Magdalini Paschali
Attention Head Entropy of LLMs Predicts Answer Correctness
Feb 14, 2026Abstract:Large language models (LLMs) often generate plausible yet incorrect answers, posing risks in safety-critical settings such as medicine. Human evaluation is expensive, and LLM-as-judge approaches risk introducing hidden errors. Recent white-box methods detect contextual hallucinations using model internals, focusing on the localization of the attention mass, but two questions remain open: do these approaches extend to predicting answer correctness, and do they generalize out-of-domains? We introduce Head Entropy, a method that predicts answer correctness from attention entropy patterns, specifically measuring the spread of the attention mass. Using sparse logistic regression on per-head 2-Renyi entropies, Head Entropy matches or exceeds baselines in-distribution and generalizes substantially better on out-of-domains, it outperforms the closest baseline on average by +8.5% AUROC. We further show that attention patterns over the question/context alone, before answer generation, already carry predictive signal using Head Entropy with on average +17.7% AUROC over the closest baseline. We evaluate across 5 instruction-tuned LLMs and 3 QA datasets spanning general knowledge, multi-hop reasoning, and medicine.
Automated Real-time Assessment of Intracranial Hemorrhage Detection AI Using an Ensembled Monitoring Model (EMM)
May 16, 2025Abstract:Artificial intelligence (AI) tools for radiology are commonly unmonitored once deployed. The lack of real-time case-by-case assessments of AI prediction confidence requires users to independently distinguish between trustworthy and unreliable AI predictions, which increases cognitive burden, reduces productivity, and potentially leads to misdiagnoses. To address these challenges, we introduce Ensembled Monitoring Model (EMM), a framework inspired by clinical consensus practices using multiple expert reviews. Designed specifically for black-box commercial AI products, EMM operates independently without requiring access to internal AI components or intermediate outputs, while still providing robust confidence measurements. Using intracranial hemorrhage detection as our test case on a large, diverse dataset of 2919 studies, we demonstrate that EMM successfully categorizes confidence in the AI-generated prediction, suggesting different actions and helping improve the overall performance of AI tools to ultimately reduce cognitive burden. Importantly, we provide key technical considerations and best practices for successfully translating EMM into clinical settings.
Foundation Models in Radiology: What, How, When, Why and Why Not
Nov 27, 2024



Abstract:Recent advances in artificial intelligence have witnessed the emergence of large-scale deep learning models capable of interpreting and generating both textual and imaging data. Such models, typically referred to as foundation models, are trained on extensive corpora of unlabeled data and demonstrate high performance across various tasks. Foundation models have recently received extensive attention from academic, industry, and regulatory bodies. Given the potentially transformative impact that foundation models can have on the field of radiology, this review aims to establish a standardized terminology concerning foundation models, with a specific focus on the requirements of training data, model training paradigms, model capabilities, and evaluation strategies. We further outline potential pathways to facilitate the training of radiology-specific foundation models, with a critical emphasis on elucidating both the benefits and challenges associated with such models. Overall, we envision that this review can unify technical advances and clinical needs in the training of foundation models for radiology in a safe and responsible manner, for ultimately benefiting patients, providers, and radiologists.
SOE: SO(3)-Equivariant 3D MRI Encoding
Oct 15, 2024



Abstract:Representation learning has become increasingly important, especially as powerful models have shifted towards learning latent representations before fine-tuning for downstream tasks. This approach is particularly valuable in leveraging the structural information within brain anatomy. However, a common limitation of recent models developed for MRIs is their tendency to ignore or remove geometric information, such as translation and rotation, thereby creating invariance with respect to geometric operations. We contend that incorporating knowledge about these geometric transformations into the model can significantly enhance its ability to learn more detailed anatomical information within brain structures. As a result, we propose a novel method for encoding 3D MRIs that enforces equivariance with respect to all rotations in 3D space, in other words, SO(3)-equivariance (SOE). By explicitly modeling this geometric equivariance in the representation space, we ensure that any rotational operation applied to the input image space is also reflected in the embedding representation space. This approach requires moving beyond traditional representation learning methods, as we need a representation vector space that allows for the application of the same SO(3) operation in that space. To facilitate this, we leverage the concept of vector neurons. The representation space formed by our method captures the brain's structural and anatomical information more effectively. We evaluate SOE pretrained on the structural MRIs of two public data sets with respect to the downstream task of predicting age and diagnosing Alzheimer's Disease from T1-weighted brain scans of the ADNI data set. We demonstrate that our approach not only outperforms other methods but is also robust against various degrees of rotation along different axes. The code is available at https://github.com/shizhehe/SOE-representation-learning.
Spectral Graph Sample Weighting for Interpretable Sub-cohort Analysis in Predictive Models for Neuroimaging
Oct 01, 2024Abstract:Recent advancements in medicine have confirmed that brain disorders often comprise multiple subtypes of mechanisms, developmental trajectories, or severity levels. Such heterogeneity is often associated with demographic aspects (e.g., sex) or disease-related contributors (e.g., genetics). Thus, the predictive power of machine learning models used for symptom prediction varies across subjects based on such factors. To model this heterogeneity, one can assign each training sample a factor-dependent weight, which modulates the subject's contribution to the overall objective loss function. To this end, we propose to model the subject weights as a linear combination of the eigenbases of a spectral population graph that captures the similarity of factors across subjects. In doing so, the learned weights smoothly vary across the graph, highlighting sub-cohorts with high and low predictability. Our proposed sample weighting scheme is evaluated on two tasks. First, we predict initiation of heavy alcohol drinking in young adulthood from imaging and neuropsychological measures from the National Consortium on Alcohol and NeuroDevelopment in Adolescence (NCANDA). Next, we detect Dementia vs. Mild Cognitive Impairment (MCI) using imaging and demographic measurements in subjects from the Alzheimer's Disease Neuroimaging Initiative (ADNI). Compared to existing sample weighting schemes, our sample weights improve interpretability and highlight sub-cohorts with distinct characteristics and varying model accuracy.
Merlin: A Vision Language Foundation Model for 3D Computed Tomography
Jun 10, 2024



Abstract:Over 85 million computed tomography (CT) scans are performed annually in the US, of which approximately one quarter focus on the abdomen. Given the current radiologist shortage, there is a large impetus to use artificial intelligence to alleviate the burden of interpreting these complex imaging studies. Prior state-of-the-art approaches for automated medical image interpretation leverage vision language models (VLMs). However, current medical VLMs are generally limited to 2D images and short reports, and do not leverage electronic health record (EHR) data for supervision. We introduce Merlin - a 3D VLM that we train using paired CT scans (6+ million images from 15,331 CTs), EHR diagnosis codes (1.8+ million codes), and radiology reports (6+ million tokens). We evaluate Merlin on 6 task types and 752 individual tasks. The non-adapted (off-the-shelf) tasks include zero-shot findings classification (31 findings), phenotype classification (692 phenotypes), and zero-shot cross-modal retrieval (image to findings and image to impressions), while model adapted tasks include 5-year disease prediction (6 diseases), radiology report generation, and 3D semantic segmentation (20 organs). We perform internal validation on a test set of 5,137 CTs, and external validation on 7,000 clinical CTs and on two public CT datasets (VerSe, TotalSegmentator). Beyond these clinically-relevant evaluations, we assess the efficacy of various network architectures and training strategies to depict that Merlin has favorable performance to existing task-specific baselines. We derive data scaling laws to empirically assess training data needs for requisite downstream task performance. Furthermore, unlike conventional VLMs that require hundreds of GPUs for training, we perform all training on a single GPU.
CheXagent: Towards a Foundation Model for Chest X-Ray Interpretation
Jan 22, 2024



Abstract:Chest X-rays (CXRs) are the most frequently performed imaging test in clinical practice. Recent advances in the development of vision-language foundation models (FMs) give rise to the possibility of performing automated CXR interpretation, which can assist physicians with clinical decision-making and improve patient outcomes. However, developing FMs that can accurately interpret CXRs is challenging due to the (1) limited availability of large-scale vision-language datasets in the medical image domain, (2) lack of vision and language encoders that can capture the complexities of medical data, and (3) absence of evaluation frameworks for benchmarking the abilities of FMs on CXR interpretation. In this work, we address these challenges by first introducing \emph{CheXinstruct} - a large-scale instruction-tuning dataset curated from 28 publicly-available datasets. We then present \emph{CheXagent} - an instruction-tuned FM capable of analyzing and summarizing CXRs. To build CheXagent, we design a clinical large language model (LLM) for parsing radiology reports, a vision encoder for representing CXR images, and a network to bridge the vision and language modalities. Finally, we introduce \emph{CheXbench} - a novel benchmark designed to systematically evaluate FMs across 8 clinically-relevant CXR interpretation tasks. Extensive quantitative evaluations and qualitative reviews with five expert radiologists demonstrate that CheXagent outperforms previously-developed general- and medical-domain FMs on CheXbench tasks. Furthermore, in an effort to improve model transparency, we perform a fairness evaluation across factors of sex, race and age to highlight potential performance disparities. Our project is at \url{https://stanford-aimi.github.io/chexagent.html}.
Self-Supervised Learning for Physiologically-Based Pharmacokinetic Modeling in Dynamic PET
May 17, 2023



Abstract:Dynamic positron emission tomography imaging (dPET) provides temporally resolved images of a tracer enabling a quantitative measure of physiological processes. Voxel-wise physiologically-based pharmacokinetic (PBPK) modeling of the time activity curves (TAC) can provide relevant diagnostic information for clinical workflow. Conventional fitting strategies for TACs are slow and ignore the spatial relation between neighboring voxels. We train a spatio-temporal UNet to estimate the kinetic parameters given TAC from F-18-fluorodeoxyglucose (FDG) dPET. This work introduces a self-supervised loss formulation to enforce the similarity between the measured TAC and those generated with the learned kinetic parameters. Our method provides quantitatively comparable results at organ-level to the significantly slower conventional approaches, while generating pixel-wise parametric images which are consistent with expected physiology. To the best of our knowledge, this is the first self-supervised network that allows voxel-wise computation of kinetic parameters consistent with a non-linear kinetic model. The code will become publicly available upon acceptance.
Investigating Pulse-Echo Sound Speed Estimation in Breast Ultrasound with Deep Learning
Feb 06, 2023



Abstract:Ultrasound is an adjunct tool to mammography that can quickly and safely aid physicians with diagnosing breast abnormalities. Clinical ultrasound often assumes a constant sound speed to form B-mode images for diagnosis. However, the various types of breast tissue, such as glandular, fat, and lesions, differ in sound speed. These differences can degrade the image reconstruction process. Alternatively, sound speed can be a powerful tool for identifying disease. To this end, we propose a deep-learning approach for sound speed estimation from in-phase and quadrature ultrasound signals. First, we develop a large-scale simulated ultrasound dataset that generates quasi-realistic breast tissue by modeling breast gland, skin, and lesions with varying echogenicity and sound speed. We developed a fully convolutional neural network architecture trained on a simulated dataset to produce an estimated sound speed map from inputting three complex-value in-phase and quadrature ultrasound images formed from plane-wave transmissions at separate angles. Furthermore, thermal noise augmentation is used during model optimization to enhance generalizability to real ultrasound data. We evaluate the model on simulated, phantom, and in-vivo breast ultrasound data, demonstrating its ability to accurately estimate sound speeds consistent with previously reported values in the literature. Our simulated dataset and model will be publicly available to provide a step towards accurate and generalizable sound speed estimation for pulse-echo ultrasound imaging.
Bridging the Gap between Deep Learning and Hypothesis-Driven Analysis via Permutation Testing
Jul 28, 2022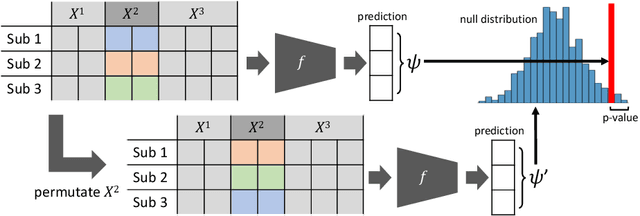
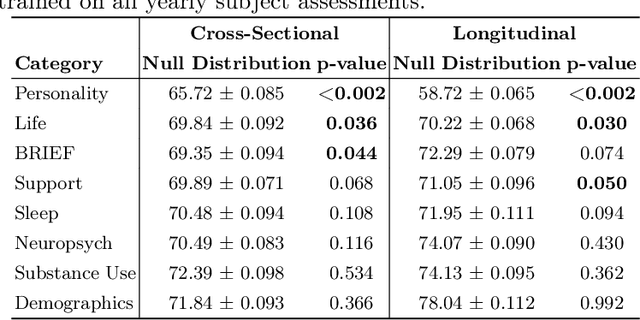

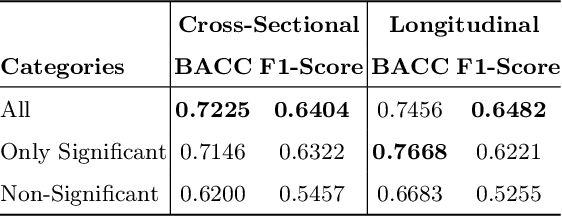
Abstract:A fundamental approach in neuroscience research is to test hypotheses based on neuropsychological and behavioral measures, i.e., whether certain factors (e.g., related to life events) are associated with an outcome (e.g., depression). In recent years, deep learning has become a potential alternative approach for conducting such analyses by predicting an outcome from a collection of factors and identifying the most "informative" ones driving the prediction. However, this approach has had limited impact as its findings are not linked to statistical significance of factors supporting hypotheses. In this article, we proposed a flexible and scalable approach based on the concept of permutation testing that integrates hypothesis testing into the data-driven deep learning analysis. We apply our approach to the yearly self-reported assessments of 621 adolescent participants of the National Consortium of Alcohol and Neurodevelopment in Adolescence (NCANDA) to predict negative valence, a symptom of major depressive disorder according to the NIMH Research Domain Criteria (RDoC). Our method successfully identifies categories of risk factors that further explain the symptom.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge