Qi Yang
MedSAM-Agent: Empowering Interactive Medical Image Segmentation with Multi-turn Agentic Reinforcement Learning
Feb 03, 2026Abstract:Medical image segmentation is evolving from task-specific models toward generalizable frameworks. Recent research leverages Multi-modal Large Language Models (MLLMs) as autonomous agents, employing reinforcement learning with verifiable reward (RLVR) to orchestrate specialized tools like the Segment Anything Model (SAM). However, these approaches often rely on single-turn, rigid interaction strategies and lack process-level supervision during training, which hinders their ability to fully exploit the dynamic potential of interactive tools and leads to redundant actions. To bridge this gap, we propose MedSAM-Agent, a framework that reformulates interactive segmentation as a multi-step autonomous decision-making process. First, we introduce a hybrid prompting strategy for expert-curated trajectory generation, enabling the model to internalize human-like decision heuristics and adaptive refinement strategies. Furthermore, we develop a two-stage training pipeline that integrates multi-turn, end-to-end outcome verification with a clinical-fidelity process reward design to promote interaction parsimony and decision efficiency. Extensive experiments across 6 medical modalities and 21 datasets demonstrate that MedSAM-Agent achieves state-of-the-art performance, effectively unifying autonomous medical reasoning with robust, iterative optimization. Code is available \href{https://github.com/CUHK-AIM-Group/MedSAM-Agent}{here}.
Molecular Representations in Implicit Functional Space via Hyper-Networks
Jan 29, 2026Abstract:Molecular representations fundamentally shape how machine learning systems reason about molecular structure and physical properties. Most existing approaches adopt a discrete pipeline: molecules are encoded as sequences, graphs, or point clouds, mapped to fixed-dimensional embeddings, and then used for task-specific prediction. This paradigm treats molecules as discrete objects, despite their intrinsically continuous and field-like physical nature. We argue that molecular learning can instead be formulated as learning in function space. Specifically, we model each molecule as a continuous function over three-dimensional (3D) space and treat this molecular field as the primary object of representation. From this perspective, conventional molecular representations arise as particular sampling schemes of an underlying continuous object. We instantiate this formulation with MolField, a hyper-network-based framework that learns distributions over molecular fields. To ensure physical consistency, these functions are defined over canonicalized coordinates, yielding invariance to global SE(3) transformations. To enable learning directly over functions, we introduce a structured weight tokenization and train a sequence-based hyper-network to model a shared prior over molecular fields. We evaluate MolField on molecular dynamics and property prediction. Our results show that treating molecules as continuous functions fundamentally changes how molecular representations generalize across tasks and yields downstream behavior that is stable to how molecules are discretized or queried.
Building from Scratch: A Multi-Agent Framework with Human-in-the-Loop for Multilingual Legal Terminology Mapping
Dec 15, 2025Abstract:Accurately mapping legal terminology across languages remains a significant challenge, especially for language pairs like Chinese and Japanese, which share a large number of homographs with different meanings. Existing resources and standardized tools for these languages are limited. To address this, we propose a human-AI collaborative approach for building a multilingual legal terminology database, based on a multi-agent framework. This approach integrates advanced large language models and legal domain experts throughout the entire process-from raw document preprocessing, article-level alignment, to terminology extraction, mapping, and quality assurance. Unlike a single automated pipeline, our approach places greater emphasis on how human experts participate in this multi-agent system. Humans and AI agents take on different roles: AI agents handle specific, repetitive tasks, such as OCR, text segmentation, semantic alignment, and initial terminology extraction, while human experts provide crucial oversight, review, and supervise the outputs with contextual knowledge and legal judgment. We tested the effectiveness of this framework using a trilingual parallel corpus comprising 35 key Chinese statutes, along with their English and Japanese translations. The experimental results show that this human-in-the-loop, multi-agent workflow not only improves the precision and consistency of multilingual legal terminology mapping but also offers greater scalability compared to traditional manual methods.
Lightweight 3D Gaussian Splatting Compression via Video Codec
Dec 12, 2025Abstract:Current video-based GS compression methods rely on using Parallel Linear Assignment Sorting (PLAS) to convert 3D GS into smooth 2D maps, which are computationally expensive and time-consuming, limiting the application of GS on lightweight devices. In this paper, we propose a Lightweight 3D Gaussian Splatting (GS) Compression method based on Video codec (LGSCV). First, a two-stage Morton scan is proposed to generate blockwise 2D maps that are friendly for canonical video codecs in which the coding units (CU) are square blocks. A 3D Morton scan is used to permute GS primitives, followed by a 2D Morton scan to map the ordered GS primitives to 2D maps in a blockwise style. However, although the blockwise 2D maps report close performance to the PLAS map in high-bitrate regions, they show a quality collapse at medium-to-low bitrates. Therefore, a principal component analysis (PCA) is used to reduce the dimensionality of spherical harmonics (SH), and a MiniPLAS, which is flexible and fast, is designed to permute the primitives within certain block sizes. Incorporating SH PCA and MiniPLAS leads to a significant gain in rate-distortion (RD) performance, especially at medium and low bitrates. MiniPLAS can also guide the setting of the codec CU size configuration and significantly reduce encoding time. Experimental results on the MPEG dataset demonstrate that the proposed LGSCV achieves over 20% RD gain compared with state-of-the-art methods, while reducing 2D map generation time to approximately 1 second and cutting encoding time by 50%. The code is available at https://github.com/Qi-Yangsjtu/LGSCV .
J-SGFT: Joint Spatial and Graph Fourier Domain Learning for Point Cloud Attribute Deblocking
Nov 07, 2025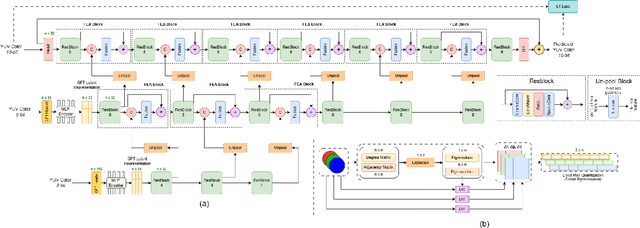
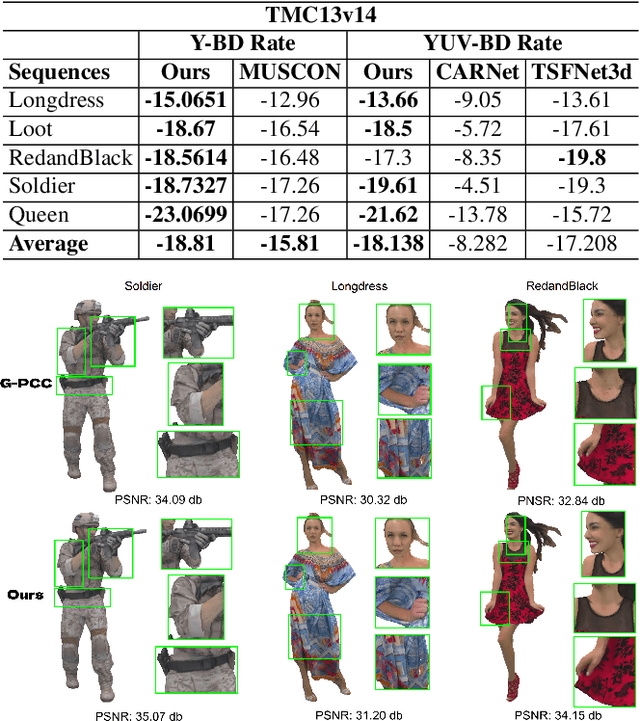
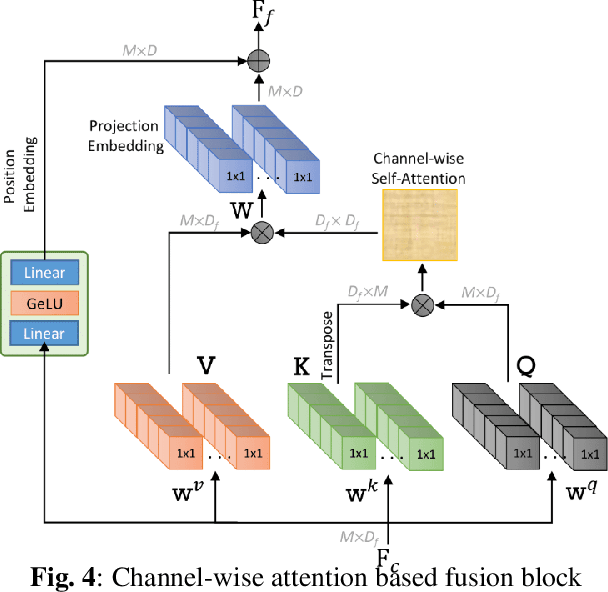
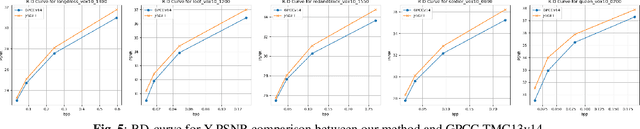

Abstract:Point clouds (PC) are essential for AR/VR and autonomous driving but challenge compression schemes with their size, irregular sampling, and sparsity. MPEG's Geometry-based Point Cloud Compression (GPCC) methods successfully reduce bitrate; however, they introduce significant blocky artifacts in the reconstructed point cloud. We introduce a novel multi-scale postprocessing framework that fuses graph-Fourier latent attribute representations with sparse convolutions and channel-wise attention to efficiently deblock reconstructed point clouds. Against the GPCC TMC13v14 baseline, our approach achieves BD-rate reduction of 18.81\% in the Y channel and 18.14\% in the joint YUV on the 8iVFBv2 dataset, delivering markedly improved visual fidelity with minimal overhead.
TVMC: Time-Varying Mesh Compression via Multi-Stage Anchor Mesh Generation
Oct 26, 2025Abstract:Time-varying meshes, characterized by dynamic connectivity and varying vertex counts, hold significant promise for applications such as augmented reality. However, their practical utilization remains challenging due to the substantial data volume required for high-fidelity representation. While various compression methods attempt to leverage temporal redundancy between consecutive mesh frames, most struggle with topological inconsistency and motion-induced artifacts. To address these issues, we propose Time-Varying Mesh Compression (TVMC), a novel framework built on multi-stage coarse-to-fine anchor mesh generation for inter-frame prediction. Specifically, the anchor mesh is progressively constructed in three stages: initial, coarse, and fine. The initial anchor mesh is obtained through fast topology alignment to exploit temporal coherence. A Kalman filter-based motion estimation module then generates a coarse anchor mesh by accurately compensating inter-frame motions. Subsequently, a Quadric Error Metric-based refinement step optimizes vertex positions to form a fine anchor mesh with improved geometric fidelity. Based on the refined anchor mesh, the inter-frame motions relative to the reference base mesh are encoded, while the residual displacements between the subdivided fine anchor mesh and the input mesh are adaptively quantized and compressed. This hierarchical strategy preserves consistent connectivity and high-quality surface approximation, while achieving an efficient and compact representation of dynamic geometry. Extensive experiments on standard MPEG dynamic mesh sequences demonstrate that TVMC achieves state-of-the-art compression performance. Compared to the latest V-DMC standard, it delivers a significant BD-rate gain of 10.2% ~ 16.9%, while preserving high reconstruction quality. The code is available at https://github.com/H-Huang774/TVMC.
R-4B: Incentivizing General-Purpose Auto-Thinking Capability in MLLMs via Bi-Mode Annealing and Reinforce Learning
Aug 28, 2025



Abstract:Multimodal Large Language Models (MLLMs) equipped with step-by-step thinking capabilities have demonstrated remarkable performance on complex reasoning problems. However, this thinking process is redundant for simple problems solvable without complex reasoning. To address this inefficiency, we propose R-4B, an auto-thinking MLLM, which can adaptively decide when to think based on problem complexity. The central idea of R-4B is to empower the model with both thinking and non-thinking capabilities using bi-mode annealing, and apply Bi-mode Policy Optimization~(BPO) to improve the model's accuracy in determining whether to activate the thinking process. Specifically, we first train the model on a carefully curated dataset spanning various topics, which contains samples from both thinking and non-thinking modes. Then it undergoes a second phase of training under an improved GRPO framework, where the policy model is forced to generate responses from both modes for each input query. Experimental results show that R-4B achieves state-of-the-art performance across 25 challenging benchmarks. It outperforms Qwen2.5-VL-7B in most tasks and achieves performance comparable to larger models such as Kimi-VL-A3B-Thinking-2506 (16B) on reasoning-intensive benchmarks with lower computational cost.
3DGS-VBench: A Comprehensive Video Quality Evaluation Benchmark for 3DGS Compression
Aug 09, 2025Abstract:3D Gaussian Splatting (3DGS) enables real-time novel view synthesis with high visual fidelity, but its substantial storage requirements hinder practical deployment, prompting state-of-the-art (SOTA) 3DGS methods to incorporate compression modules. However, these 3DGS generative compression techniques introduce unique distortions lacking systematic quality assessment research. To this end, we establish 3DGS-VBench, a large-scale Video Quality Assessment (VQA) Dataset and Benchmark with 660 compressed 3DGS models and video sequences generated from 11 scenes across 6 SOTA 3DGS compression algorithms with systematically designed parameter levels. With annotations from 50 participants, we obtained MOS scores with outlier removal and validated dataset reliability. We benchmark 6 3DGS compression algorithms on storage efficiency and visual quality, and evaluate 15 quality assessment metrics across multiple paradigms. Our work enables specialized VQA model training for 3DGS, serving as a catalyst for compression and quality assessment research. The dataset is available at https://github.com/YukeXing/3DGS-VBench.
Rasterizing Wireless Radiance Field via Deformable 2D Gaussian Splatting
Jun 15, 2025Abstract:Modeling the wireless radiance field (WRF) is fundamental to modern communication systems, enabling key tasks such as localization, sensing, and channel estimation. Traditional approaches, which rely on empirical formulas or physical simulations, often suffer from limited accuracy or require strong scene priors. Recent neural radiance field (NeRF-based) methods improve reconstruction fidelity through differentiable volumetric rendering, but their reliance on computationally expensive multilayer perceptron (MLP) queries hinders real-time deployment. To overcome these challenges, we introduce Gaussian splatting (GS) to the wireless domain, leveraging its efficiency in modeling optical radiance fields to enable compact and accurate WRF reconstruction. Specifically, we propose SwiftWRF, a deformable 2D Gaussian splatting framework that synthesizes WRF spectra at arbitrary positions under single-sided transceiver mobility. SwiftWRF employs CUDA-accelerated rasterization to render spectra at over 100000 fps and uses a lightweight MLP to model the deformation of 2D Gaussians, effectively capturing mobility-induced WRF variations. In addition to novel spectrum synthesis, the efficacy of SwiftWRF is further underscored in its applications in angle-of-arrival (AoA) and received signal strength indicator (RSSI) prediction. Experiments conducted on both real-world and synthetic indoor scenes demonstrate that SwiftWRF can reconstruct WRF spectra up to 500x faster than existing state-of-the-art methods, while significantly enhancing its signal quality. Code and datasets will be released.
Re-ranking Reasoning Context with Tree Search Makes Large Vision-Language Models Stronger
Jun 09, 2025Abstract:Recent advancements in Large Vision Language Models (LVLMs) have significantly improved performance in Visual Question Answering (VQA) tasks through multimodal Retrieval-Augmented Generation (RAG). However, existing methods still face challenges, such as the scarcity of knowledge with reasoning examples and erratic responses from retrieved knowledge. To address these issues, in this study, we propose a multimodal RAG framework, termed RCTS, which enhances LVLMs by constructing a Reasoning Context-enriched knowledge base and a Tree Search re-ranking method. Specifically, we introduce a self-consistent evaluation mechanism to enrich the knowledge base with intrinsic reasoning patterns. We further propose a Monte Carlo Tree Search with Heuristic Rewards (MCTS-HR) to prioritize the most relevant examples. This ensures that LVLMs can leverage high-quality contextual reasoning for better and more consistent responses. Extensive experiments demonstrate that our framework achieves state-of-the-art performance on multiple VQA datasets, significantly outperforming In-Context Learning (ICL) and Vanilla-RAG methods. It highlights the effectiveness of our knowledge base and re-ranking method in improving LVLMs. Our code is available at https://github.com/yannqi/RCTS-RAG.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge