Juntang Zhuang
Tony
Adaptive Checkpoint Adjoint Method for Gradient Estimation in Neural ODE
Jun 03, 2020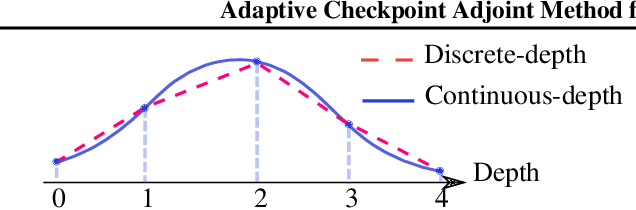
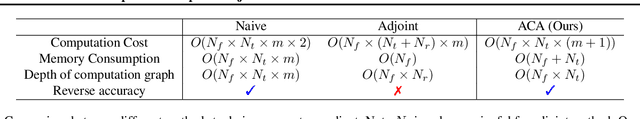
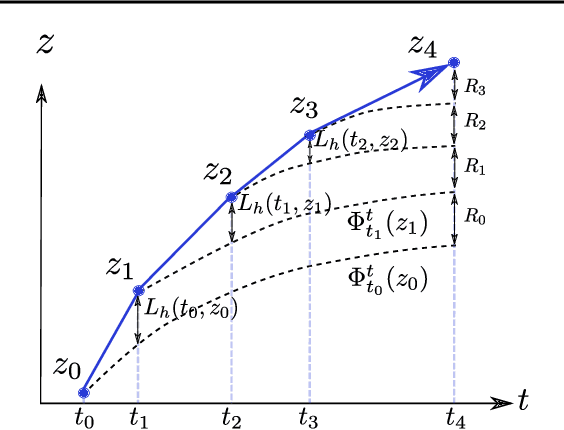
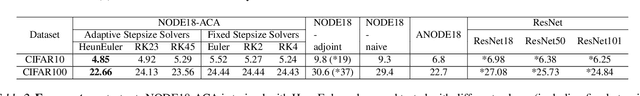
Abstract:Neural ordinary differential equations (NODEs) have recently attracted increasing attention; however, their empirical performance on benchmark tasks (e.g. image classification) are significantly inferior to discrete-layer models. We demonstrate an explanation for their poorer performance is the inaccuracy of existing gradient estimation methods: the adjoint method has numerical errors in reverse-mode integration; the naive method directly back-propagates through ODE solvers, but suffers from a redundantly deep computation graph when searching for the optimal stepsize. We propose the Adaptive Checkpoint Adjoint (ACA) method: in automatic differentiation, ACA applies a trajectory checkpoint strategy which records the forward-mode trajectory as the reverse-mode trajectory to guarantee accuracy; ACA deletes redundant components for shallow computation graphs; and ACA supports adaptive solvers. On image classification tasks, compared with the adjoint and naive method, ACA achieves half the error rate in half the training time; NODE trained with ACA outperforms ResNet in both accuracy and test-retest reliability. On time-series modeling, ACA outperforms competing methods. Finally, in an example of the three-body problem, we show NODE with ACA can incorporate physical knowledge to achieve better accuracy. We provide the PyTorch implementation of ACA: \url{https://github.com/juntang-zhuang/torch-ACA}.
2018 Robotic Scene Segmentation Challenge
Feb 03, 2020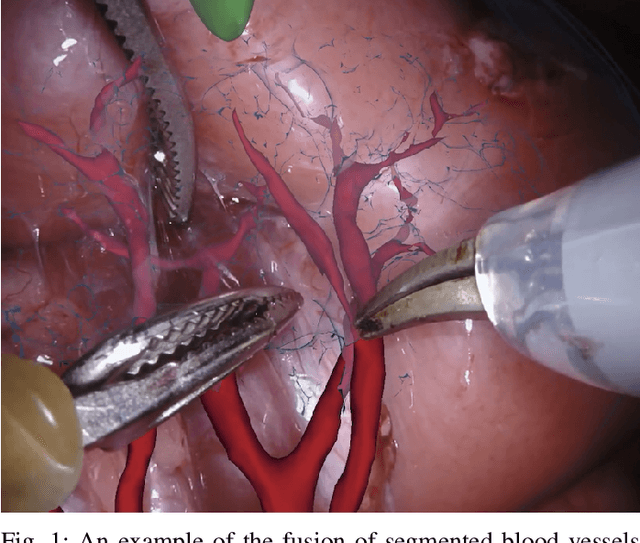
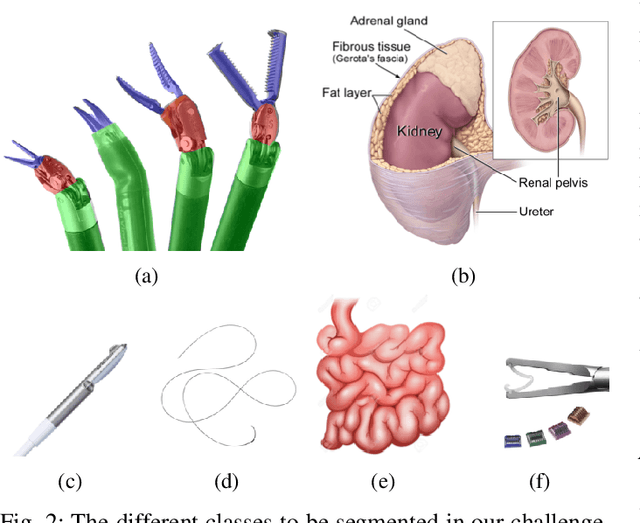
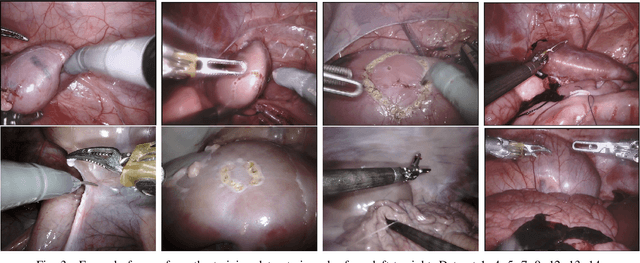
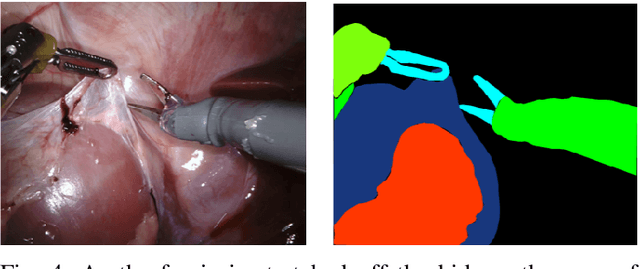
Abstract:In 2015 we began a sub-challenge at the EndoVis workshop at MICCAI in Munich using endoscope images of ex-vivo tissue with automatically generated annotations from robot forward kinematics and instrument CAD models. However, the limited background variation and simple motion rendered the dataset uninformative in learning about which techniques would be suitable for segmentation in real surgery. In 2017, at the same workshop in Quebec we introduced the robotic instrument segmentation dataset with 10 teams participating in the challenge to perform binary, articulating parts and type segmentation of da Vinci instruments. This challenge included realistic instrument motion and more complex porcine tissue as background and was widely addressed with modifications on U-Nets and other popular CNN architectures. In 2018 we added to the complexity by introducing a set of anatomical objects and medical devices to the segmented classes. To avoid over-complicating the challenge, we continued with porcine data which is dramatically simpler than human tissue due to the lack of fatty tissue occluding many organs.
Jointly Discriminative and Generative Recurrent Neural Networks for Learning from fMRI
Oct 15, 2019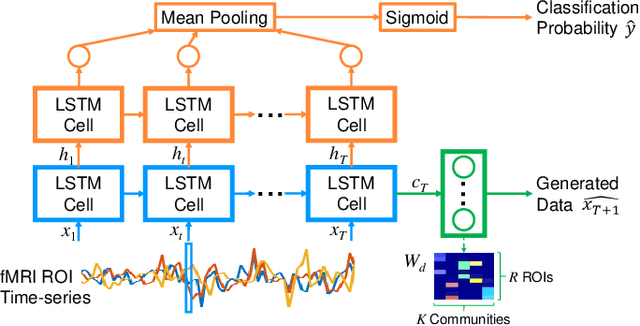
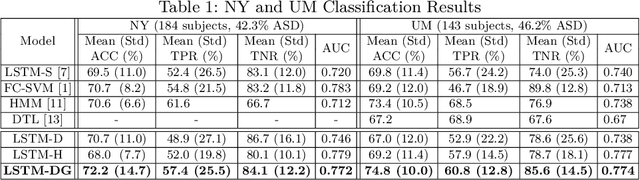
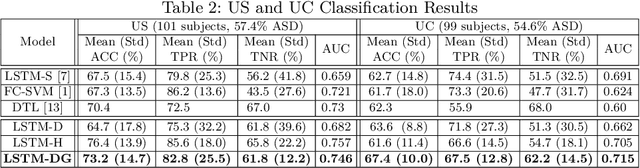
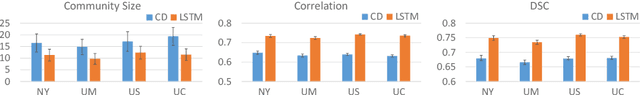
Abstract:Recurrent neural networks (RNNs) were designed for dealing with time-series data and have recently been used for creating predictive models from functional magnetic resonance imaging (fMRI) data. However, gathering large fMRI datasets for learning is a difficult task. Furthermore, network interpretability is unclear. To address these issues, we utilize multitask learning and design a novel RNN-based model that learns to discriminate between classes while simultaneously learning to generate the fMRI time-series data. Employing the long short-term memory (LSTM) structure, we develop a discriminative model based on the hidden state and a generative model based on the cell state. The addition of the generative model constrains the network to learn functional communities represented by the LSTM nodes that are both consistent with the data generation as well as useful for the classification task. We apply our approach to the classification of subjects with autism vs. healthy controls using several datasets from the Autism Brain Imaging Data Exchange. Experiments show that our jointly discriminative and generative model improves classification learning while also producing robust and meaningful functional communities for better model understanding.
Decision Explanation and Feature Importance for Invertible Networks
Oct 15, 2019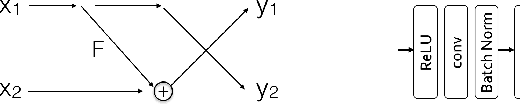

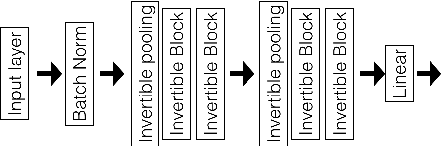

Abstract:Deep neural networks are vulnerable to adversarial attacks and hard to interpret because of their black-box nature. The recently proposed invertible network is able to accurately reconstruct the inputs to a layer from its outputs, thus has the potential to unravel the black-box model. An invertible network classifier can be viewed as a two-stage model: (1) invertible transformation from input space to the feature space; (2) a linear classifier in the feature space. We can determine the decision boundary of a linear classifier in the feature space; since the transform is invertible, we can invert the decision boundary from the feature space to the input space. Furthermore, we propose to determine the projection of a data point onto the decision boundary, and define explanation as the difference between data and its projection. Finally, we propose to locally approximate a neural network with its first-order Taylor expansion, and define feature importance using a local linear model. We provide the implementation of our method: \url{https://github.com/juntang-zhuang/explain_invertible}.
* Correct notations
Domain-Agnostic Learning with Anatomy-Consistent Embedding for Cross-Modality Liver Segmentation
Aug 27, 2019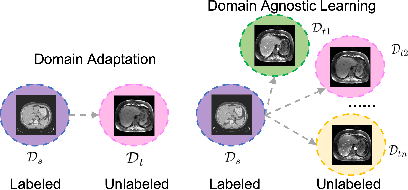
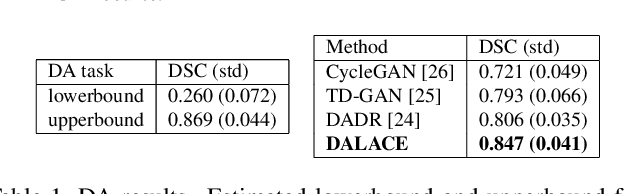
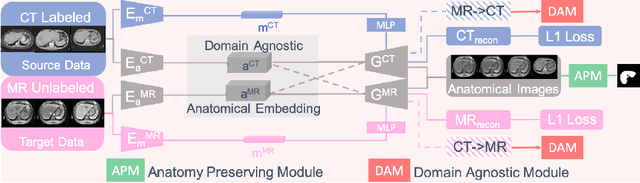
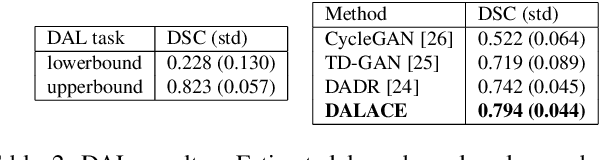
Abstract:Domain Adaptation (DA) has the potential to greatly help the generalization of deep learning models. However, the current literature usually assumes to transfer the knowledge from the source domain to a specific known target domain. Domain Agnostic Learning (DAL) proposes a new task of transferring knowledge from the source domain to data from multiple heterogeneous target domains. In this work, we propose the Domain-Agnostic Learning framework with Anatomy-Consistent Embedding (DALACE) that works on both domain-transfer and task-transfer to learn a disentangled representation, aiming to not only be invariant to different modalities but also preserve anatomical structures for the DA and DAL tasks in cross-modality liver segmentation. We validated and compared our model with state-of-the-art methods, including CycleGAN, Task Driven Generative Adversarial Network (TD-GAN), and Domain Adaptation via Disentangled Representations (DADR). For the DA task, our DALACE model outperformed CycleGAN, TD-GAN ,and DADR with DSC of 0.847 compared to 0.721, 0.793 and 0.806. For the DAL task, our model improved the performance with DSC of 0.794 from 0.522, 0.719 and 0.742 by CycleGAN, TD-GAN, and DADR. Further, we visualized the success of disentanglement, which added human interpretability of the learned meaningful representations. Through ablation analysis, we specifically showed the concrete benefits of disentanglement for downstream tasks and the role of supervision for better disentangled representation with segmentation consistency to be invariant to domains with the proposed Domain-Agnostic Module (DAM) and to preserve anatomical information with the proposed Anatomy-Preserving Module (APM).
Graph Embedding Using Infomax for ASD Classification and Brain Functional Difference Detection
Aug 14, 2019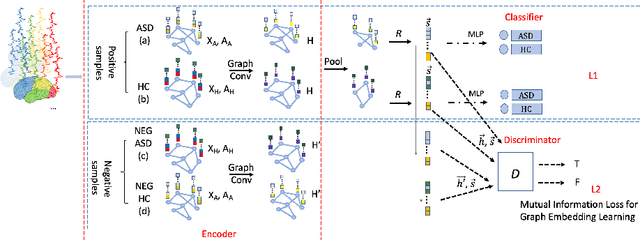

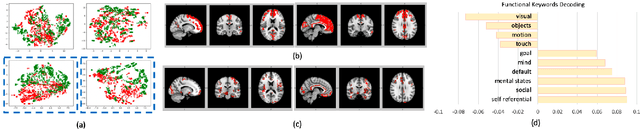
Abstract:Significant progress has been made using fMRI to characterize the brain changes that occur in ASD, a complex neuro-developmental disorder. However, due to the high dimensionality and low signal-to-noise ratio of fMRI, embedding informative and robust brain regional fMRI representations for both graph-level classification and region-level functional difference detection tasks between ASD and healthy control (HC) groups is difficult. Here, we model the whole brain fMRI as a graph, which preserves geometrical and temporal information and use a Graph Neural Network (GNN) to learn from the graph-structured fMRI data. We investigate the potential of including mutual information (MI) loss (Infomax), which is an unsupervised term encouraging large MI of each nodal representation and its corresponding graph-level summarized representation to learn a better graph embedding. Specifically, this work developed a pipeline including a GNN encoder, a classifier and a discriminator, which forces the encoded nodal representations to both benefit classification and reveal the common nodal patterns in a graph. We simultaneously optimize graph-level classification loss and Infomax. We demonstrated that Infomax graph embedding improves classification performance as a regularization term. Furthermore, we found separable nodal representations of ASD and HC groups in prefrontal cortex, cingulate cortex, visual regions, and other social, emotional and execution related brain regions. In contrast with GNN with classification loss only, the proposed pipeline can facilitate training more robust ASD classification models. Moreover, the separable nodal representations can detect the functional differences between the two groups and contribute to revealing new ASD biomarkers.
Invertible Network for Classification and Biomarker Selection for ASD
Jul 23, 2019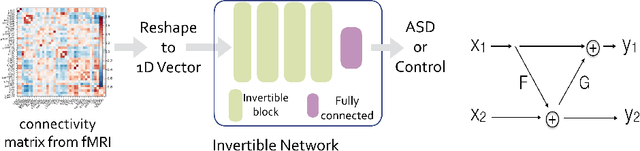
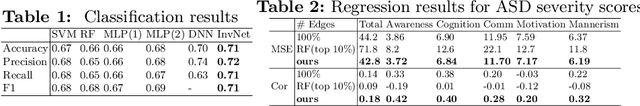
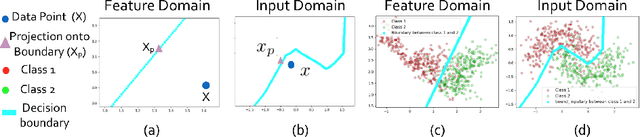
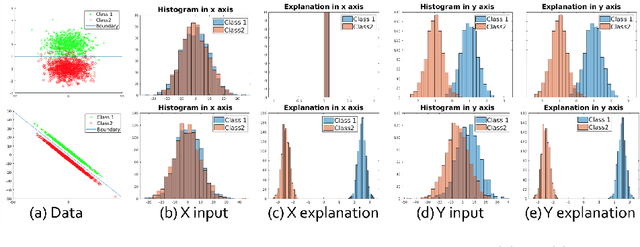
Abstract:Determining biomarkers for autism spectrum disorder (ASD) is crucial to understanding its mechanisms. Recently deep learning methods have achieved success in the classification task of ASD using fMRI data. However, due to the black-box nature of most deep learning models, it's hard to perform biomarker selection and interpret model decisions. The recently proposed invertible networks can accurately reconstruct the input from its output, and have the potential to unravel the black-box representation. Therefore, we propose a novel method to classify ASD and identify biomarkers for ASD using the connectivity matrix calculated from fMRI as the input. Specifically, with invertible networks, we explicitly determine the decision boundary and the projection of data points onto the boundary. Like linear classifiers, the difference between a point and its projection onto the decision boundary can be viewed as the explanation. We then define the importance as the explanation weighted by the gradient of prediction $w.r.t$ the input, and identify biomarkers based on this importance measure. We perform a regression task to further validate our biomarker selection: compared to using all edges in the connectivity matrix, using the top 10\% important edges we generate a lower regression error on 6 different severity scores. Our experiments show that the invertible network is both effective at ASD classification and interpretable, allowing for discovery of reliable biomarkers.
Graph Neural Network for Interpreting Task-fMRI Biomarkers
Jul 12, 2019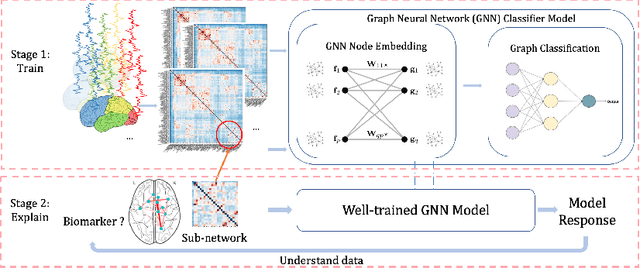
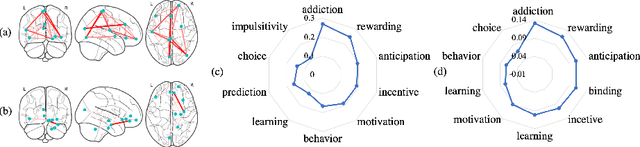
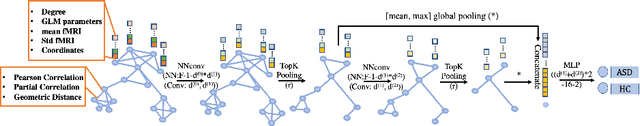
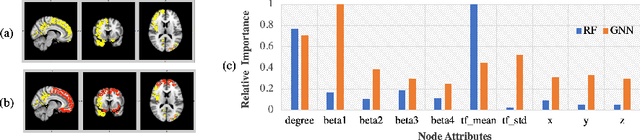

Abstract:Finding the biomarkers associated with ASD is helpful for understanding the underlying roots of the disorder and can lead to earlier diagnosis and more targeted treatment. A promising approach to identify biomarkers is using Graph Neural Networks (GNNs), which can be used to analyze graph structured data, i.e. brain networks constructed by fMRI. One way to interpret important features is through looking at how the classification probability changes if the features are occluded or replaced. The major limitation of this approach is that replacing values may change the distribution of the data and lead to serious errors. Therefore, we develop a 2-stage pipeline to eliminate the need to replace features for reliable biomarker interpretation. Specifically, we propose an inductive GNN to embed the graphs containing different properties of task-fMRI for identifying ASD and then discover the brain regions/sub-graphs used as evidence for the GNN classifier. We first show GNN can achieve high accuracy in identifying ASD. Next, we calculate the feature importance scores using GNN and compare the interpretation ability with Random Forest. Finally, we run with different atlases and parameters, proving the robustness of the proposed method. The detected biomarkers reveal their association with social behaviors. We also show the potential of discovering new informative biomarkers. Our pipeline can be generalized to other graph feature importance interpretation problems.
Efficient Interpretation of Deep Learning Models Using Graph Structure and Cooperative Game Theory: Application to ASD Biomarker Discovery
Dec 14, 2018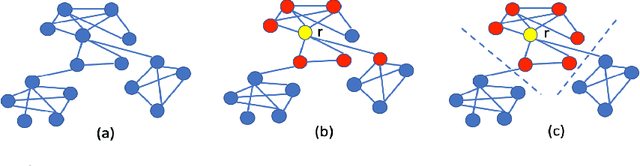
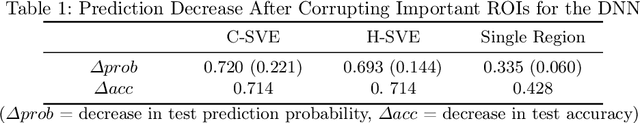
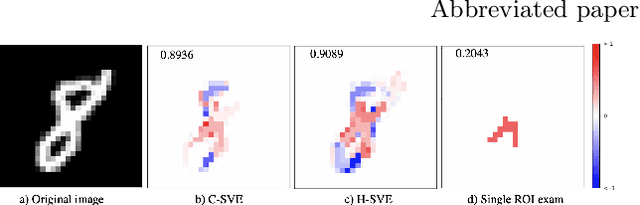
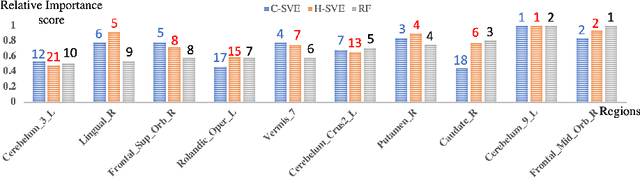
Abstract:Discovering imaging biomarkers for autism spectrum disorder (ASD) is critical to help explain ASD and predict or monitor treatment outcomes. Toward this end, deep learning classifiers have recently been used for identifying ASD from functional magnetic resonance imaging (fMRI) with higher accuracy than traditional learning strategies. However, a key challenge with deep learning models is understanding just what image features the network is using, which can in turn be used to define the biomarkers. Current methods extract biomarkers, i.e., important features, by looking at how the prediction changes if "ignoring" one feature at a time. In this work, we go beyond looking at only individual features by using Shapley value explanation (SVE) from cooperative game theory. Cooperative game theory is advantageous here because it directly considers the interaction between features and can be applied to any machine learning method, making it a novel, more accurate way of determining instance-wise biomarker importance from deep learning models. A barrier to using SVE is its computational complexity: $2^N$ given $N$ features. We explicitly reduce the complexity of SVE computation by two approaches based on the underlying graph structure of the input data: 1) only consider the centralized coalition of each feature; 2) a hierarchical pipeline which first clusters features into small communities, then applies SVE in each community. Monte Carlo approximation can be used for large permutation sets. We first validate our methods on the MNIST dataset and compare to human perception. Next, to insure plausibility of our biomarker results, we train a Random Forest (RF) to classify ASD/control subjects from fMRI and compare SVE results to standard RF-based feature importance. Finally, we show initial results on ranked fMRI biomarkers using SVE on a deep learning classifier for the ASD/control dataset.
Multi-path segmentation network
Dec 13, 2018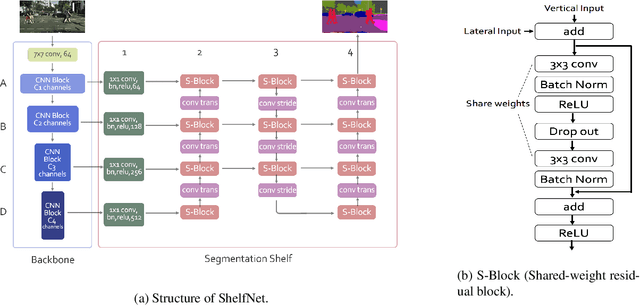
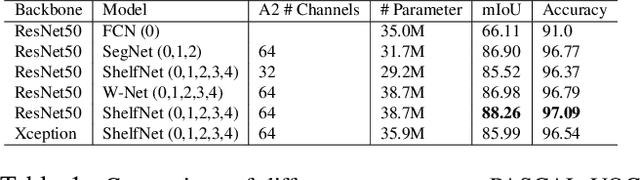
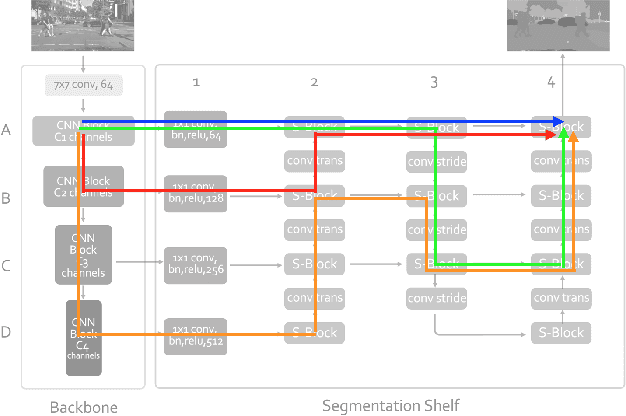
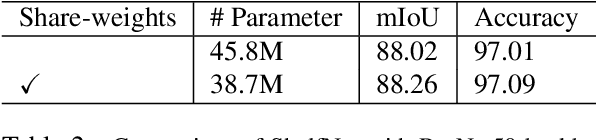
Abstract:We present a new type of convolutional network for semantic segmentation here. We tested it on several benchmark datasets, including PASCAL VOC, PASCAL Context and Cityscapes. It achieved superior performance compared to state-of-the-art segmentation methods. To increase segmentation accuracy, we design a special structure with multiple columns. The special structure creates much more paths for information flow. Therefore, it has the potential for more accurate segmentation. We propose the idea of multi-path design here, and hope it can help inspire new ideas.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge