S. Sara Mahdavi
Advancing Conversational Diagnostic AI with Multimodal Reasoning
May 06, 2025



Abstract:Large Language Models (LLMs) have demonstrated great potential for conducting diagnostic conversations but evaluation has been largely limited to language-only interactions, deviating from the real-world requirements of remote care delivery. Instant messaging platforms permit clinicians and patients to upload and discuss multimodal medical artifacts seamlessly in medical consultation, but the ability of LLMs to reason over such data while preserving other attributes of competent diagnostic conversation remains unknown. Here we advance the conversational diagnosis and management performance of the Articulate Medical Intelligence Explorer (AMIE) through a new capability to gather and interpret multimodal data, and reason about this precisely during consultations. Leveraging Gemini 2.0 Flash, our system implements a state-aware dialogue framework, where conversation flow is dynamically controlled by intermediate model outputs reflecting patient states and evolving diagnoses. Follow-up questions are strategically directed by uncertainty in such patient states, leading to a more structured multimodal history-taking process that emulates experienced clinicians. We compared AMIE to primary care physicians (PCPs) in a randomized, blinded, OSCE-style study of chat-based consultations with patient actors. We constructed 105 evaluation scenarios using artifacts like smartphone skin photos, ECGs, and PDFs of clinical documents across diverse conditions and demographics. Our rubric assessed multimodal capabilities and other clinically meaningful axes like history-taking, diagnostic accuracy, management reasoning, communication, and empathy. Specialist evaluation showed AMIE to be superior to PCPs on 7/9 multimodal and 29/32 non-multimodal axes (including diagnostic accuracy). The results show clear progress in multimodal conversational diagnostic AI, but real-world translation needs further research.
Towards Conversational AI for Disease Management
Mar 08, 2025



Abstract:While large language models (LLMs) have shown promise in diagnostic dialogue, their capabilities for effective management reasoning - including disease progression, therapeutic response, and safe medication prescription - remain under-explored. We advance the previously demonstrated diagnostic capabilities of the Articulate Medical Intelligence Explorer (AMIE) through a new LLM-based agentic system optimised for clinical management and dialogue, incorporating reasoning over the evolution of disease and multiple patient visit encounters, response to therapy, and professional competence in medication prescription. To ground its reasoning in authoritative clinical knowledge, AMIE leverages Gemini's long-context capabilities, combining in-context retrieval with structured reasoning to align its output with relevant and up-to-date clinical practice guidelines and drug formularies. In a randomized, blinded virtual Objective Structured Clinical Examination (OSCE) study, AMIE was compared to 21 primary care physicians (PCPs) across 100 multi-visit case scenarios designed to reflect UK NICE Guidance and BMJ Best Practice guidelines. AMIE was non-inferior to PCPs in management reasoning as assessed by specialist physicians and scored better in both preciseness of treatments and investigations, and in its alignment with and grounding of management plans in clinical guidelines. To benchmark medication reasoning, we developed RxQA, a multiple-choice question benchmark derived from two national drug formularies (US, UK) and validated by board-certified pharmacists. While AMIE and PCPs both benefited from the ability to access external drug information, AMIE outperformed PCPs on higher difficulty questions. While further research would be needed before real-world translation, AMIE's strong performance across evaluations marks a significant step towards conversational AI as a tool in disease management.
Sleepless Nights, Sugary Days: Creating Synthetic Users with Health Conditions for Realistic Coaching Agent Interactions
Feb 18, 2025Abstract:We present an end-to-end framework for generating synthetic users for evaluating interactive agents designed to encourage positive behavior changes, such as in health and lifestyle coaching. The synthetic users are grounded in health and lifestyle conditions, specifically sleep and diabetes management in this study, to ensure realistic interactions with the health coaching agent. Synthetic users are created in two stages: first, structured data are generated grounded in real-world health and lifestyle factors in addition to basic demographics and behavioral attributes; second, full profiles of the synthetic users are developed conditioned on the structured data. Interactions between synthetic users and the coaching agent are simulated using generative agent-based models such as Concordia, or directly by prompting a language model. Using two independently-developed agents for sleep and diabetes coaching as case studies, the validity of this framework is demonstrated by analyzing the coaching agent's understanding of the synthetic users' needs and challenges. Finally, through multiple blinded evaluations of user-coach interactions by human experts, we demonstrate that our synthetic users with health and behavioral attributes more accurately portray real human users with the same attributes, compared to generic synthetic users not grounded in such attributes. The proposed framework lays the foundation for efficient development of conversational agents through extensive, realistic, and grounded simulated interactions.
Tx-LLM: A Large Language Model for Therapeutics
Jun 10, 2024



Abstract:Developing therapeutics is a lengthy and expensive process that requires the satisfaction of many different criteria, and AI models capable of expediting the process would be invaluable. However, the majority of current AI approaches address only a narrowly defined set of tasks, often circumscribed within a particular domain. To bridge this gap, we introduce Tx-LLM, a generalist large language model (LLM) fine-tuned from PaLM-2 which encodes knowledge about diverse therapeutic modalities. Tx-LLM is trained using a collection of 709 datasets that target 66 tasks spanning various stages of the drug discovery pipeline. Using a single set of weights, Tx-LLM simultaneously processes a wide variety of chemical or biological entities(small molecules, proteins, nucleic acids, cell lines, diseases) interleaved with free-text, allowing it to predict a broad range of associated properties, achieving competitive with state-of-the-art (SOTA) performance on 43 out of 66 tasks and exceeding SOTA on 22. Among these, Tx-LLM is particularly powerful and exceeds best-in-class performance on average for tasks combining molecular SMILES representations with text such as cell line names or disease names, likely due to context learned during pretraining. We observe evidence of positive transfer between tasks with diverse drug types (e.g.,tasks involving small molecules and tasks involving proteins), and we study the impact of model size, domain finetuning, and prompting strategies on performance. We believe Tx-LLM represents an important step towards LLMs encoding biochemical knowledge and could have a future role as an end-to-end tool across the drug discovery development pipeline.
Advancing Multimodal Medical Capabilities of Gemini
May 06, 2024



Abstract:Many clinical tasks require an understanding of specialized data, such as medical images and genomics, which is not typically found in general-purpose large multimodal models. Building upon Gemini's multimodal models, we develop several models within the new Med-Gemini family that inherit core capabilities of Gemini and are optimized for medical use via fine-tuning with 2D and 3D radiology, histopathology, ophthalmology, dermatology and genomic data. Med-Gemini-2D sets a new standard for AI-based chest X-ray (CXR) report generation based on expert evaluation, exceeding previous best results across two separate datasets by an absolute margin of 1% and 12%, where 57% and 96% of AI reports on normal cases, and 43% and 65% on abnormal cases, are evaluated as "equivalent or better" than the original radiologists' reports. We demonstrate the first ever large multimodal model-based report generation for 3D computed tomography (CT) volumes using Med-Gemini-3D, with 53% of AI reports considered clinically acceptable, although additional research is needed to meet expert radiologist reporting quality. Beyond report generation, Med-Gemini-2D surpasses the previous best performance in CXR visual question answering (VQA) and performs well in CXR classification and radiology VQA, exceeding SoTA or baselines on 17 of 20 tasks. In histopathology, ophthalmology, and dermatology image classification, Med-Gemini-2D surpasses baselines across 18 out of 20 tasks and approaches task-specific model performance. Beyond imaging, Med-Gemini-Polygenic outperforms the standard linear polygenic risk score-based approach for disease risk prediction and generalizes to genetically correlated diseases for which it has never been trained. Although further development and evaluation are necessary in the safety-critical medical domain, our results highlight the potential of Med-Gemini across a wide range of medical tasks.
Capabilities of Gemini Models in Medicine
May 01, 2024



Abstract:Excellence in a wide variety of medical applications poses considerable challenges for AI, requiring advanced reasoning, access to up-to-date medical knowledge and understanding of complex multimodal data. Gemini models, with strong general capabilities in multimodal and long-context reasoning, offer exciting possibilities in medicine. Building on these core strengths of Gemini, we introduce Med-Gemini, a family of highly capable multimodal models that are specialized in medicine with the ability to seamlessly use web search, and that can be efficiently tailored to novel modalities using custom encoders. We evaluate Med-Gemini on 14 medical benchmarks, establishing new state-of-the-art (SoTA) performance on 10 of them, and surpass the GPT-4 model family on every benchmark where a direct comparison is viable, often by a wide margin. On the popular MedQA (USMLE) benchmark, our best-performing Med-Gemini model achieves SoTA performance of 91.1% accuracy, using a novel uncertainty-guided search strategy. On 7 multimodal benchmarks including NEJM Image Challenges and MMMU (health & medicine), Med-Gemini improves over GPT-4V by an average relative margin of 44.5%. We demonstrate the effectiveness of Med-Gemini's long-context capabilities through SoTA performance on a needle-in-a-haystack retrieval task from long de-identified health records and medical video question answering, surpassing prior bespoke methods using only in-context learning. Finally, Med-Gemini's performance suggests real-world utility by surpassing human experts on tasks such as medical text summarization, alongside demonstrations of promising potential for multimodal medical dialogue, medical research and education. Taken together, our results offer compelling evidence for Med-Gemini's potential, although further rigorous evaluation will be crucial before real-world deployment in this safety-critical domain.
Towards Expert-Level Medical Question Answering with Large Language Models
May 16, 2023



Abstract:Recent artificial intelligence (AI) systems have reached milestones in "grand challenges" ranging from Go to protein-folding. The capability to retrieve medical knowledge, reason over it, and answer medical questions comparably to physicians has long been viewed as one such grand challenge. Large language models (LLMs) have catalyzed significant progress in medical question answering; Med-PaLM was the first model to exceed a "passing" score in US Medical Licensing Examination (USMLE) style questions with a score of 67.2% on the MedQA dataset. However, this and other prior work suggested significant room for improvement, especially when models' answers were compared to clinicians' answers. Here we present Med-PaLM 2, which bridges these gaps by leveraging a combination of base LLM improvements (PaLM 2), medical domain finetuning, and prompting strategies including a novel ensemble refinement approach. Med-PaLM 2 scored up to 86.5% on the MedQA dataset, improving upon Med-PaLM by over 19% and setting a new state-of-the-art. We also observed performance approaching or exceeding state-of-the-art across MedMCQA, PubMedQA, and MMLU clinical topics datasets. We performed detailed human evaluations on long-form questions along multiple axes relevant to clinical applications. In pairwise comparative ranking of 1066 consumer medical questions, physicians preferred Med-PaLM 2 answers to those produced by physicians on eight of nine axes pertaining to clinical utility (p < 0.001). We also observed significant improvements compared to Med-PaLM on every evaluation axis (p < 0.001) on newly introduced datasets of 240 long-form "adversarial" questions to probe LLM limitations. While further studies are necessary to validate the efficacy of these models in real-world settings, these results highlight rapid progress towards physician-level performance in medical question answering.
Large Language Models Encode Clinical Knowledge
Dec 26, 2022Abstract:Large language models (LLMs) have demonstrated impressive capabilities in natural language understanding and generation, but the quality bar for medical and clinical applications is high. Today, attempts to assess models' clinical knowledge typically rely on automated evaluations on limited benchmarks. There is no standard to evaluate model predictions and reasoning across a breadth of tasks. To address this, we present MultiMedQA, a benchmark combining six existing open question answering datasets spanning professional medical exams, research, and consumer queries; and HealthSearchQA, a new free-response dataset of medical questions searched online. We propose a framework for human evaluation of model answers along multiple axes including factuality, precision, possible harm, and bias. In addition, we evaluate PaLM (a 540-billion parameter LLM) and its instruction-tuned variant, Flan-PaLM, on MultiMedQA. Using a combination of prompting strategies, Flan-PaLM achieves state-of-the-art accuracy on every MultiMedQA multiple-choice dataset (MedQA, MedMCQA, PubMedQA, MMLU clinical topics), including 67.6% accuracy on MedQA (US Medical License Exam questions), surpassing prior state-of-the-art by over 17%. However, human evaluation reveals key gaps in Flan-PaLM responses. To resolve this we introduce instruction prompt tuning, a parameter-efficient approach for aligning LLMs to new domains using a few exemplars. The resulting model, Med-PaLM, performs encouragingly, but remains inferior to clinicians. We show that comprehension, recall of knowledge, and medical reasoning improve with model scale and instruction prompt tuning, suggesting the potential utility of LLMs in medicine. Our human evaluations reveal important limitations of today's models, reinforcing the importance of both evaluation frameworks and method development in creating safe, helpful LLM models for clinical applications.
Photorealistic Text-to-Image Diffusion Models with Deep Language Understanding
May 23, 2022
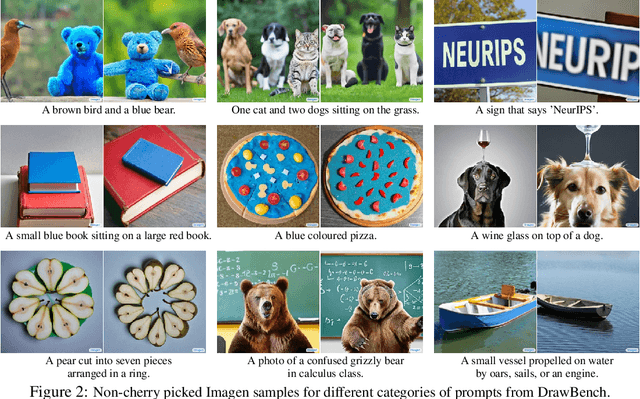

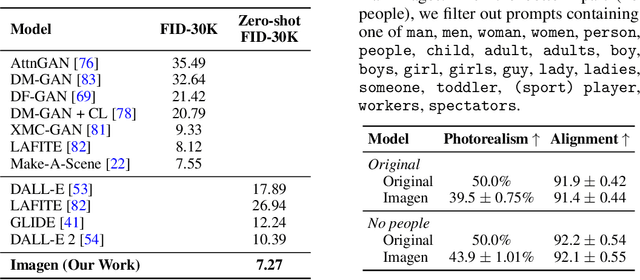
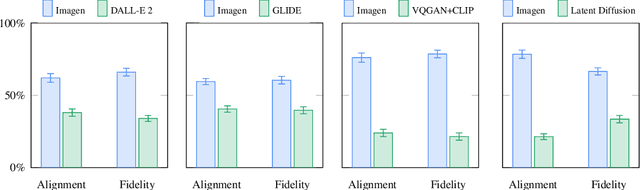
Abstract:We present Imagen, a text-to-image diffusion model with an unprecedented degree of photorealism and a deep level of language understanding. Imagen builds on the power of large transformer language models in understanding text and hinges on the strength of diffusion models in high-fidelity image generation. Our key discovery is that generic large language models (e.g. T5), pretrained on text-only corpora, are surprisingly effective at encoding text for image synthesis: increasing the size of the language model in Imagen boosts both sample fidelity and image-text alignment much more than increasing the size of the image diffusion model. Imagen achieves a new state-of-the-art FID score of 7.27 on the COCO dataset, without ever training on COCO, and human raters find Imagen samples to be on par with the COCO data itself in image-text alignment. To assess text-to-image models in greater depth, we introduce DrawBench, a comprehensive and challenging benchmark for text-to-image models. With DrawBench, we compare Imagen with recent methods including VQ-GAN+CLIP, Latent Diffusion Models, and DALL-E 2, and find that human raters prefer Imagen over other models in side-by-side comparisons, both in terms of sample quality and image-text alignment. See https://imagen.research.google/ for an overview of the results.
Robust and Efficient Medical Imaging with Self-Supervision
May 19, 2022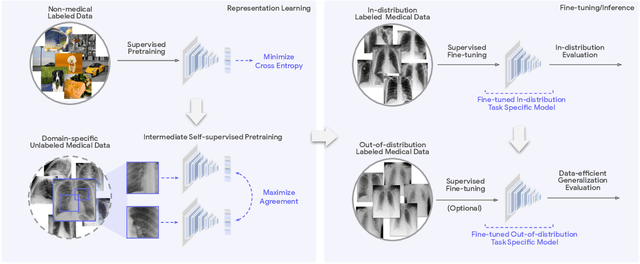
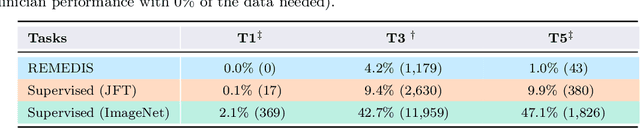
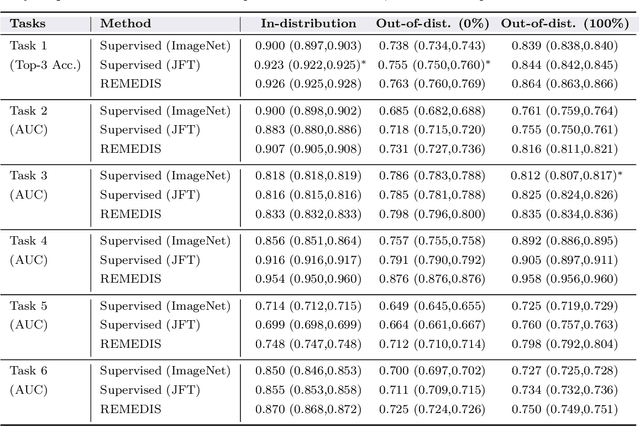
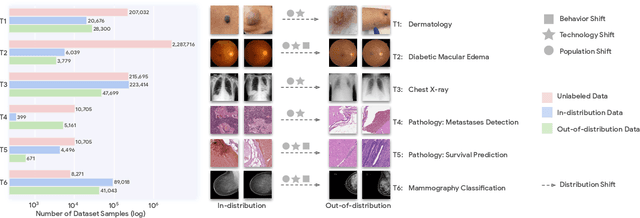

Abstract:Recent progress in Medical Artificial Intelligence (AI) has delivered systems that can reach clinical expert level performance. However, such systems tend to demonstrate sub-optimal "out-of-distribution" performance when evaluated in clinical settings different from the training environment. A common mitigation strategy is to develop separate systems for each clinical setting using site-specific data [1]. However, this quickly becomes impractical as medical data is time-consuming to acquire and expensive to annotate [2]. Thus, the problem of "data-efficient generalization" presents an ongoing difficulty for Medical AI development. Although progress in representation learning shows promise, their benefits have not been rigorously studied, specifically for out-of-distribution settings. To meet these challenges, we present REMEDIS, a unified representation learning strategy to improve robustness and data-efficiency of medical imaging AI. REMEDIS uses a generic combination of large-scale supervised transfer learning with self-supervised learning and requires little task-specific customization. We study a diverse range of medical imaging tasks and simulate three realistic application scenarios using retrospective data. REMEDIS exhibits significantly improved in-distribution performance with up to 11.5% relative improvement in diagnostic accuracy over a strong supervised baseline. More importantly, our strategy leads to strong data-efficient generalization of medical imaging AI, matching strong supervised baselines using between 1% to 33% of retraining data across tasks. These results suggest that REMEDIS can significantly accelerate the life-cycle of medical imaging AI development thereby presenting an important step forward for medical imaging AI to deliver broad impact.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge