Nassir Navab
Computer Aided Medical Procedures, Technische Universit Munchen, Germany, Johns Hopkins University, Baltimore MD, USA
OperA: Attention-Regularized Transformers for Surgical Phase Recognition
Mar 05, 2021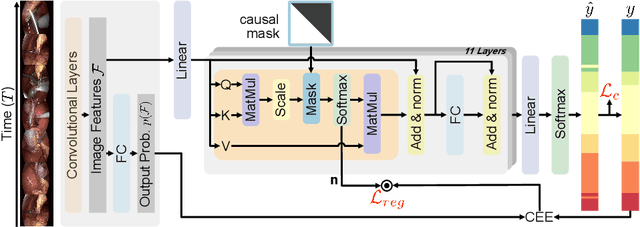
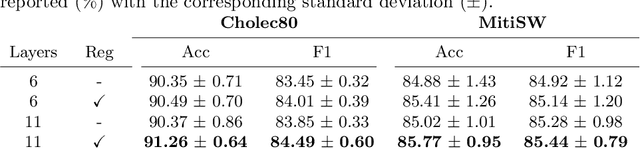
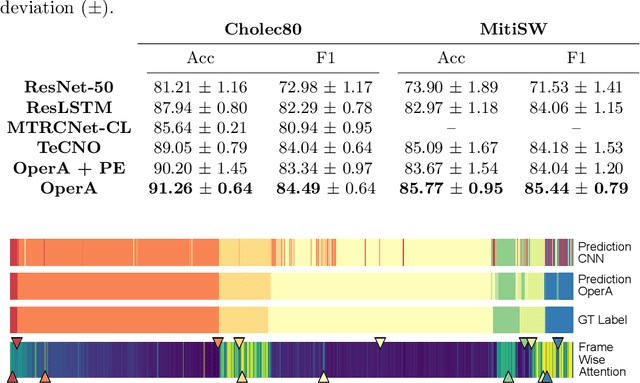
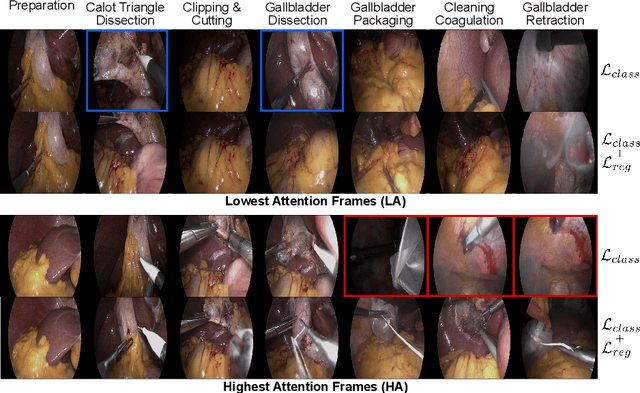
Abstract:In this paper we introduce OperA, a transformer-based model that accurately predicts surgical phases from long video sequences. A novel attention regularization loss encourages the model to focus on high-quality frames during training. Moreover, the attention weights are utilized to identify characteristic high attention frames for each surgical phase, which could further be used for surgery summarization. OperA is thoroughly evaluated on two datasets of laparoscopic cholecystectomy videos, outperforming various state-of-the-art temporal refinement approaches.
RA-GCN: Graph Convolutional Network for Disease Prediction Problems with Imbalanced Data
Feb 27, 2021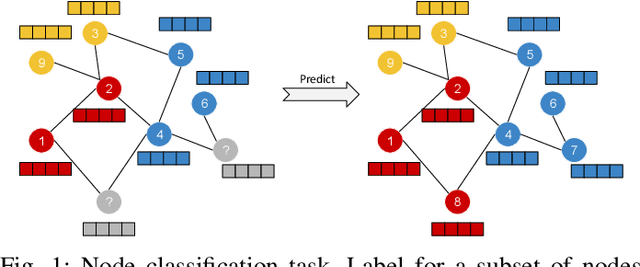
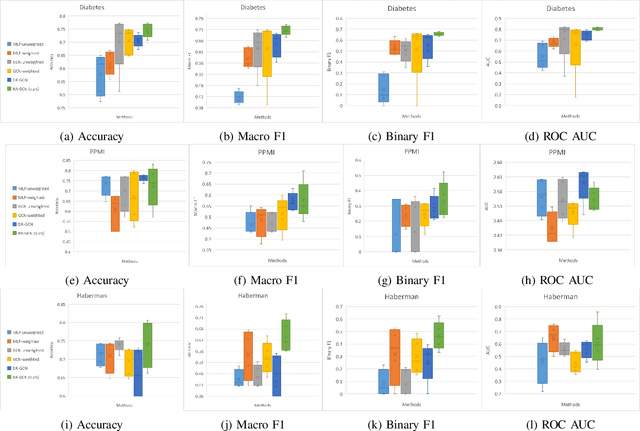
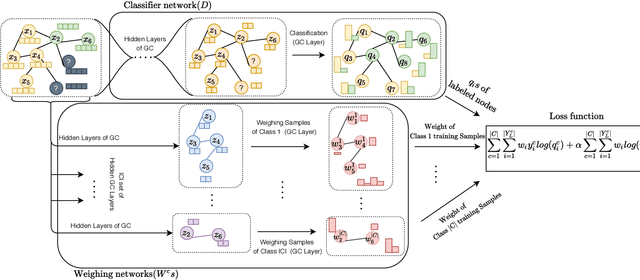
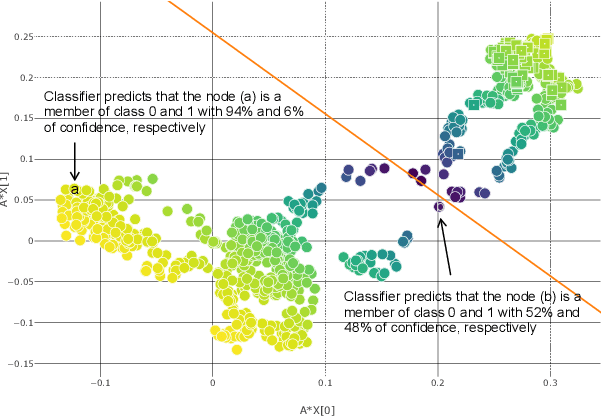
Abstract:Disease prediction is a well-known classification problem in medical applications. Graph neural networks provide a powerful tool for analyzing the patients' features relative to each other. Recently, Graph Convolutional Networks (GCNs) have particularly been studied in the field of disease prediction. Due to the nature of such medical datasets, the class imbalance is a familiar issue in the field of disease prediction. When the class imbalance is present in the data, the existing graph-based classifiers tend to be biased towards the major class(es). Meanwhile, the correct diagnosis of the rare true-positive cases among all the patients is vital. In conventional methods, such imbalance is tackled by assigning appropriate weights to classes in the loss function; however, this solution is still dependent on the relative values of weights, sensitive to outliers, and in some cases biased towards the minor class(es). In this paper, we propose Re-weighted Adversarial Graph Convolutional Network (RA-GCN) to enhance the performance of the graph-based classifier and prevent it from emphasizing the samples of any particular class. This is accomplished by automatically learning to weigh the samples of the classes. For this purpose, a graph-based network is associated with each class, which is responsible for weighing the class samples and informing the classifier about the importance of each sample. Therefore, the classifier adjusts itself and determines the boundary between classes with more attention to the important samples. The parameters of the classifier and weighing networks are trained by an adversarial approach. At the end of the adversarial training process, the boundary of the classifier is more accurate and unbiased. We show the superiority of RA-GCN on synthetic and three publicly available medical datasets compared to the recent method.
Deep Bingham Networks: Dealing with Uncertainty and Ambiguity in Pose Estimation
Dec 20, 2020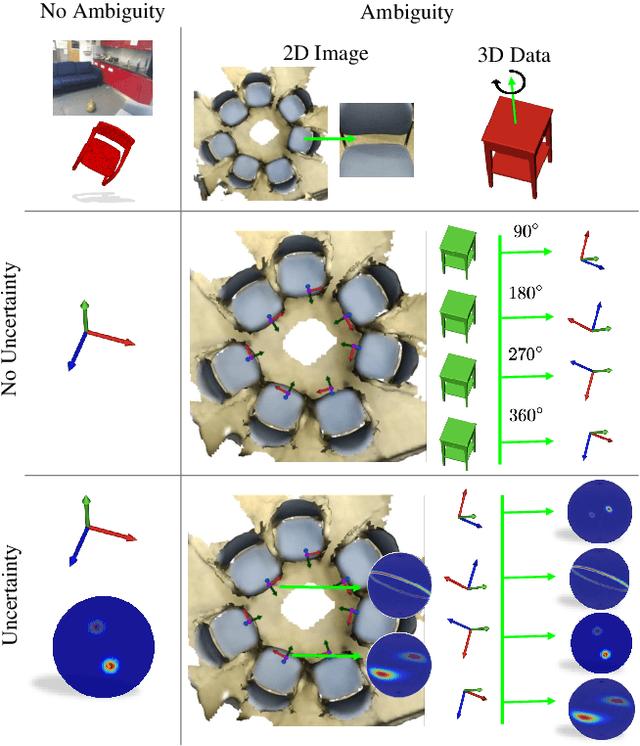
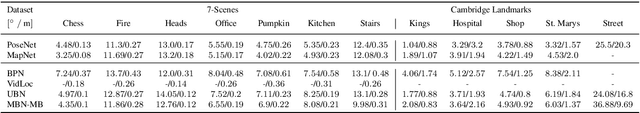
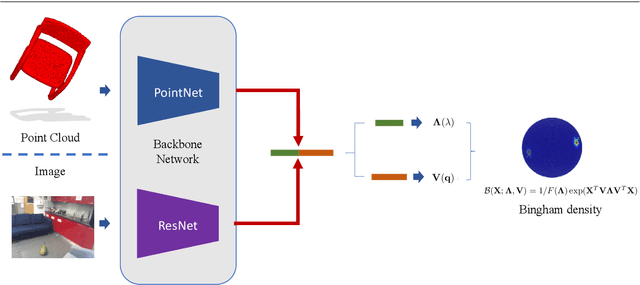
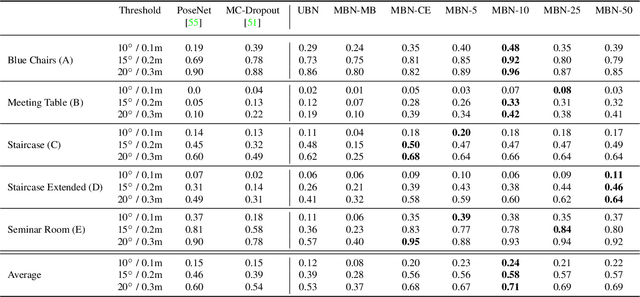
Abstract:In this work, we introduce Deep Bingham Networks (DBN), a generic framework that can naturally handle pose-related uncertainties and ambiguities arising in almost all real life applications concerning 3D data. While existing works strive to find a single solution to the pose estimation problem, we make peace with the ambiguities causing high uncertainty around which solutions to identify as the best. Instead, we report a family of poses which capture the nature of the solution space. DBN extends the state of the art direct pose regression networks by (i) a multi-hypotheses prediction head which can yield different distribution modes; and (ii) novel loss functions that benefit from Bingham distributions on rotations. This way, DBN can work both in unambiguous cases providing uncertainty information, and in ambiguous scenes where an uncertainty per mode is desired. On a technical front, our network regresses continuous Bingham mixture models and is applicable to both 2D data such as images and to 3D data such as point clouds. We proposed new training strategies so as to avoid mode or posterior collapse during training and to improve numerical stability. Our methods are thoroughly tested on two different applications exploiting two different modalities: (i) 6D camera relocalization from images; and (ii) object pose estimation from 3D point clouds, demonstrating decent advantages over the state of the art. For the former we contributed our own dataset composed of five indoor scenes where it is unavoidable to capture images corresponding to views that are hard to uniquely identify. For the latter we achieve the top results especially for symmetric objects of ModelNet dataset.
An Uncertainty-Driven GCN Refinement Strategy for Organ Segmentation
Dec 06, 2020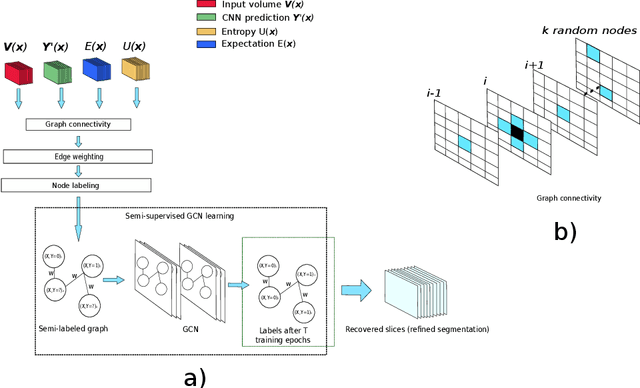
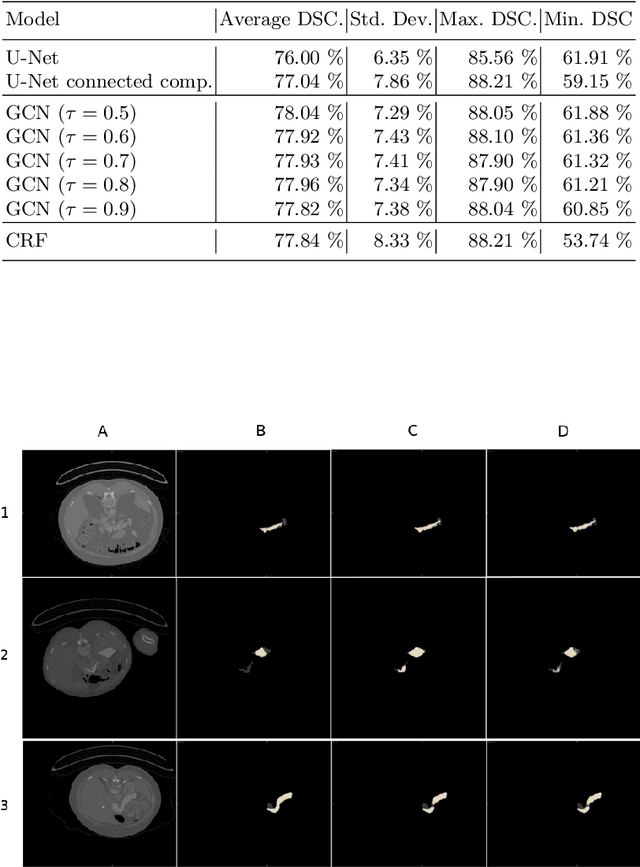
Abstract:Organ segmentation in CT volumes is an important pre-processing step in many computer assisted intervention and diagnosis methods. In recent years, convolutional neural networks have dominated the state of the art in this task. However, since this problem presents a challenging environment due to high variability in the organ's shape and similarity between tissues, the generation of false negative and false positive regions in the output segmentation is a common issue. Recent works have shown that the uncertainty analysis of the model can provide us with useful information about potential errors in the segmentation. In this context, we proposed a segmentation refinement method based on uncertainty analysis and graph convolutional networks. We employ the uncertainty levels of the convolutional network in a particular input volume to formulate a semi-supervised graph learning problem that is solved by training a graph convolutional network. To test our method we refine the initial output of a 2D U-Net. We validate our framework with the NIH pancreas dataset and the spleen dataset of the medical segmentation decathlon. We show that our method outperforms the state-of-the-art CRF refinement method by improving the dice score by 1% for the pancreas and 2% for spleen, with respect to the original U-Net's prediction. Finally, we perform a sensitivity analysis on the parameters of our proposal and discuss the applicability to other CNN architectures, the results, and current limitations of the model for future work in this research direction. For reproducibility purposes, we make our code publicly available at https://github.com/rodsom22/gcn_refinement.
Rethinking Positive Aggregation and Propagation of Gradients in Gradient-based Saliency Methods
Dec 01, 2020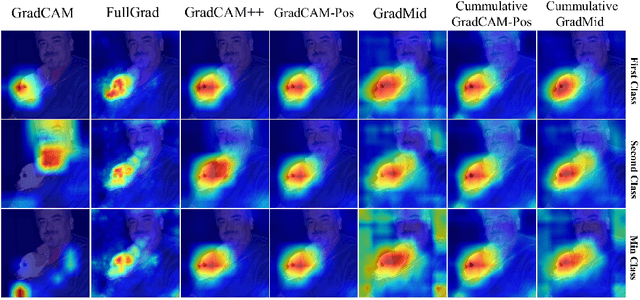
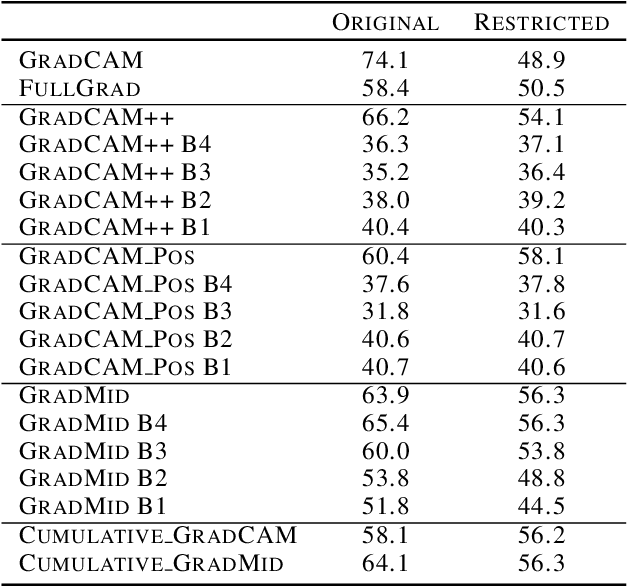

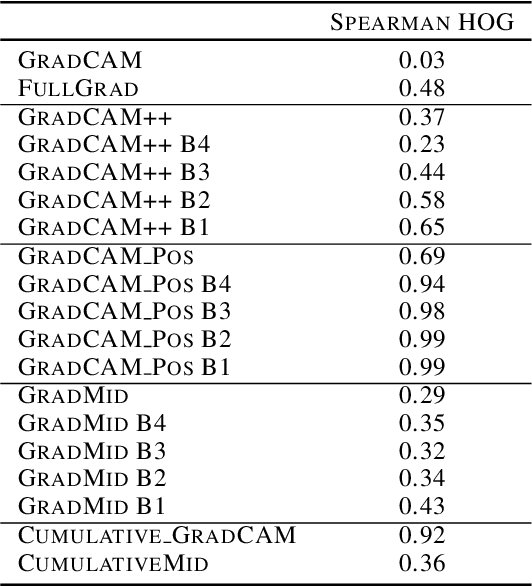
Abstract:Saliency methods interpret the prediction of a neural network by showing the importance of input elements for that prediction. A popular family of saliency methods utilize gradient information. In this work, we empirically show that two approaches for handling the gradient information, namely positive aggregation, and positive propagation, break these methods. Though these methods reflect visually salient information in the input, they do not explain the model prediction anymore as the generated saliency maps are insensitive to the predicted output and are insensitive to model parameter randomization. Specifically for methods that aggregate the gradients of a chosen layer such as GradCAM++ and FullGrad, exclusively aggregating positive gradients is detrimental. We further support this by proposing several variants of aggregation methods with positive handling of gradient information. For methods that backpropagate gradient information such as LRP, RectGrad, and Guided Backpropagation, we show the destructive effect of exclusively propagating positive gradient information.
Pose-dependent weights and Domain Randomization for fully automatic X-ray to CT Registration
Nov 14, 2020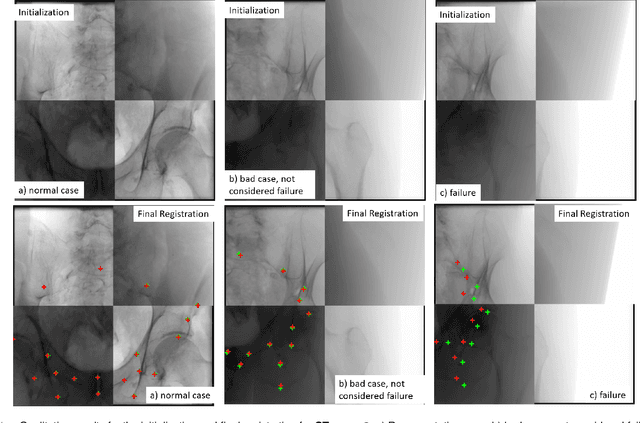
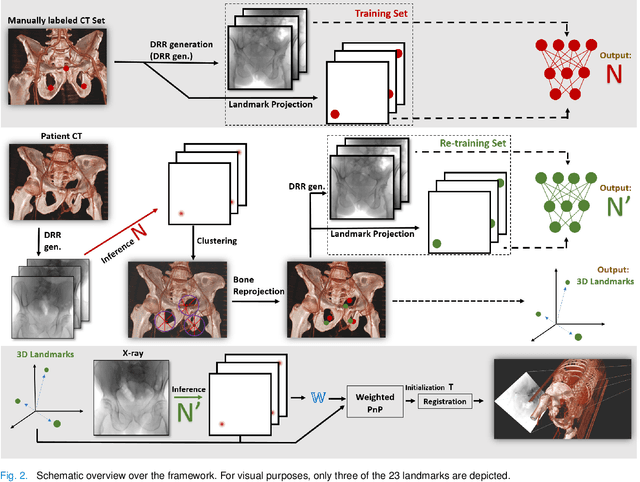
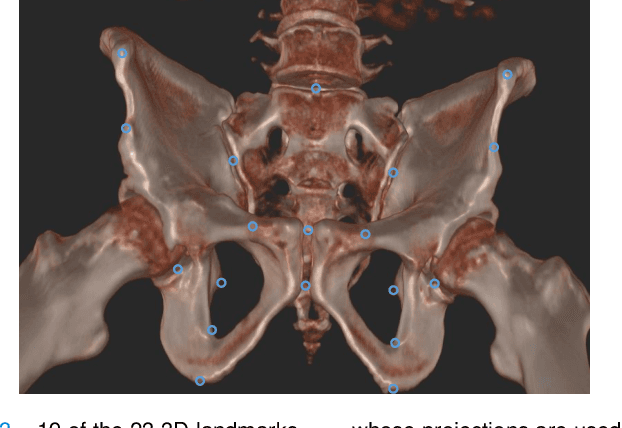
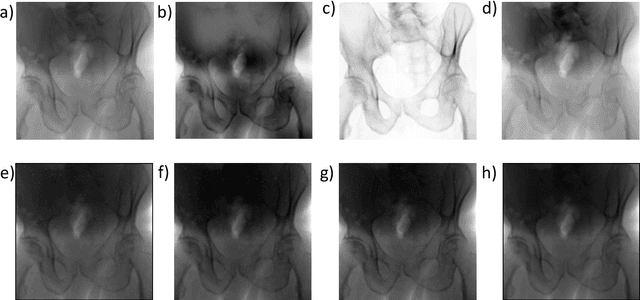
Abstract:Fully automatic X-ray to CT registration requires a solid initialization to provide an initial alignment within the capture range of existing intensity-based registrations. This work adresses that need by providing a novel automatic initialization, which enables end to end registration. First, a neural network is trained once to detect a set of anatomical landmarks on simulated X-rays. A domain randomization scheme is proposed to enable the network to overcome the challenge of being trained purely on simulated data and run inference on real Xrays. Then, for each patient CT, a patient-specific landmark extraction scheme is used. It is based on backprojecting and clustering the previously trained networks predictions on a set of simulated X-rays. Next, the network is retrained to detect the new landmarks. Finally the combination of network and 3D landmark locations is used to compute the initialization using a perspective-n-point algorithm. During the computation of the pose, a weighting scheme is introduced to incorporate the confidence of the network in detecting the landmarks. The algorithm is evaluated on the pelvis using both real and simulated x-rays. The mean (+-standard deviation) target registration error in millimetres is 4.1 +- 4.3 for simulated X-rays with a success rate of 92% and 4.2 +- 3.9 for real X-rays with a success rate of 86.8%, where a success is defined as a translation error of less than 30mm.
Self-Supervised Out-of-Distribution Detection in Brain CT Scans
Nov 10, 2020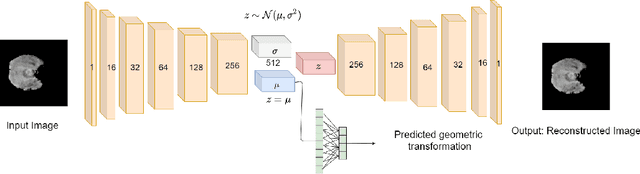
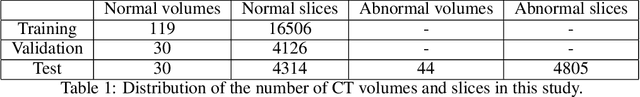
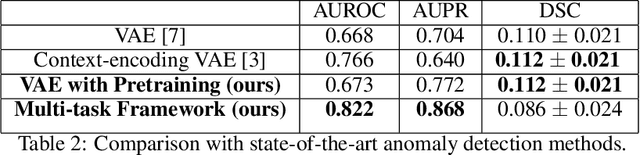
Abstract:Medical imaging data suffers from the limited availability of annotation because annotating 3D medical data is a time-consuming and expensive task. Moreover, even if the annotation is available, supervised learning-based approaches suffer highly imbalanced data. Most of the scans during the screening are from normal subjects, but there are also large variations in abnormal cases. To address these issues, recently, unsupervised deep anomaly detection methods that train the model on large-sized normal scans and detect abnormal scans by calculating reconstruction error have been reported. In this paper, we propose a novel self-supervised learning technique for anomaly detection. Our architecture largely consists of two parts: 1) Reconstruction and 2) predicting geometric transformations. By training the network to predict geometric transformations, the model could learn better image features and distribution of normal scans. In the test time, the geometric transformation predictor can assign the anomaly score by calculating the error between geometric transformation and prediction. Moreover, we further use self-supervised learning with context restoration for pretraining our model. By comparative experiments on clinical brain CT scans, the effectiveness of the proposed method has been verified.
Autonomous Robotic Screening of Tubular Structures based only on Real-Time Ultrasound Imaging Feedback
Oct 30, 2020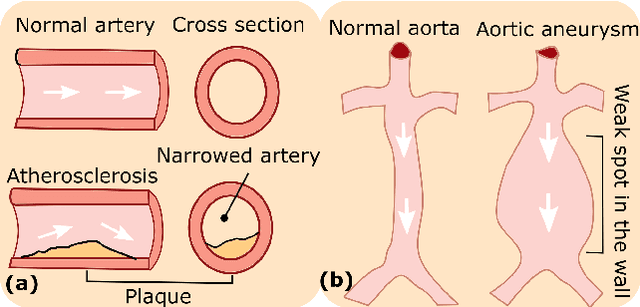
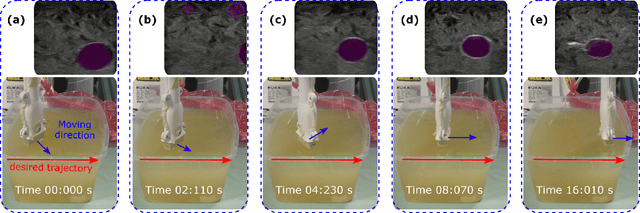

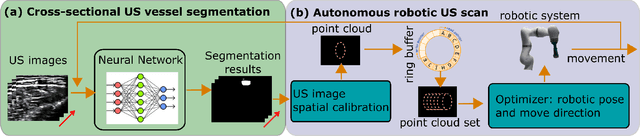
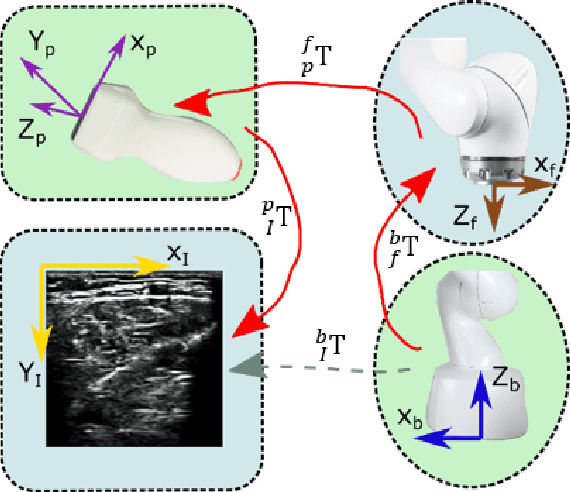
Abstract:Ultrasound (US) imaging is widely employed for diagnosis and staging of peripheral vascular diseases (PVD), mainly due to its high availability and the fact it does not emit radiation. However, high inter-operator variability and a lack of repeatability of US image acquisition hinder the implementation of extensive screening programs. To address this challenge, we propose an end-to-end workflow for automatic robotic US screening of tubular structures using only the real-time US imaging feedback. We first train a U-Net for real-time segmentation of the vascular structure from cross-sectional US images. Then, we represent the detected vascular structure as a 3D point cloud and use it to estimate the longitudinal axis of the target tubular structure and its mean radius by solving a constrained non-linear optimization problem. Iterating the previous processes, the US probe is automatically aligned to the orientation normal to the target tubular tissue and adjusted online to center the tracked tissue based on the spatial calibration. The real-time segmentation result is evaluated both on a phantom and in-vivo on brachial arteries of volunteers. In addition, the whole process is validated both in simulation and physical phantoms. The mean absolute radius error and orientation error ($\pm$ SD) in the simulation are $1.16\pm0.1~mm$ and $2.7\pm3.3^{\circ}$, respectively. On a gel phantom, these errors are $1.95\pm2.02~mm$ and $3.3\pm2.4^{\circ}$. This shows that the method is able to automatically screen tubular tissues with an optimal probe orientation (i.e. normal to the vessel) and at the same to accurately estimate the mean radius, both in real-time.
Surgical Data Science -- from Concepts to Clinical Translation
Oct 30, 2020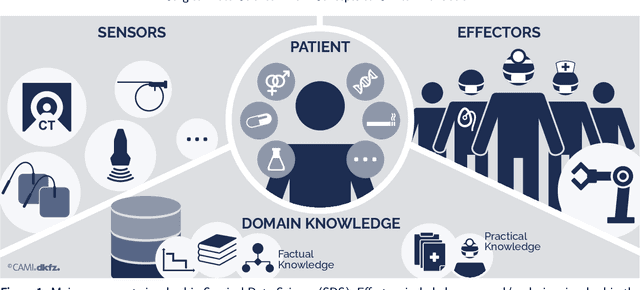
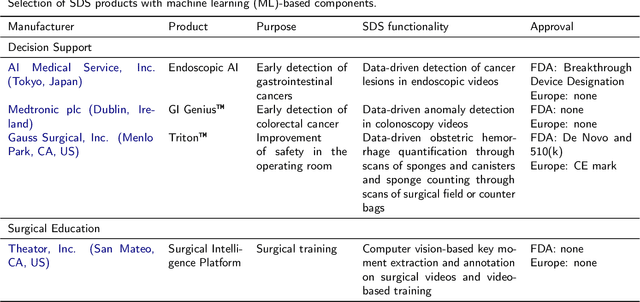
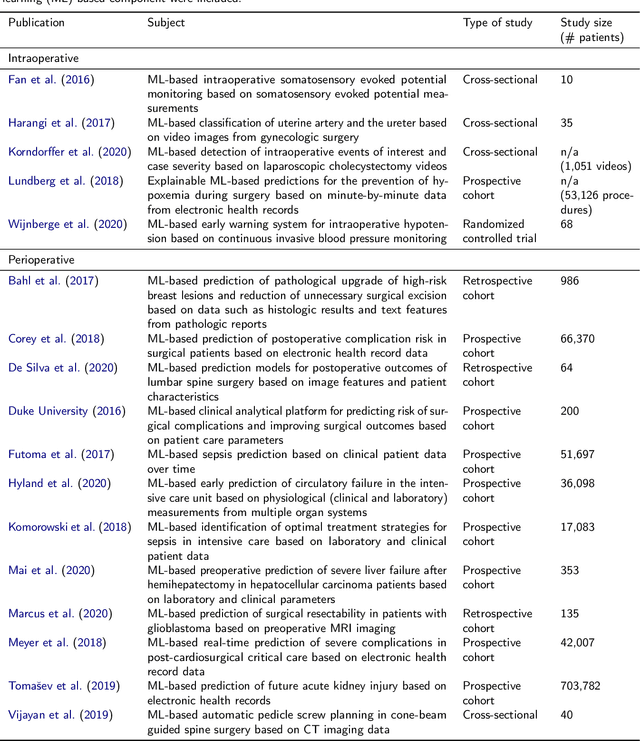
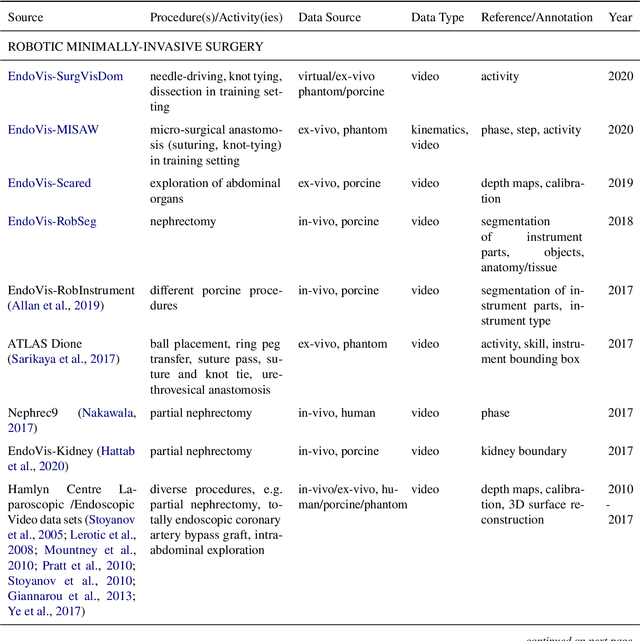
Abstract:Recent developments in data science in general and machine learning in particular have transformed the way experts envision the future of surgery. Surgical data science is a new research field that aims to improve the quality of interventional healthcare through the capture, organization, analysis and modeling of data. While an increasing number of data-driven approaches and clinical applications have been studied in the fields of radiological and clinical data science, translational success stories are still lacking in surgery. In this publication, we shed light on the underlying reasons and provide a roadmap for future advances in the field. Based on an international workshop involving leading researchers in the field of surgical data science, we review current practice, key achievements and initiatives as well as available standards and tools for a number of topics relevant to the field, namely (1) technical infrastructure for data acquisition, storage and access in the presence of regulatory constraints, (2) data annotation and sharing and (3) data analytics. Drawing from this extensive review, we present current challenges for technology development and (4) describe a roadmap for faster clinical translation and exploitation of the full potential of surgical data science.
Panoster: End-to-end Panoptic Segmentation of LiDAR Point Clouds
Oct 28, 2020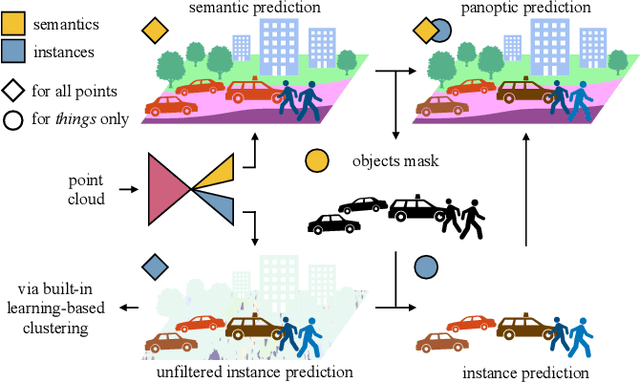
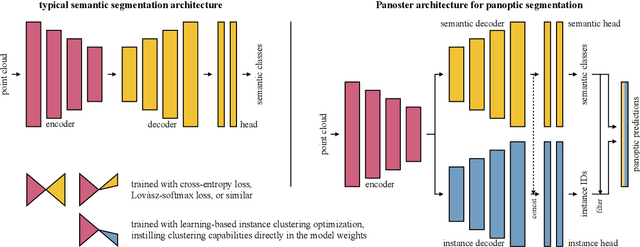
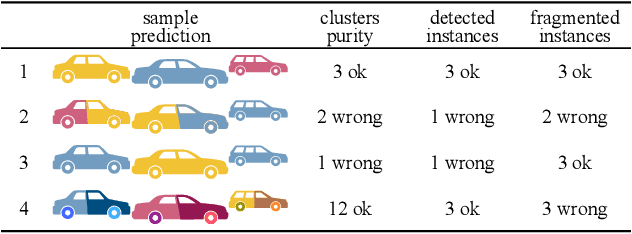
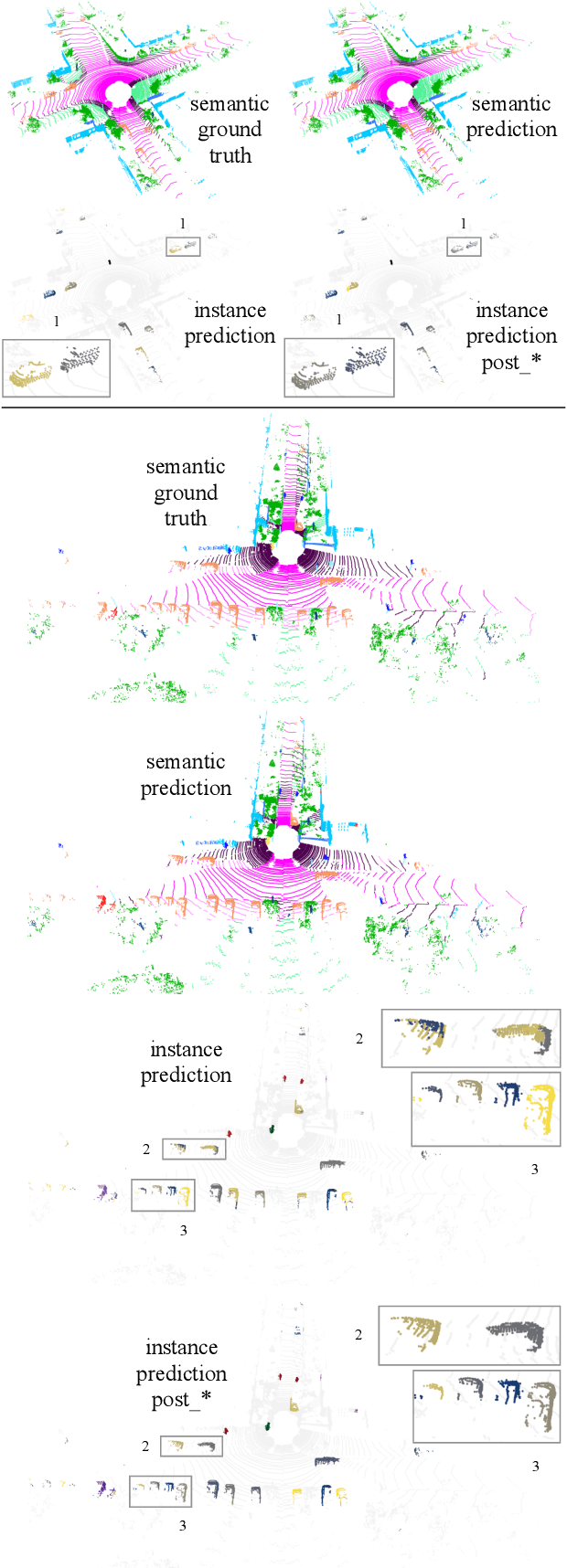
Abstract:Panoptic segmentation has recently unified semantic and instance segmentation, previously addressed separately, thus taking a step further towards creating more comprehensive and efficient perception systems. In this paper, we present Panoster, a novel proposal-free panoptic segmentation method for point clouds. Unlike previous approaches relying on several steps to group pixels or points into objects, Panoster proposes a simplified framework incorporating a learning-based clustering solution to identify instances. At inference time, this acts as a class-agnostic semantic segmentation, allowing Panoster to be fast, while outperforming prior methods in terms of accuracy. Additionally, we showcase how our approach can be flexibly and effectively applied on diverse existing semantic architectures to deliver panoptic predictions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge