Mathias Unberath
Open-H-Embodiment: A Large-Scale Dataset for Enabling Foundation Models in Medical Robotics
Apr 22, 2026Abstract:Autonomous medical robots hold promise to improve patient outcomes, reduce provider workload, democratize access to care, and enable superhuman precision. However, autonomous medical robotics has been limited by a fundamental data problem: existing medical robotic datasets are small, single-embodiment, and rarely shared openly, restricting the development of foundation models that the field needs to advance. We introduce Open-H-Embodiment, the largest open dataset of medical robotic video with synchronized kinematics to date, spanning more than 49 institutions and multiple robotic platforms including the CMR Versius, Intuitive Surgical's da Vinci, da Vinci Research Kit (dVRK), Rob Surgical BiTrack, Virtual Incision's MIRA, Moon Surgical Maestro, and a variety of custom systems, spanning surgical manipulation, robotic ultrasound, and endoscopy procedures. We demonstrate the research enabled by this dataset through two foundation models. GR00T-H is the first open foundation vision-language-action model for medical robotics, which is the only evaluated model to achieve full end-to-end task completion on a structured suturing benchmark (25% of trials vs. 0% for all others) and achieves 64% average success across a 29-step ex vivo suturing sequence. We also train Cosmos-H-Surgical-Simulator, the first action-conditioned world model to enable multi-embodiment surgical simulation from a single checkpoint, spanning nine robotic platforms and supporting in silico policy evaluation and synthetic data generation for the medical domain. These results suggest that open, large-scale medical robot data collection can serve as critical infrastructure for the research community, enabling advances in robot learning, world modeling, and beyond.
AffordTissue: Dense Affordance Prediction for Tool-Action Specific Tissue Interaction
Apr 01, 2026Abstract:Surgical action automation has progressed rapidly toward achieving surgeon-like dexterous control, driven primarily by advances in learning from demonstration and vision-language-action models. While these have demonstrated success in table-top experiments, translating them to clinical deployment remains challenging: current methods offer limited predictability on where instruments will interact on tissue surfaces and lack explicit conditioning inputs to enforce tool-action-specific safe interaction regions. Addressing this gap, we introduce AffordTissue, a multimodal framework for predicting tool-action specific tissue affordance regions as dense heatmaps during cholecystectomy. Our approach combines a temporal vision encoder capturing tool motion and tissue dynamics across multiple viewpoints, language conditioning enabling generalization across diverse instrument-action pairs, and a DiT-style decoder for dense affordance prediction. We establish the first tissue affordance benchmark by curating and annotating 15,638 video clips across 103 cholecystectomy procedures, covering six unique tool-action pairs involving four instruments (hook, grasper, scissors, clipper) and their associated tasks: dissection, grasping, clipping, and cutting. Experiments demonstrate substantial improvement over vision-language model baselines (20.6 px ASSD vs. 60.2 px for Molmo-VLM), showing that our task-specific architecture outperforms large-scale foundation models for dense surgical affordance prediction. By predicting tool-action specific tissue affordance regions, AffordTissue provides explicit spatial reasoning for safe surgical automation, potentially unlocking explicit policy guidance toward appropriate tissue regions and early safe stop when instruments deviate outside predicted safe zones.
Investigating a Policy-Based Formulation for Endoscopic Camera Pose Recovery
Mar 20, 2026Abstract:In endoscopic surgery, surgeons continuously locate the endoscopic view relative to the anatomy by interpreting the evolving visual appearance of the intraoperative scene in the context of their prior knowledge. Vision-based navigation systems seek to replicate this capability by recovering camera pose directly from endoscopic video, but most approaches do not embody the same principles of reasoning about new frames that makes surgeons successful. Instead, they remain grounded in feature matching and geometric optimization over keyframes, an approach that has been shown to degrade under the challenging conditions of endoscopic imaging like low texture and rapid illumination changes. Here, we pursue an alternative approach and investigate a policy-based formulation of endoscopic camera pose recovery that seeks to imitate experts in estimating trajectories conditioned on the previous camera state. Our approach directly predicts short-horizon relative motions without maintaining an explicit geometric representation at inference time. It thus addresses, by design, some of the notorious challenges of geometry-based approaches, such as brittle correspondence matching, instability in texture-sparse regions, and limited pose coverage due to reconstruction failure. We evaluate the proposed formulation on cadaveric sinus endoscopy. Under oracle state conditioning, we compare short-horizon motion prediction quality to geometric baselines achieving lowest mean translation error and competitive rotational accuracy. We analyze robustness by grouping prediction windows according to texture richness and illumination change indicating reduced sensitivity to low-texture conditions. These findings suggest that a learned motion policy offers a viable alternative formulation for endoscopic camera pose recovery.
SAW: Toward a Surgical Action World Model via Controllable and Scalable Video Generation
Mar 13, 2026Abstract:A surgical world model capable of generating realistic surgical action videos with precise control over tool-tissue interactions can address fundamental challenges in surgical AI and simulation -- from data scarcity and rare event synthesis to bridging the sim-to-real gap for surgical automation. However, current video generation methods, the very core of such surgical world models, require expensive annotations or complex structured intermediates as conditioning signals at inference, limiting their scalability. Other approaches exhibit limited temporal consistency across complex laparoscopic scenes and do not possess sufficient realism. We propose Surgical Action World (SAW) -- a step toward surgical action world modeling through video diffusion conditioned on four lightweight signals: language prompts encoding tool-action context, a reference surgical scene, tissue affordance mask, and 2D tool-tip trajectories. We design a conditional video diffusion approach that reformulates video-to-video diffusion into trajectory-conditioned surgical action synthesis. The backbone diffusion model is fine-tuned on a custom-curated dataset of 12,044 laparoscopic clips with lightweight spatiotemporal conditioning signals, leveraging a depth consistency loss to enforce geometric plausibility without requiring depth at inference. SAW achieves state-of-the-art temporal consistency (CD-FVD: 199.19 vs. 546.82) and strong visual quality on held-out test data. Furthermore, we demonstrate its downstream utility for (a) surgical AI, where augmenting rare actions with SAW-generated videos improves action recognition (clipping F1-score: 20.93% to 43.14%; cutting: 0.00% to 8.33%) on real test data, and (b) surgical simulation, where rendering tool-tissue interaction videos from simulator-derived trajectory points toward a visually faithful simulation engine.
Humanoid Robots as First Assistants in Endoscopic Surgery
Feb 27, 2026Abstract:Humanoid robots have become a focal point of technological ambition, with claims of surgical capability within years in mainstream discourse. These projections are aspirational yet lack empirical grounding. To date, no humanoid has assisted a surgeon through an actual procedure, let alone performed one. The work described here breaks this new ground. Here we report a proof of concept in which a teleoperated Unitree G1 provided endoscopic visualization while an attending otolaryngologist performed a cadaveric sphenoidectomy. The procedure was completed successfully, with stable visualization maintained throughout. Teleoperation allowed assessment of whether the humanoid form factor could meet the physical demands of surgical assistance in terms of sustenance and precision; the cognitive demands were satisfied -- for now -- by the operator. Post-procedure analysis identified engineering targets for clinical translation, alongside near-term opportunities such as autonomous diagnostic scoping. This work establishes form-factor feasibility for humanoid surgical assistance while identifying challenges for continued development.
Towards Controllable Video Synthesis of Routine and Rare OR Events
Feb 24, 2026Abstract:Purpose: Curating large-scale datasets of operating room (OR) workflow, encompassing rare, safety-critical, or atypical events, remains operationally and ethically challenging. This data bottleneck complicates the development of ambient intelligence for detecting, understanding, and mitigating rare or safety-critical events in the OR. Methods: This work presents an OR video diffusion framework that enables controlled synthesis of rare and safety-critical events. The framework integrates a geometric abstraction module, a conditioning module, and a fine-tuned diffusion model to first transform OR scenes into abstract geometric representations, then condition the synthesis process, and finally generate realistic OR event videos. Using this framework, we also curate a synthetic dataset to train and validate AI models for detecting near-misses of sterile-field violations. Results: In synthesizing routine OR events, our method outperforms off-the-shelf video diffusion baselines, achieving lower FVD/LPIPS and higher SSIM/PSNR in both in- and out-of-domain datasets. Through qualitative results, we illustrate its ability for controlled video synthesis of counterfactual events. An AI model trained and validated on the generated synthetic data achieved a RECALL of 70.13% in detecting near safety-critical events. Finally, we conduct an ablation study to quantify performance gains from key design choices. Conclusion: Our solution enables controlled synthesis of routine and rare OR events from abstract geometric representations. Beyond demonstrating its capability to generate rare and safety-critical scenarios, we show its potential to support the development of ambient intelligence models.
Text-Driven Reasoning Video Editing via Reinforcement Learning on Digital Twin Representations
Nov 18, 2025Abstract:Text-driven video editing enables users to modify video content only using text queries. While existing methods can modify video content if explicit descriptions of editing targets with precise spatial locations and temporal boundaries are provided, these requirements become impractical when users attempt to conceptualize edits through implicit queries referencing semantic properties or object relationships. We introduce reasoning video editing, a task where video editing models must interpret implicit queries through multi-hop reasoning to infer editing targets before executing modifications, and a first model attempting to solve this complex task, RIVER (Reasoning-based Implicit Video Editor). RIVER decouples reasoning from generation through digital twin representations of video content that preserve spatial relationships, temporal trajectories, and semantic attributes. A large language model then processes this representation jointly with the implicit query, performing multi-hop reasoning to determine modifications, then outputs structured instructions that guide a diffusion-based editor to execute pixel-level changes. RIVER training uses reinforcement learning with rewards that evaluate reasoning accuracy and generation quality. Finally, we introduce RVEBenchmark, a benchmark of 100 videos with 519 implicit queries spanning three levels and categories of reasoning complexity specifically for reasoning video editing. RIVER demonstrates best performance on the proposed RVEBenchmark and also achieves state-of-the-art performance on two additional video editing benchmarks (VegGIE and FiVE), where it surpasses six baseline methods.
Fast Reasoning Segmentation for Images and Videos
Nov 15, 2025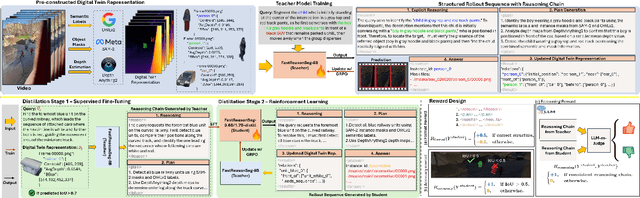
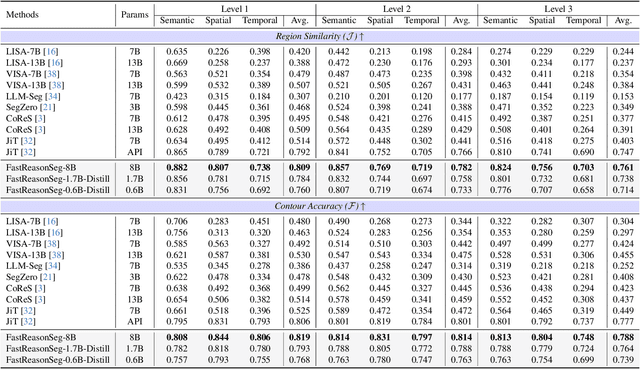
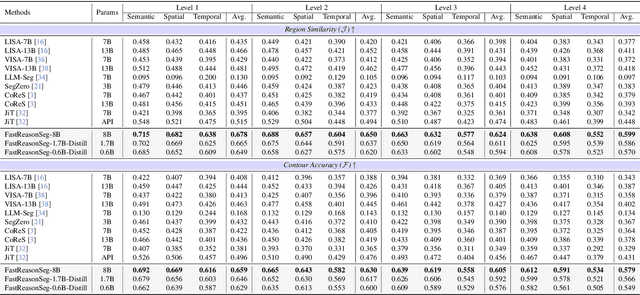

Abstract:Reasoning segmentation enables open-set object segmentation via implicit text queries, therefore serving as a foundation for embodied agents that should operate autonomously in real-world environments. However, existing methods for reasoning segmentation require multimodal large language models with billions of parameters that exceed the computational capabilities of edge devices that typically deploy the embodied AI systems. Distillation offers a pathway to compress these models while preserving their capabilities. Yet, existing distillation approaches fail to transfer the multi-step reasoning capabilities that reasoning segmentation demands, as they focus on matching output predictions and intermediate features rather than preserving reasoning chains. The emerging paradigm of reasoning over digital twin representations presents an opportunity for more effective distillation by re-framing the problem. Consequently, we propose FastReasonSeg, which employs digital twin representations that decouple perception from reasoning to enable more effective distillation. Our distillation scheme first relies on supervised fine-tuning on teacher-generated reasoning chains. Then it is followed by reinforcement fine-tuning with joint rewards evaluating both segmentation accuracy and reasoning quality alignment. Experiments on two video (JiTBench, RVTBench) and two image benchmarks (ReasonSeg, LLM-Seg40K) demonstrate that our FastReasonSeg achieves state-of-the-art reasoning segmentation performance. Moreover, the distilled 0.6B variant outperforms models with 20 times more parameters while achieving 7.79 FPS throughput with only 2.1GB memory consumption. This efficiency enables deployment in resource-constrained environments to enable real-time reasoning segmentation.
Reasoning Text-to-Video Retrieval via Digital Twin Video Representations and Large Language Models
Nov 15, 2025Abstract:The goal of text-to-video retrieval is to search large databases for relevant videos based on text queries. Existing methods have progressed to handling explicit queries where the visual content of interest is described explicitly; however, they fail with implicit queries where identifying videos relevant to the query requires reasoning. We introduce reasoning text-to-video retrieval, a paradigm that extends traditional retrieval to process implicit queries through reasoning while providing object-level grounding masks that identify which entities satisfy the query conditions. Instead of relying on vision-language models directly, we propose representing video content as digital twins, i.e., structured scene representations that decompose salient objects through specialist vision models. This approach is beneficial because it enables large language models to reason directly over long-horizon video content without visual token compression. Specifically, our two-stage framework first performs compositional alignment between decomposed sub-queries and digital twin representations for candidate identification, then applies large language model-based reasoning with just-in-time refinement that invokes additional specialist models to address information gaps. We construct a benchmark of 447 manually created implicit queries with 135 videos (ReasonT2VBench-135) and another more challenging version of 1000 videos (ReasonT2VBench-1000). Our method achieves 81.2% R@1 on ReasonT2VBench-135, outperforming the strongest baseline by greater than 50 percentage points, and maintains 81.7% R@1 on the extended configuration while establishing state-of-the-art results in three conventional benchmarks (MSR-VTT, MSVD, and VATEX).
Constructing and Interpreting Digital Twin Representations for Visual Reasoning via Reinforcement Learning
Nov 15, 2025Abstract:Visual reasoning may require models to interpret images and videos and respond to implicit text queries across diverse output formats, from pixel-level segmentation masks to natural language descriptions. Existing approaches rely on supervised fine-tuning with task-specific architectures. For example, reasoning segmentation, grounding, summarization, and visual question answering each demand distinct model designs and training, preventing unified solutions and limiting cross-task and cross-modality generalization. Hence, we propose DT-R1, a reinforcement learning framework that trains large language models to construct digital twin representations of complex multi-modal visual inputs and then reason over these high-level representations as a unified approach to visual reasoning. Specifically, we train DT-R1 using GRPO with a novel reward that validates both structural integrity and output accuracy. Evaluations in six visual reasoning benchmarks, covering two modalities and four task types, demonstrate that DT-R1 consistently achieves improvements over state-of-the-art task-specific models. DT-R1 opens a new direction where visual reasoning emerges from reinforcement learning with digital twin representations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge