Jing Xiao
nnSpeech: Speaker-Guided Conditional Variational Autoencoder for Zero-shot Multi-speaker Text-to-Speech
Feb 22, 2022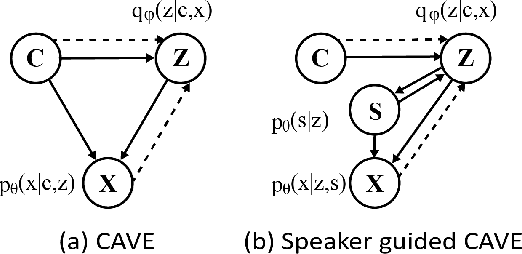
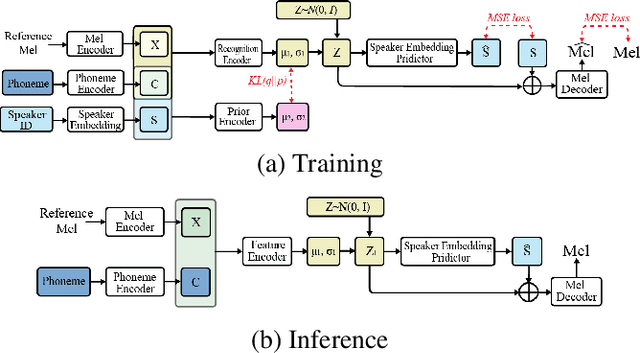
Abstract:Multi-speaker text-to-speech (TTS) using a few adaption data is a challenge in practical applications. To address that, we propose a zero-shot multi-speaker TTS, named nnSpeech, that could synthesis a new speaker voice without fine-tuning and using only one adaption utterance. Compared with using a speaker representation module to extract the characteristics of new speakers, our method bases on a speaker-guided conditional variational autoencoder and can generate a variable Z, which contains both speaker characteristics and content information. The latent variable Z distribution is approximated by another variable conditioned on reference mel-spectrogram and phoneme. Experiments on the English corpus, Mandarin corpus, and cross-dataset proves that our model could generate natural and similar speech with only one adaption speech.
r-G2P: Evaluating and Enhancing Robustness of Grapheme to Phoneme Conversion by Controlled noise introducing and Contextual information incorporation
Feb 21, 2022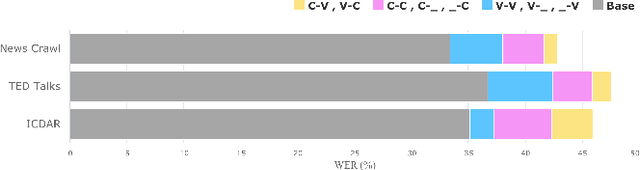
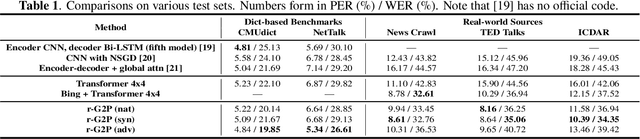
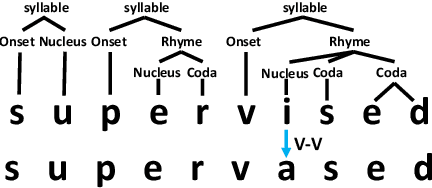
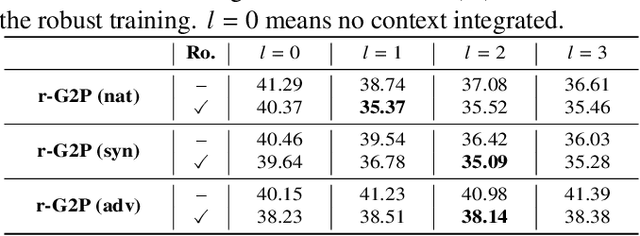
Abstract:Grapheme-to-phoneme (G2P) conversion is the process of converting the written form of words to their pronunciations. It has an important role for text-to-speech (TTS) synthesis and automatic speech recognition (ASR) systems. In this paper, we aim to evaluate and enhance the robustness of G2P models. We show that neural G2P models are extremely sensitive to orthographical variations in graphemes like spelling mistakes. To solve this problem, we propose three controlled noise introducing methods to synthesize noisy training data. Moreover, we incorporate the contextual information with the baseline and propose a robust training strategy to stabilize the training process. The experimental results demonstrate that our proposed robust G2P model (r-G2P) outperforms the baseline significantly (-2.73\% WER on Dict-based benchmarks and -9.09\% WER on Real-world sources).
AVQVC: One-shot Voice Conversion by Vector Quantization with applying contrastive learning
Feb 21, 2022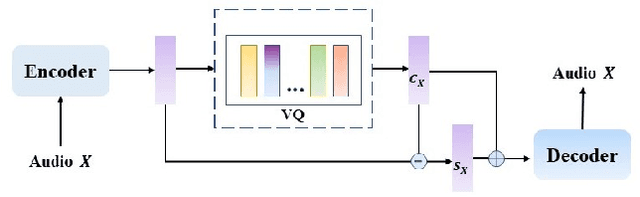
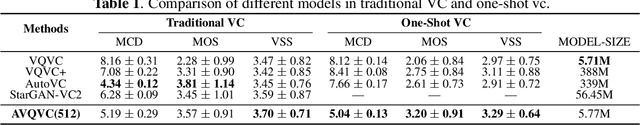
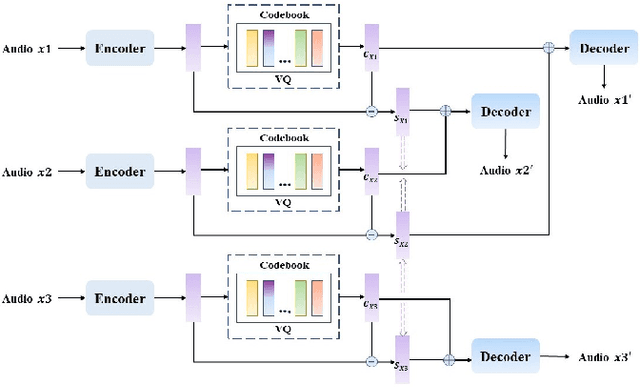
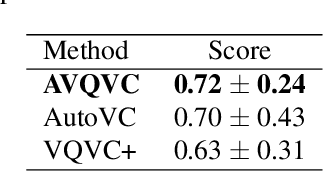
Abstract:Voice Conversion(VC) refers to changing the timbre of a speech while retaining the discourse content. Recently, many works have focused on disentangle-based learning techniques to separate the timbre and the linguistic content information from a speech signal. Once successful, voice conversion will be feasible and straightforward. This paper proposed a novel one-shot voice conversion framework based on vector quantization voice conversion (VQVC) and AutoVC, called AVQVC. A new training method is applied to VQVC to separate content and timbre information from speech more effectively. The result shows that this approach has better performance than VQVC in separating content and timbre to improve the sound quality of generated speech.
Lumbar Bone Mineral Density Estimation from Chest X-ray Images: Anatomy-aware Attentive Multi-ROI Modeling
Jan 05, 2022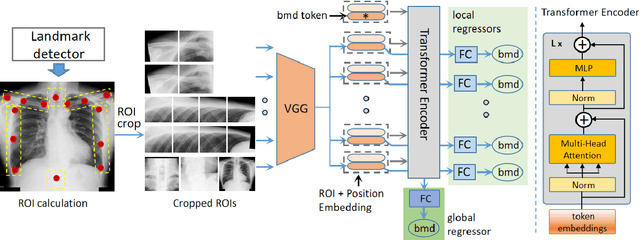
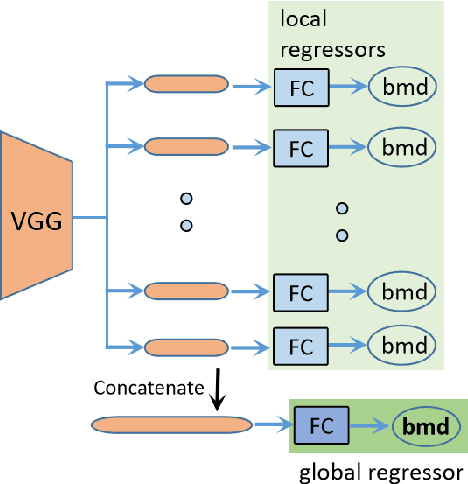
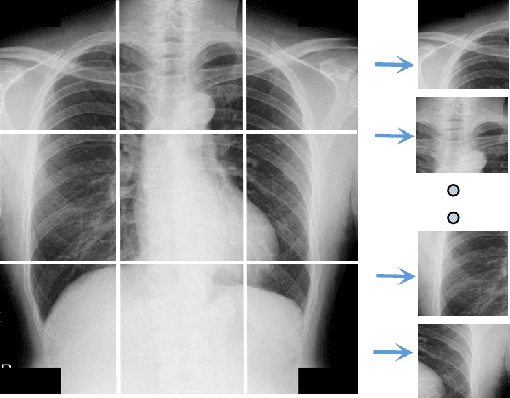
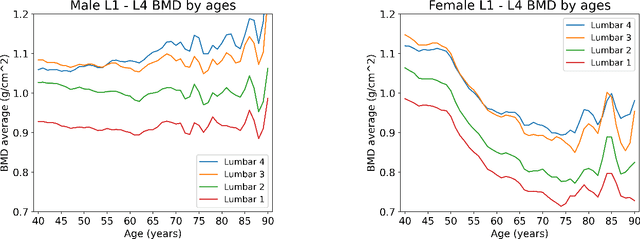
Abstract:Osteoporosis is a common chronic metabolic bone disease that is often under-diagnosed and under-treated due to the limited access to bone mineral density (BMD) examinations, e.g. via Dual-energy X-ray Absorptiometry (DXA). In this paper, we propose a method to predict BMD from Chest X-ray (CXR), one of the most commonly accessible and low-cost medical imaging examinations. Our method first automatically detects Regions of Interest (ROIs) of local and global bone structures from the CXR. Then a multi-ROI deep model with transformer encoder is developed to exploit both local and global information in the chest X-ray image for accurate BMD estimation. Our method is evaluated on 13719 CXR patient cases with their ground truth BMD scores measured by gold-standard DXA. The model predicted BMD has a strong correlation with the ground truth (Pearson correlation coefficient 0.889 on lumbar 1). When applied for osteoporosis screening, it achieves a high classification performance (AUC 0.963 on lumbar 1). As the first effort in the field using CXR scans to predict the BMD, the proposed algorithm holds strong potential in early osteoporosis screening and public health promotion.
Coherence Learning using Keypoint-based Pooling Network for Accurately Assessing Radiographic Knee Osteoarthritis
Dec 16, 2021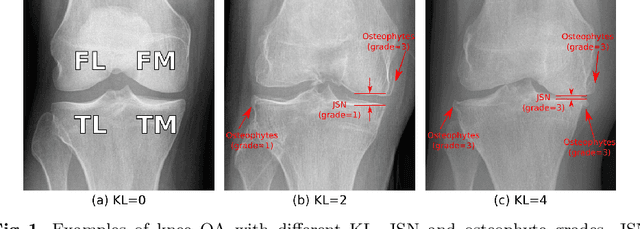
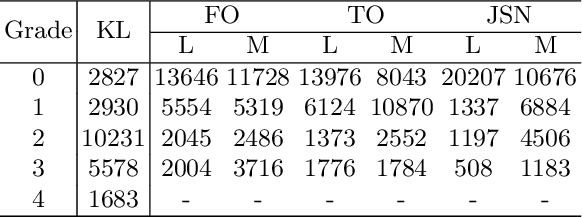
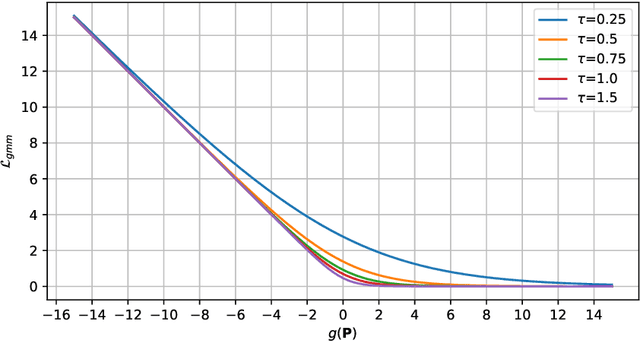
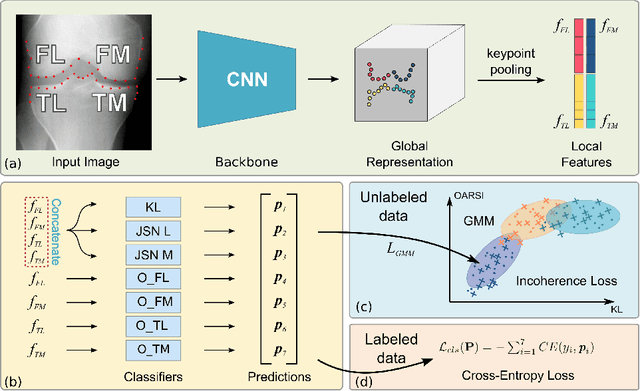
Abstract:Knee osteoarthritis (OA) is a common degenerate joint disorder that affects a large population of elderly people worldwide. Accurate radiographic assessment of knee OA severity plays a critical role in chronic patient management. Current clinically-adopted knee OA grading systems are observer subjective and suffer from inter-rater disagreements. In this work, we propose a computer-aided diagnosis approach to provide more accurate and consistent assessments of both composite and fine-grained OA grades simultaneously. A novel semi-supervised learning method is presented to exploit the underlying coherence in the composite and fine-grained OA grades by learning from unlabeled data. By representing the grade coherence using the log-probability of a pre-trained Gaussian Mixture Model, we formulate an incoherence loss to incorporate unlabeled data in training. The proposed method also describes a keypoint-based pooling network, where deep image features are pooled from the disease-targeted keypoints (extracted along the knee joint) to provide more aligned and pathologically informative feature representations, for accurate OA grade assessments. The proposed method is comprehensively evaluated on the public Osteoarthritis Initiative (OAI) data, a multi-center ten-year observational study on 4,796 subjects. Experimental results demonstrate that our method leads to significant improvements over previous strong whole image-based deep classification network baselines (like ResNet-50).
Comprehensive and Clinically Accurate Head and Neck Organs at Risk Delineation via Stratified Deep Learning: A Large-scale Multi-Institutional Study
Nov 01, 2021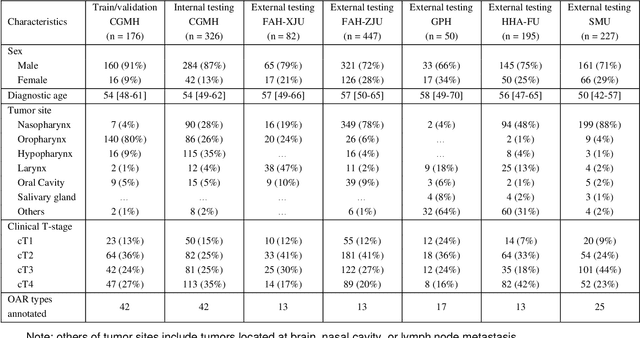
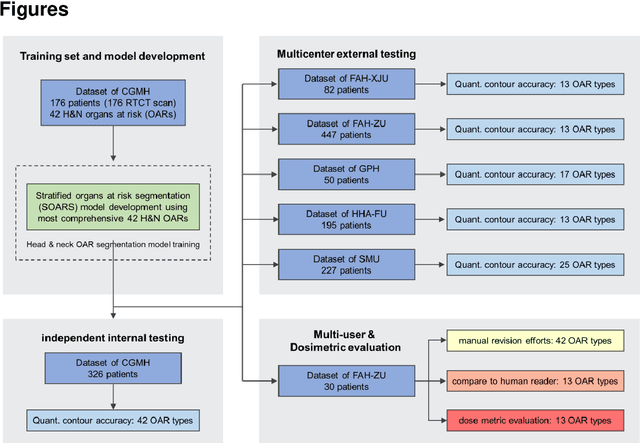
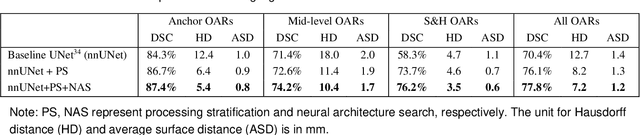
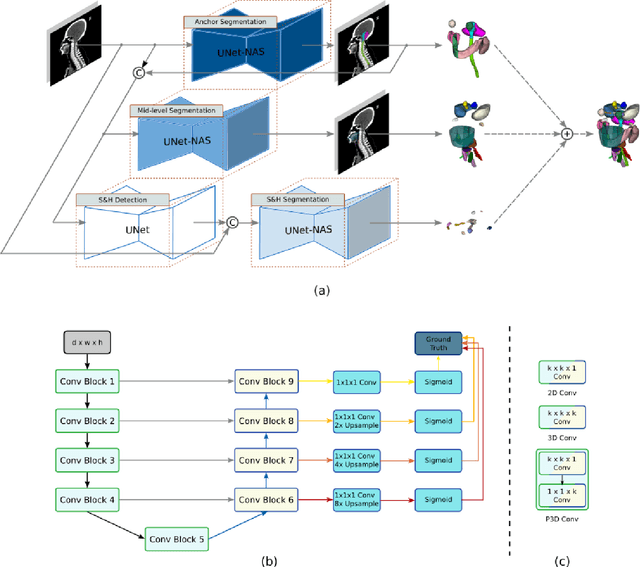
Abstract:Accurate organ at risk (OAR) segmentation is critical to reduce the radiotherapy post-treatment complications. Consensus guidelines recommend a set of more than 40 OARs in the head and neck (H&N) region, however, due to the predictable prohibitive labor-cost of this task, most institutions choose a substantially simplified protocol by delineating a smaller subset of OARs and neglecting the dose distributions associated with other OARs. In this work we propose a novel, automated and highly effective stratified OAR segmentation (SOARS) system using deep learning to precisely delineate a comprehensive set of 42 H&N OARs. SOARS stratifies 42 OARs into anchor, mid-level, and small & hard subcategories, with specifically derived neural network architectures for each category by neural architecture search (NAS) principles. We built SOARS models using 176 training patients in an internal institution and independently evaluated on 1327 external patients across six different institutions. It consistently outperformed other state-of-the-art methods by at least 3-5% in Dice score for each institutional evaluation (up to 36% relative error reduction in other metrics). More importantly, extensive multi-user studies evidently demonstrated that 98% of the SOARS predictions need only very minor or no revisions for direct clinical acceptance (saving 90% radiation oncologists workload), and their segmentation and dosimetric accuracy are within or smaller than the inter-user variation. These findings confirmed the strong clinical applicability of SOARS for the OAR delineation process in H&N cancer radiotherapy workflows, with improved efficiency, comprehensiveness, and quality.
A deep learning pipeline for localization, differentiation, and uncertainty estimation of liver lesions using multi-phasic and multi-sequence MRI
Oct 17, 2021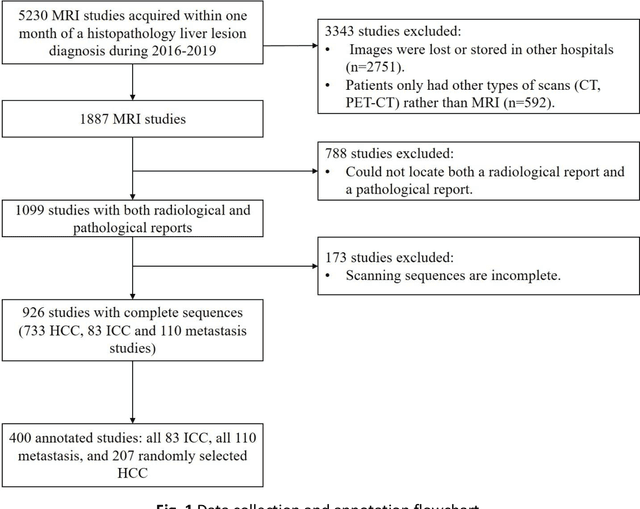
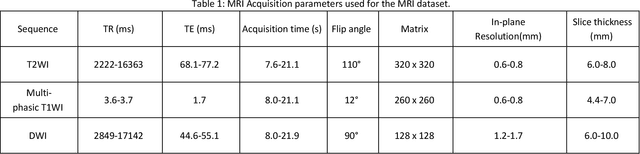
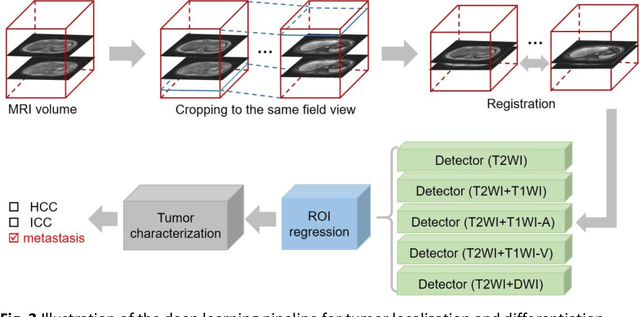
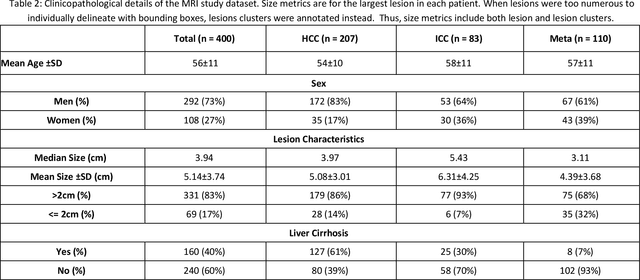
Abstract:Objectives: to propose a fully-automatic computer-aided diagnosis (CAD) solution for liver lesion characterization, with uncertainty estimation. Methods: we enrolled 400 patients who had either liver resection or a biopsy and was diagnosed with either hepatocellular carcinoma (HCC), intrahepatic cholangiocarcinoma, or secondary metastasis, from 2006 to 2019. Each patient was scanned with T1WI, T2WI, T1WI venous phase (T2WI-V), T1WI arterial phase (T1WI-A), and DWI MRI sequences. We propose a fully-automatic deep CAD pipeline that localizes lesions from 3D MRI studies using key-slice parsing and provides a confidence measure for its diagnoses. We evaluate using five-fold cross validation and compare performance against three radiologists, including a senior hepatology radiologist, a junior hepatology radiologist and an abdominal radiologist. Results: the proposed CAD solution achieves a mean F1 score of 0.62, outperforming the abdominal radiologist (0.47), matching the junior hepatology radiologist (0.61), and underperforming the senior hepatology radiologist (0.68). The CAD system can informatively assess its diagnostic confidence, i.e., when only evaluating on the 70% most confident cases the mean f1 score and sensitivity at 80% specificity for HCC vs. others are boosted from 0.62 to 0.71 and 0.84 to 0.92, respectively. Conclusion: the proposed fully-automatic CAD solution can provide good diagnostic performance with informative confidence assessments in finding and discriminating liver lesions from MRI studies.
Accurate and Generalizable Quantitative Scoring of Liver Steatosis from Ultrasound Images via Scalable Deep Learning
Oct 12, 2021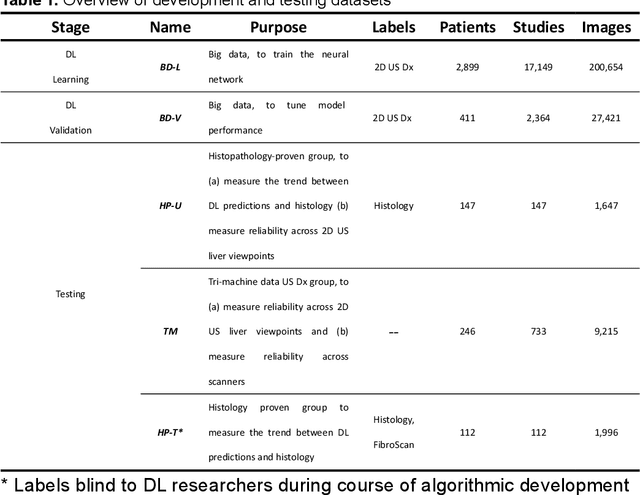
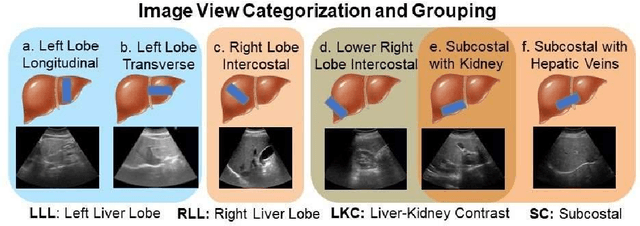
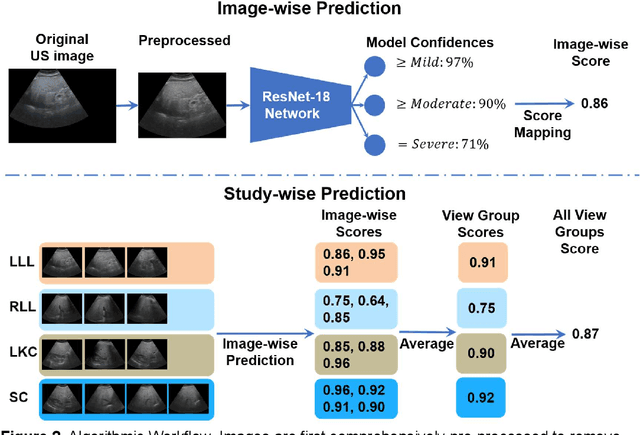
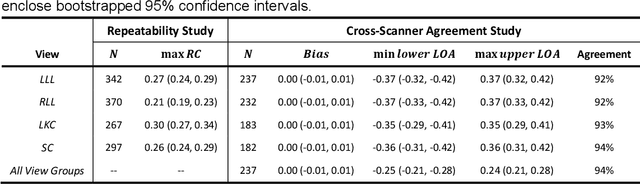
Abstract:Background & Aims: Hepatic steatosis is a major cause of chronic liver disease. 2D ultrasound is the most widely used non-invasive tool for screening and monitoring, but associated diagnoses are highly subjective. We developed a scalable deep learning (DL) algorithm for quantitative scoring of liver steatosis from 2D ultrasound images. Approach & Results: Using retrospectively collected multi-view ultrasound data from 3,310 patients, 19,513 studies, and 228,075 images, we trained a DL algorithm to diagnose steatosis stages (healthy, mild, moderate, or severe) from ultrasound diagnoses. Performance was validated on two multi-scanner unblinded and blinded (initially to DL developer) histology-proven cohorts (147 and 112 patients) with histopathology fatty cell percentage diagnoses, and a subset with FibroScan diagnoses. We also quantified reliability across scanners and viewpoints. Results were evaluated using Bland-Altman and receiver operating characteristic (ROC) analysis. The DL algorithm demonstrates repeatable measurements with a moderate number of images (3 for each viewpoint) and high agreement across 3 premium ultrasound scanners. High diagnostic performance was observed across all viewpoints: area under the curves of the ROC to classify >=mild, >=moderate, =severe steatosis grades were 0.85, 0.90, and 0.93, respectively. The DL algorithm outperformed or performed at least comparably to FibroScan with statistically significant improvements for all levels on the unblinded histology-proven cohort, and for =severe steatosis on the blinded histology-proven cohort. Conclusions: The DL algorithm provides a reliable quantitative steatosis assessment across view and scanners on two multi-scanner cohorts. Diagnostic performance was high with comparable or better performance than FibroScan.
Multi-institutional Validation of Two-Streamed Deep Learning Method for Automated Delineation of Esophageal Gross Tumor Volume using planning-CT and FDG-PETCT
Oct 11, 2021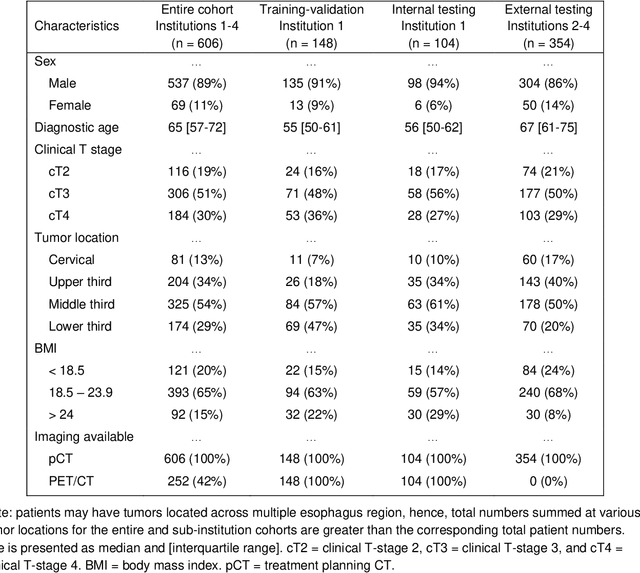
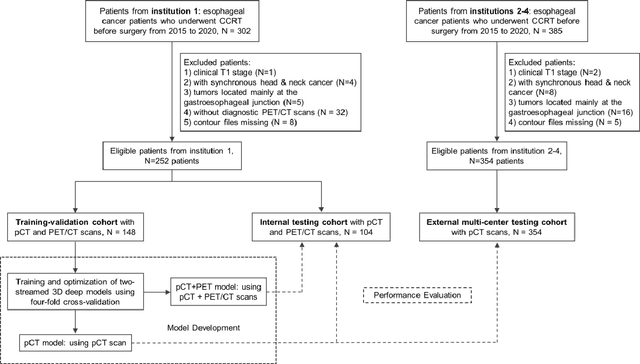
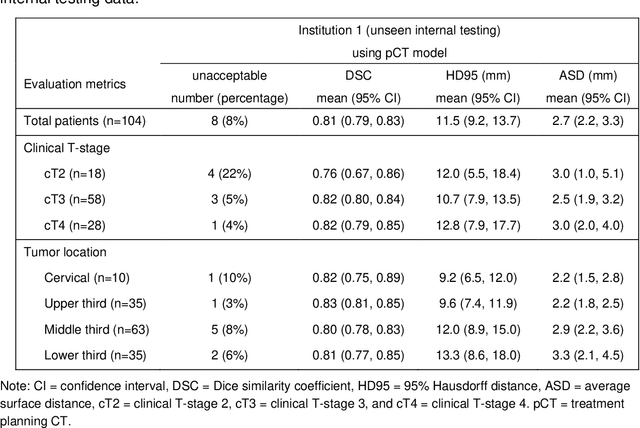
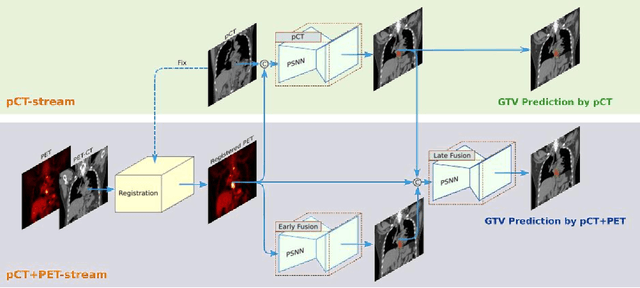
Abstract:Background: The current clinical workflow for esophageal gross tumor volume (GTV) contouring relies on manual delineation of high labor-costs and interuser variability. Purpose: To validate the clinical applicability of a deep learning (DL) multi-modality esophageal GTV contouring model, developed at 1 institution whereas tested at multiple ones. Methods and Materials: We collected 606 esophageal cancer patients from four institutions. 252 institution-1 patients had a treatment planning-CT (pCT) and a pair of diagnostic FDG-PETCT; 354 patients from other 3 institutions had only pCT. A two-streamed DL model for GTV segmentation was developed using pCT and PETCT scans of a 148 patient institution-1 subset. This built model had the flexibility of segmenting GTVs via only pCT or pCT+PETCT combined. For independent evaluation, the rest 104 institution-1 patients behaved as unseen internal testing, and 354 institutions 2-4 patients were used for external testing. We evaluated manual revision degrees by human experts to assess the contour-editing effort. The performance of the deep model was compared against 4 radiation oncologists in a multiuser study with 20 random external patients. Contouring accuracy and time were recorded for the pre-and post-DL assisted delineation process. Results: Our model achieved high segmentation accuracy in internal testing (mean Dice score: 0.81 using pCT and 0.83 using pCT+PET) and generalized well to external evaluation (mean DSC: 0.80). Expert assessment showed that the predicted contours of 88% patients need only minor or no revision. In multi-user evaluation, with the assistance of a deep model, inter-observer variation and required contouring time were reduced by 37.6% and 48.0%, respectively. Conclusions: Deep learning predicted GTV contours were in close agreement with the ground truth and could be adopted clinically with mostly minor or no changes.
SAME: Deformable Image Registration based on Self-supervised Anatomical Embeddings
Sep 23, 2021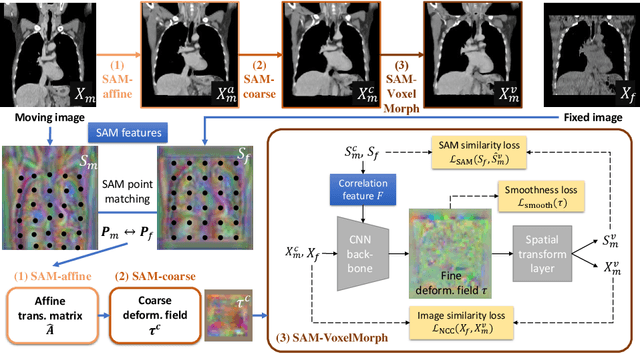
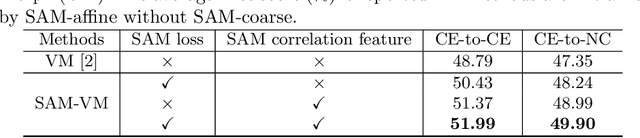
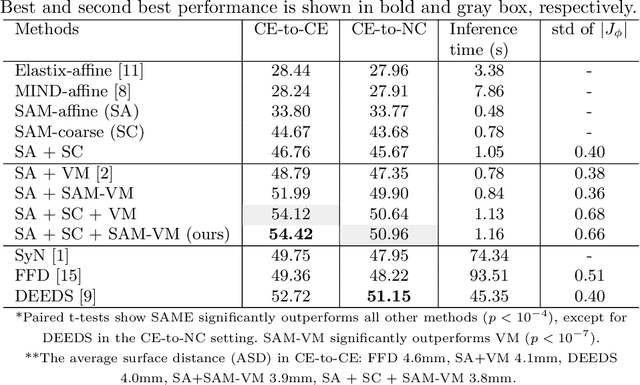

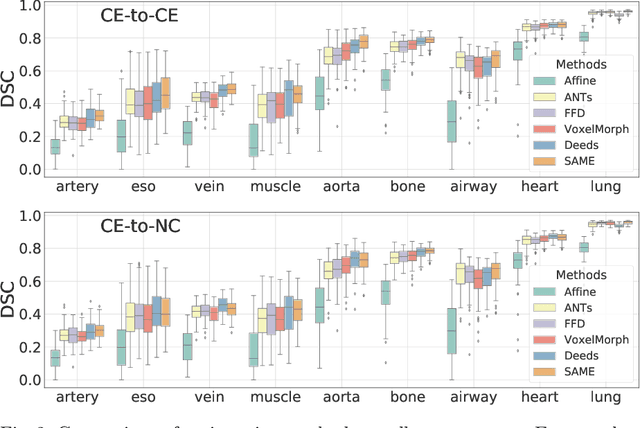
Abstract:In this work, we introduce a fast and accurate method for unsupervised 3D medical image registration. This work is built on top of a recent algorithm SAM, which is capable of computing dense anatomical/semantic correspondences between two images at the pixel level. Our method is named SAME, which breaks down image registration into three steps: affine transformation, coarse deformation, and deep deformable registration. Using SAM embeddings, we enhance these steps by finding more coherent correspondences, and providing features and a loss function with better semantic guidance. We collect a multi-phase chest computed tomography dataset with 35 annotated organs for each patient and conduct inter-subject registration for quantitative evaluation. Results show that SAME outperforms widely-used traditional registration techniques (Elastix FFD, ANTs SyN) and learning based VoxelMorph method by at least 4.7% and 2.7% in Dice scores for two separate tasks of within-contrast-phase and across-contrast-phase registration, respectively. SAME achieves the comparable performance to the best traditional registration method, DEEDS (from our evaluation), while being orders of magnitude faster (from 45 seconds to 1.2 seconds).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge