Islem Rekik
Reservoir-Based Graph Convolutional Networks
Mar 25, 2026Abstract:Message passing is a core mechanism in Graph Neural Networks (GNNs), enabling the iterative update of node embeddings by aggregating information from neighboring nodes. Graph Convolutional Networks (GCNs) exemplify this approach by adapting convolutional operations for graph structures, allowing features from adjacent nodes to be combined effectively. However, GCNs encounter challenges with complex or dynamic data. Capturing long-range dependencies often requires deeper layers, which not only increase computational costs but also lead to over-smoothing, where node embeddings become indistinguishable. To overcome these challenges, reservoir computing has been integrated into GNNs, leveraging iterative message-passing dynamics for stable information propagation without extensive parameter tuning. Despite its promise, existing reservoir-based models lack structured convolutional mechanisms, limiting their ability to accurately aggregate multi-hop neighborhood information. To address these limitations, we propose RGC-Net (Reservoir-based Graph Convolutional Network), which integrates reservoir dynamics with structured graph convolution. Key contributions include: (i) a reimagined convolutional framework with fixed random reservoir weights and a leaky integrator to enhance feature retention; (ii) a robust, adaptable model for graph classification; and (iii) an RGC-Net-powered transformer for graph generation with application to dynamic brain connectivity. Extensive experiments show that RGC-Net achieves state-of-the-art performance in classification and generative tasks, including brain graph evolution, with faster convergence and reduced over-smoothing. Source code is available at https://github.com/basiralab/RGC-Net .
HGNet: Scalable Foundation Model for Automated Knowledge Graph Generation from Scientific Literature
Mar 24, 2026Abstract:Automated knowledge graph (KG) construction is essential for navigating the rapidly expanding body of scientific literature. However, existing approaches struggle to recognize long multi-word entities, often fail to generalize across domains, and typically overlook the hierarchical nature of scientific knowledge. While general-purpose large language models (LLMs) offer adaptability, they are computationally expensive and yield inconsistent accuracy on specialized tasks. As a result, current KGs are shallow and inconsistent, limiting their utility for exploration and synthesis. We propose a two-stage framework for scalable, zero-shot scientific KG construction. The first stage, Z-NERD, introduces (i) Orthogonal Semantic Decomposition (OSD), which promotes domain-agnostic entity recognition by isolating semantic "turns" in text, and (ii) a Multi-Scale TCQK attention mechanism that captures coherent multi-word entities through n-gram-aware attention heads. The second stage, HGNet, performs relation extraction with hierarchy-aware message passing, explicitly modeling parent, child, and peer relations. To enforce global consistency, we introduce two complementary objectives: a Differentiable Hierarchy Loss to discourage cycles and shortcut edges, and a Continuum Abstraction Field (CAF) Loss that embeds abstraction levels along a learnable axis in Euclidean space. This is the first approach to formalize hierarchical abstraction as a continuous property within standard Euclidean embeddings, offering a simpler alternative to hyperbolic methods. We release SPHERE (https://github.com/basiralab/SPHERE), a multi-domain benchmark for hierarchical relation extraction. Our framework establishes a new state of the art on SciERC, SciER, and SPHERE, improving NER by 8.08% and RE by 5.99% on out-of-distribution tests. In zero-shot settings, gains reach 10.76% for NER and 26.2% for RE.
XAI-CLIP: ROI-Guided Perturbation Framework for Explainable Medical Image Segmentation in Multimodal Vision-Language Models
Feb 01, 2026Abstract:Medical image segmentation is a critical component of clinical workflows, enabling accurate diagnosis, treatment planning, and disease monitoring. However, despite the superior performance of transformer-based models over convolutional architectures, their limited interpretability remains a major obstacle to clinical trust and deployment. Existing explainable artificial intelligence (XAI) techniques, including gradient-based saliency methods and perturbation-based approaches, are often computationally expensive, require numerous forward passes, and frequently produce noisy or anatomically irrelevant explanations. To address these limitations, we propose XAI-CLIP, an ROI-guided perturbation framework that leverages multimodal vision-language model embeddings to localize clinically meaningful anatomical regions and guide the explanation process. By integrating language-informed region localization with medical image segmentation and applying targeted, region-aware perturbations, the proposed method generates clearer, boundary-aware saliency maps while substantially reducing computational overhead. Experiments conducted on the FLARE22 and CHAOS datasets demonstrate that XAI-CLIP achieves up to a 60\% reduction in runtime, a 44.6\% improvement in dice score, and a 96.7\% increase in Intersection-over-Union for occlusion-based explanations compared to conventional perturbation methods. Qualitative results further confirm cleaner and more anatomically consistent attribution maps with fewer artifacts, highlighting that the incorporation of multimodal vision-language representations into perturbation-based XAI frameworks significantly enhances both interpretability and efficiency, thereby enabling transparent and clinically deployable medical image segmentation systems.
Rethinking Graph Super-resolution: Dual Frameworks for Topological Fidelity
Nov 12, 2025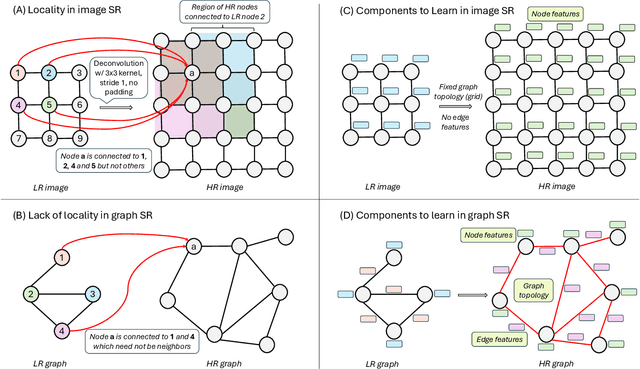
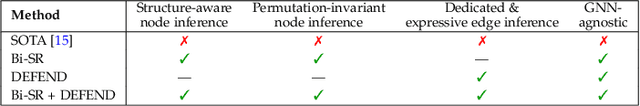
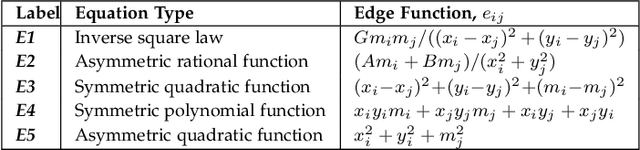

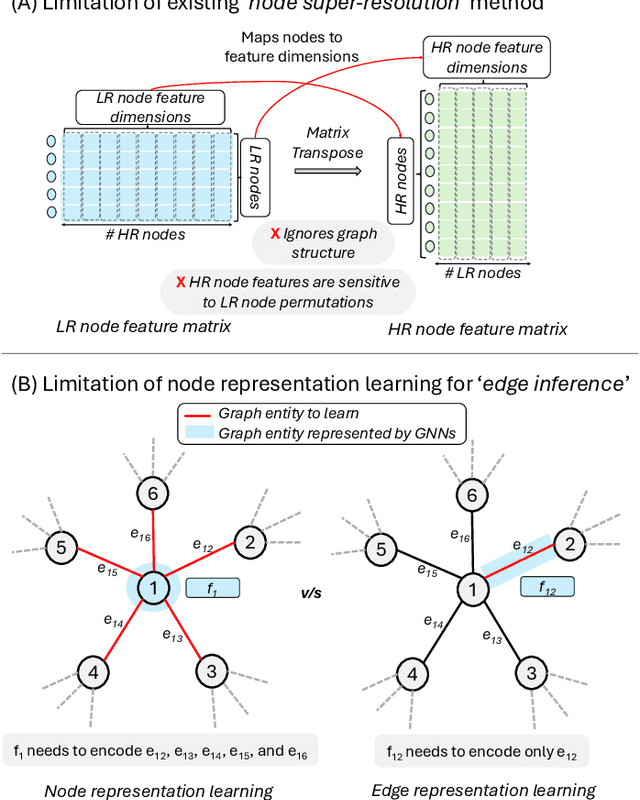

Abstract:Graph super-resolution, the task of inferring high-resolution (HR) graphs from low-resolution (LR) counterparts, is an underexplored yet crucial research direction that circumvents the need for costly data acquisition. This makes it especially desirable for resource-constrained fields such as the medical domain. While recent GNN-based approaches show promise, they suffer from two key limitations: (1) matrix-based node super-resolution that disregards graph structure and lacks permutation invariance; and (2) reliance on node representations to infer edge weights, which limits scalability and expressivity. In this work, we propose two GNN-agnostic frameworks to address these issues. First, Bi-SR introduces a bipartite graph connecting LR and HR nodes to enable structure-aware node super-resolution that preserves topology and permutation invariance. Second, DEFEND learns edge representations by mapping HR edges to nodes of a dual graph, allowing edge inference via standard node-based GNNs. We evaluate both frameworks on a real-world brain connectome dataset, where they achieve state-of-the-art performance across seven topological measures. To support generalization, we introduce twelve new simulated datasets that capture diverse topologies and LR-HR relationships. These enable comprehensive benchmarking of graph super-resolution methods.
UnifiedFL: A Dynamic Unified Learning Framework for Equitable Federation
Oct 30, 2025Abstract:Federated learning (FL) has emerged as a key paradigm for collaborative model training across multiple clients without sharing raw data, enabling privacy-preserving applications in areas such as radiology and pathology. However, works on collaborative training across clients with fundamentally different neural architectures and non-identically distributed datasets remain scarce. Existing FL frameworks face several limitations. Despite claiming to support architectural heterogeneity, most recent FL methods only tolerate variants within a single model family (e.g., shallower, deeper, or wider CNNs), still presuming a shared global architecture and failing to accommodate federations where clients deploy fundamentally different network types (e.g., CNNs, GNNs, MLPs). Moreover, existing approaches often address only statistical heterogeneity while overlooking the domain-fracture problem, where each client's data distribution differs markedly from that faced at testing time, undermining model generalizability. When clients use different architectures, have non-identically distributed data, and encounter distinct test domains, current methods perform poorly. To address these challenges, we propose UnifiedFL, a dynamic federated learning framework that represents heterogeneous local networks as nodes and edges in a directed model graph optimized by a shared graph neural network (GNN). UnifiedFL introduces (i) a common GNN to parameterize all architectures, (ii) distance-driven clustering via Euclidean distances between clients' parameters, and (iii) a two-tier aggregation policy balancing convergence and diversity. Experiments on MedMNIST classification and hippocampus segmentation benchmarks demonstrate UnifiedFL's superior performance. Code and data: https://github.com/basiralab/UnifiedFL
DeltaGNN: Graph Neural Network with Information Flow Control
Jan 10, 2025Abstract:Graph Neural Networks (GNNs) are popular deep learning models designed to process graph-structured data through recursive neighborhood aggregations in the message passing process. When applied to semi-supervised node classification, the message-passing enables GNNs to understand short-range spatial interactions, but also causes them to suffer from over-smoothing and over-squashing. These challenges hinder model expressiveness and prevent the use of deeper models to capture long-range node interactions (LRIs) within the graph. Popular solutions for LRIs detection are either too expensive to process large graphs due to high time complexity or fail to generalize across diverse graph structures. To address these limitations, we propose a mechanism called \emph{information flow control}, which leverages a novel connectivity measure, called \emph{information flow score}, to address over-smoothing and over-squashing with linear computational overhead, supported by theoretical evidence. Finally, to prove the efficacy of our methodology we design DeltaGNN, the first scalable and generalizable approach for detecting long-range and short-range interactions. We benchmark our model across 10 real-world datasets, including graphs with varying sizes, topologies, densities, and homophilic ratios, showing superior performance with limited computational complexity. The implementation of the proposed methods are publicly available at https://github.com/basiralab/DeltaGNN.
Strongly Topology-preserving GNNs for Brain Graph Super-resolution
Nov 01, 2024Abstract:Brain graph super-resolution (SR) is an under-explored yet highly relevant task in network neuroscience. It circumvents the need for costly and time-consuming medical imaging data collection, preparation, and processing. Current SR methods leverage graph neural networks (GNNs) thanks to their ability to natively handle graph-structured datasets. However, most GNNs perform node feature learning, which presents two significant limitations: (1) they require computationally expensive methods to learn complex node features capable of inferring connectivity strength or edge features, which do not scale to larger graphs; and (2) computations in the node space fail to adequately capture higher-order brain topologies such as cliques and hubs. However, numerous studies have shown that brain graph topology is crucial in identifying the onset and presence of various neurodegenerative disorders like Alzheimer and Parkinson. Motivated by these challenges and applications, we propose our STP-GSR framework. It is the first graph SR architecture to perform representation learning in higher-order topological space. Specifically, using the primal-dual graph formulation from graph theory, we develop an efficient mapping from the edge space of our low-resolution (LR) brain graphs to the node space of a high-resolution (HR) dual graph. This approach ensures that node-level computations on this dual graph correspond naturally to edge-level learning on our HR brain graphs, thereby enforcing strong topological consistency within our framework. Additionally, our framework is GNN layer agnostic and can easily learn from smaller, scalable GNNs, reducing computational requirements. We comprehensively benchmark our framework across seven key topological measures and observe that it significantly outperforms the previous state-of-the-art methods and baselines.
Metadata-Driven Federated Learning of Connectional Brain Templates in Non-IID Multi-Domain Scenarios
Mar 14, 2024



Abstract:A connectional brain template (CBT) is a holistic representation of a population of multi-view brain connectivity graphs, encoding shared patterns and normalizing typical variations across individuals. The federation of CBT learning allows for an inclusive estimation of the representative center of multi-domain brain connectivity datasets in a fully data-preserving manner. However, existing methods overlook the non-independent and identically distributed (non-IDD) issue stemming from multidomain brain connectivity heterogeneity, in which data domains are drawn from different hospitals and imaging modalities. To overcome this limitation, we unprecedentedly propose a metadata-driven federated learning framework, called MetaFedCBT, for cross-domain CBT learning. Given the data drawn from a specific domain (i.e., hospital), our model aims to learn metadata in a fully supervised manner by introducing a local client-based regressor network. The generated meta-data is forced to meet the statistical attributes (e.g., mean) of other domains, while preserving their privacy. Our supervised meta-data generation approach boosts the unsupervised learning of a more centered, representative, and holistic CBT of a particular brain state across diverse domains. As the federated learning progresses over multiple rounds, the learned metadata and associated generated connectivities are continuously updated to better approximate the target domain information. MetaFedCBT overcomes the non-IID issue of existing methods by generating informative brain connectivities for privacy-preserving holistic CBT learning with guidance using metadata. Extensive experiments on multi-view morphological brain networks of normal and patient subjects demonstrate that our MetaFedCBT is a superior federated CBT learning model and significantly advances the state-of-the-art performance.
Predicting Infant Brain Connectivity with Federated Multi-Trajectory GNNs using Scarce Data
Jan 08, 2024



Abstract:The understanding of the convoluted evolution of infant brain networks during the first postnatal year is pivotal for identifying the dynamics of early brain connectivity development. Existing deep learning solutions suffer from three major limitations. First, they cannot generalize to multi-trajectory prediction tasks, where each graph trajectory corresponds to a particular imaging modality or connectivity type (e.g., T1-w MRI). Second, existing models require extensive training datasets to achieve satisfactory performance which are often challenging to obtain. Third, they do not efficiently utilize incomplete time series data. To address these limitations, we introduce FedGmTE-Net++, a federated graph-based multi-trajectory evolution network. Using the power of federation, we aggregate local learnings among diverse hospitals with limited datasets. As a result, we enhance the performance of each hospital's local generative model, while preserving data privacy. The three key innovations of FedGmTE-Net++ are: (i) presenting the first federated learning framework specifically designed for brain multi-trajectory evolution prediction in a data-scarce environment, (ii) incorporating an auxiliary regularizer in the local objective function to exploit all the longitudinal brain connectivity within the evolution trajectory and maximize data utilization, (iii) introducing a two-step imputation process, comprising a preliminary KNN-based precompletion followed by an imputation refinement step that employs regressors to improve similarity scores and refine imputations. Our comprehensive experimental results showed the outperformance of FedGmTE-Net++ in brain multi-trajectory prediction from a single baseline graph in comparison with benchmark methods.
Replica Tree-based Federated Learning using Limited Data
Dec 28, 2023



Abstract:Learning from limited data has been extensively studied in machine learning, considering that deep neural networks achieve optimal performance when trained using a large amount of samples. Although various strategies have been proposed for centralized training, the topic of federated learning with small datasets remains largely unexplored. Moreover, in realistic scenarios, such as settings where medical institutions are involved, the number of participating clients is also constrained. In this work, we propose a novel federated learning framework, named RepTreeFL. At the core of the solution is the concept of a replica, where we replicate each participating client by copying its model architecture and perturbing its local data distribution. Our approach enables learning from limited data and a small number of clients by aggregating a larger number of models with diverse data distributions. Furthermore, we leverage the hierarchical structure of the client network (both original and virtual), alongside the model diversity across replicas, and introduce a diversity-based tree aggregation, where replicas are combined in a tree-like manner and the aggregation weights are dynamically updated based on the model discrepancy. We evaluated our method on two tasks and two types of data, graph generation and image classification (binary and multi-class), with both homogeneous and heterogeneous model architectures. Experimental results demonstrate the effectiveness and outperformance of RepTreeFL in settings where both data and clients are limited. Our code is available at https://github.com/basiralab/RepTreeFL.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge