? To address this gap in the literature, we performed a multi-center study with all 80 competitions that were conducted in the scope of IEEE ISBI 2021 and MICCAI 2021. Statistical analyses performed based on comprehensive descriptions of the submitted algorithms linked to their rank as well as the underlying participation strategies revealed common characteristics of winning solutions. These typically include the use of multi-task learning (63%) and/or multi-stage pipelines (61%), and a focus on augmentation (100%), image preprocessing (97%), data curation (79%), and postprocessing (66%). The "typical" lead of a winning team is a computer scientist with a doctoral degree, five years of experience in biomedical image analysis, and four years of experience in deep learning. Two core general development strategies stood out for highly-ranked teams: the reflection of the metrics in the method design and the focus on analyzing and handling failure cases. According to the organizers, 43% of the winning algorithms exceeded the state of the art but only 11% completely solved the respective domain problem. The insights of our study could help researchers (1) improve algorithm development strategies when approaching new problems, and (2) focus on open research questions revealed by this work.
CoNIC Challenge: Pushing the Frontiers of Nuclear Detection, Segmentation, Classification and Counting
Mar 14, 2023



Abstract:Nuclear detection, segmentation and morphometric profiling are essential in helping us further understand the relationship between histology and patient outcome. To drive innovation in this area, we setup a community-wide challenge using the largest available dataset of its kind to assess nuclear segmentation and cellular composition. Our challenge, named CoNIC, stimulated the development of reproducible algorithms for cellular recognition with real-time result inspection on public leaderboards. We conducted an extensive post-challenge analysis based on the top-performing models using 1,658 whole-slide images of colon tissue. With around 700 million detected nuclei per model, associated features were used for dysplasia grading and survival analysis, where we demonstrated that the challenge's improvement over the previous state-of-the-art led to significant boosts in downstream performance. Our findings also suggest that eosinophils and neutrophils play an important role in the tumour microevironment. We release challenge models and WSI-level results to foster the development of further methods for biomarker discovery.
Unsupervised Domain Adaptation for MRI Volume Segmentation and Classification Using Image-to-Image Translation
Feb 16, 2023
Abstract:Unsupervised domain adaptation is a type of domain adaptation and exploits labeled data from the source domain and unlabeled data from the target one. In the Cross-Modality Domain Adaptation for Medical Image Segmenta-tion challenge (crossMoDA2022), contrast enhanced T1 MRI volumes for brain are provided as the source domain data, and high-resolution T2 MRI volumes are provided as the target domain data. The crossMoDA2022 challenge contains two tasks, segmentation of vestibular schwannoma (VS) and cochlea, and clas-sification of VS with Koos grade. In this report, we presented our solution for the crossMoDA2022 challenge. We employ an image-to-image translation method for unsupervised domain adaptation and residual U-Net the segmenta-tion task. We use SVM for the classification task. The experimental results show that the mean DSC and ASSD are 0.614 and 2.936 for the segmentation task and MA-MAE is 0.84 for the classification task.
Automated Lesion Segmentation in Whole-Body FDG-PET/CT with Multi-modality Deep Neural Networks
Feb 16, 2023Abstract:Recent progress in automated PET/CT lesion segmentation using deep learning methods has demonstrated the feasibility of this task. However, tumor lesion detection and segmentation in whole-body PET/CT is still a chal-lenging task. To promote research on machine learning-based automated tumor lesion segmentation on whole-body FDG-PET/CT data, Automated Lesion Segmentation in Whole-Body FDG-PET/CT (autoPET) challenge is held, and a large, publicly available training dataset is provided. In this report, we present our solution to the autoPET challenge. We employ multi-modal residual U-Net with deep super vision. The experimental results for five preliminary test cases show that Dice score is 0.79 +/- 0.21.
CholecTriplet2022: Show me a tool and tell me the triplet -- an endoscopic vision challenge for surgical action triplet detection
Feb 13, 2023



Abstract:Formalizing surgical activities as triplets of the used instruments, actions performed, and target anatomies is becoming a gold standard approach for surgical activity modeling. The benefit is that this formalization helps to obtain a more detailed understanding of tool-tissue interaction which can be used to develop better Artificial Intelligence assistance for image-guided surgery. Earlier efforts and the CholecTriplet challenge introduced in 2021 have put together techniques aimed at recognizing these triplets from surgical footage. Estimating also the spatial locations of the triplets would offer a more precise intraoperative context-aware decision support for computer-assisted intervention. This paper presents the CholecTriplet2022 challenge, which extends surgical action triplet modeling from recognition to detection. It includes weakly-supervised bounding box localization of every visible surgical instrument (or tool), as the key actors, and the modeling of each tool-activity in the form of <instrument, verb, target> triplet. The paper describes a baseline method and 10 new deep learning algorithms presented at the challenge to solve the task. It also provides thorough methodological comparisons of the methods, an in-depth analysis of the obtained results, their significance, and useful insights for future research directions and applications in surgery.
AIROGS: Artificial Intelligence for RObust Glaucoma Screening Challenge
Feb 10, 2023



Abstract:The early detection of glaucoma is essential in preventing visual impairment. Artificial intelligence (AI) can be used to analyze color fundus photographs (CFPs) in a cost-effective manner, making glaucoma screening more accessible. While AI models for glaucoma screening from CFPs have shown promising results in laboratory settings, their performance decreases significantly in real-world scenarios due to the presence of out-of-distribution and low-quality images. To address this issue, we propose the Artificial Intelligence for Robust Glaucoma Screening (AIROGS) challenge. This challenge includes a large dataset of around 113,000 images from about 60,000 patients and 500 different screening centers, and encourages the development of algorithms that are robust to ungradable and unexpected input data. We evaluated solutions from 14 teams in this paper, and found that the best teams performed similarly to a set of 20 expert ophthalmologists and optometrists. The highest-scoring team achieved an area under the receiver operating characteristic curve of 0.99 (95% CI: 0.98-0.99) for detecting ungradable images on-the-fly. Additionally, many of the algorithms showed robust performance when tested on three other publicly available datasets. These results demonstrate the feasibility of robust AI-enabled glaucoma screening.
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Abstract:The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
Objective Surgical Skills Assessment and Tool Localization: Results from the MICCAI 2021 SimSurgSkill Challenge
Dec 08, 2022Abstract:Timely and effective feedback within surgical training plays a critical role in developing the skills required to perform safe and efficient surgery. Feedback from expert surgeons, while especially valuable in this regard, is challenging to acquire due to their typically busy schedules, and may be subject to biases. Formal assessment procedures like OSATS and GEARS attempt to provide objective measures of skill, but remain time-consuming. With advances in machine learning there is an opportunity for fast and objective automated feedback on technical skills. The SimSurgSkill 2021 challenge (hosted as a sub-challenge of EndoVis at MICCAI 2021) aimed to promote and foster work in this endeavor. Using virtual reality (VR) surgical tasks, competitors were tasked with localizing instruments and predicting surgical skill. Here we summarize the winning approaches and how they performed. Using this publicly available dataset and results as a springboard, future work may enable more efficient training of surgeons with advances in surgical data science. The dataset can be accessed from https://console.cloud.google.com/storage/browser/isi-simsurgskill-2021.
Source-Free Unsupervised Domain Adaptation with Norm and Shape Constraints for Medical Image Segmentation
Sep 03, 2022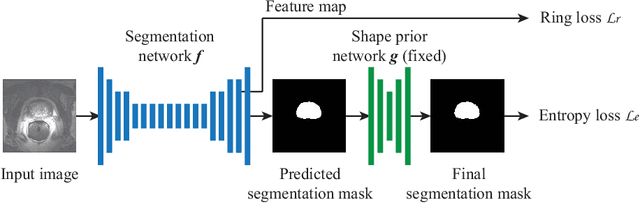
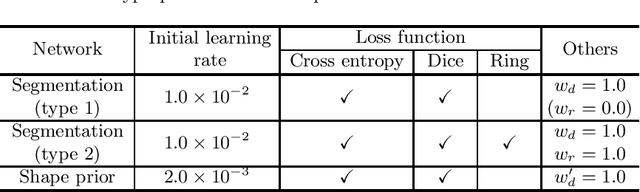

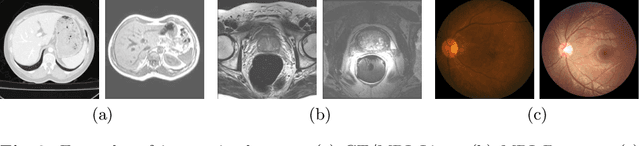
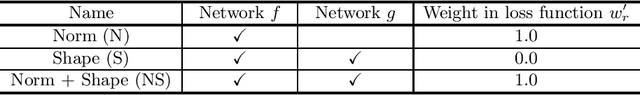
Abstract:Unsupervised domain adaptation (UDA) is one of the key technologies to solve a problem where it is hard to obtain ground truth labels needed for supervised learning. In general, UDA assumes that all samples from source and target domains are available during the training process. However, this is not a realistic assumption under applications where data privacy issues are concerned. To overcome this limitation, UDA without source data, referred to source-free unsupervised domain adaptation (SFUDA) has been recently proposed. Here, we propose a SFUDA method for medical image segmentation. In addition to the entropy minimization method, which is commonly used in UDA, we introduce a loss function for avoiding feature norms in the target domain small and a prior to preserve shape constraints of the target organ. We conduct experiments using datasets including multiple types of source-target domain combinations in order to show the versatility and robustness of our method. We confirm that our method outperforms the state-of-the-art in all datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge