Donglai Wei
An Out-of-Domain Synapse Detection Challenge for Microwasp Brain Connectomes
Feb 01, 2023



Abstract:The size of image stacks in connectomics studies now reaches the terabyte and often petabyte scales with a great diversity of appearance across brain regions and samples. However, manual annotation of neural structures, e.g., synapses, is time-consuming, which leads to limited training data often smaller than 0.001\% of the test data in size. Domain adaptation and generalization approaches were proposed to address similar issues for natural images, which were less evaluated on connectomics data due to a lack of out-of-domain benchmarks.
QuantArt: Quantizing Image Style Transfer Towards High Visual Fidelity
Dec 20, 2022Abstract:The mechanism of existing style transfer algorithms is by minimizing a hybrid loss function to push the generated image toward high similarities in both content and style. However, this type of approach cannot guarantee visual fidelity, i.e., the generated artworks should be indistinguishable from real ones. In this paper, we devise a new style transfer framework called QuantArt for high visual-fidelity stylization. QuantArt pushes the latent representation of the generated artwork toward the centroids of the real artwork distribution with vector quantization. By fusing the quantized and continuous latent representations, QuantArt allows flexible control over the generated artworks in terms of content preservation, style similarity, and visual fidelity. Experiments on various style transfer settings show that our QuantArt framework achieves significantly higher visual fidelity compared with the existing style transfer methods.
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Abstract:The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
RibSeg v2: A Large-scale Benchmark for Rib Labeling and Anatomical Centerline Extraction
Oct 18, 2022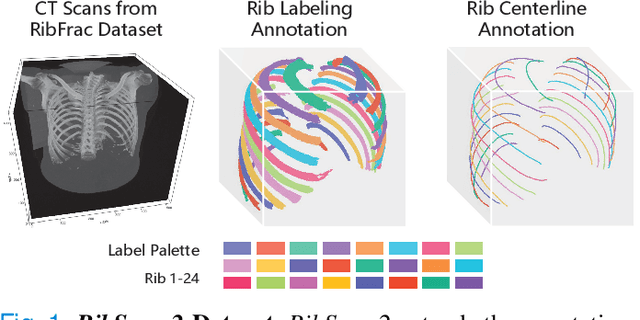
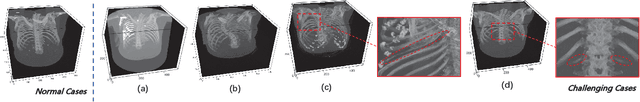
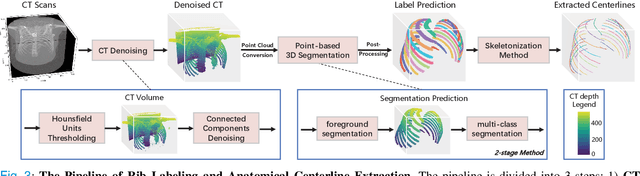
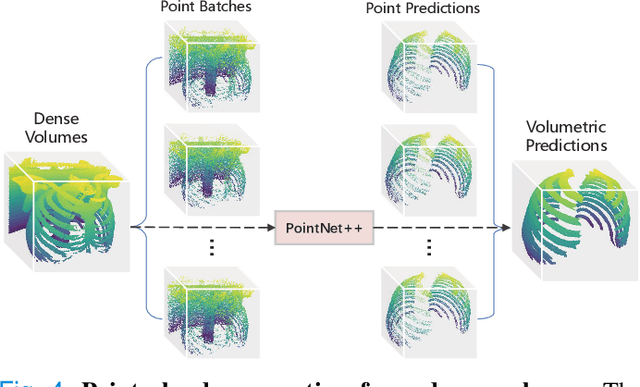
Abstract:Automatic rib labeling and anatomical centerline extraction are common prerequisites for various clinical applications. Prior studies either use in-house datasets that are inaccessible to communities, or focus on rib segmentation that neglects the clinical significance of rib labeling. To address these issues, we extend our prior dataset (RibSeg) on the binary rib segmentation task to a comprehensive benchmark, named RibSeg v2, with 660 CT scans (15,466 individual ribs in total) and annotations manually inspected by experts for rib labeling and anatomical centerline extraction. Based on the RibSeg v2, we develop a pipeline including deep learning-based methods for rib labeling, and a skeletonization-based method for centerline extraction. To improve computational efficiency, we propose a sparse point cloud representation of CT scans and compare it with standard dense voxel grids. Moreover, we design and analyze evaluation metrics to address the key challenges of each task. Our dataset, code, and model are available online to facilitate open research at https://github.com/M3DV/RibSeg
Instance Segmentation of Unlabeled Modalities via Cyclic Segmentation GAN
Apr 06, 2022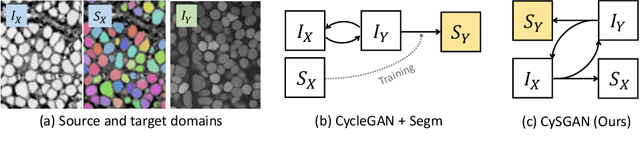
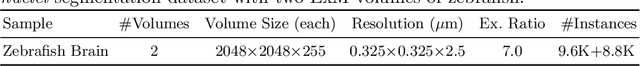
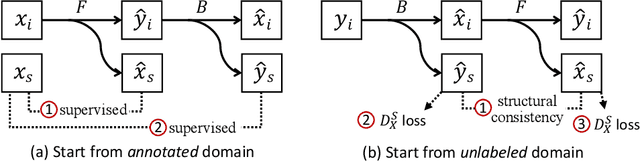
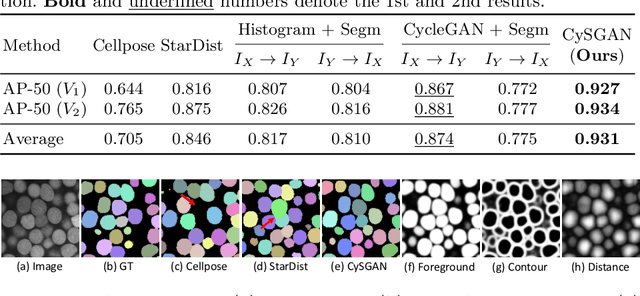
Abstract:Instance segmentation for unlabeled imaging modalities is a challenging but essential task as collecting expert annotation can be expensive and time-consuming. Existing works segment a new modality by either deploying a pre-trained model optimized on diverse training data or conducting domain translation and image segmentation as two independent steps. In this work, we propose a novel Cyclic Segmentation Generative Adversarial Network (CySGAN) that conducts image translation and instance segmentation jointly using a unified framework. Besides the CycleGAN losses for image translation and supervised losses for the annotated source domain, we introduce additional self-supervised and segmentation-based adversarial objectives to improve the model performance by leveraging unlabeled target domain images. We benchmark our approach on the task of 3D neuronal nuclei segmentation with annotated electron microscopy (EM) images and unlabeled expansion microscopy (ExM) data. Our CySGAN outperforms both pretrained generalist models and the baselines that sequentially conduct image translation and segmentation. Our implementation and the newly collected, densely annotated ExM nuclei dataset, named NucExM, are available at https://connectomics-bazaar.github.io/proj/CySGAN/index.html.
Learning to Generate Realistic Noisy Images via Pixel-level Noise-aware Adversarial Training
Apr 06, 2022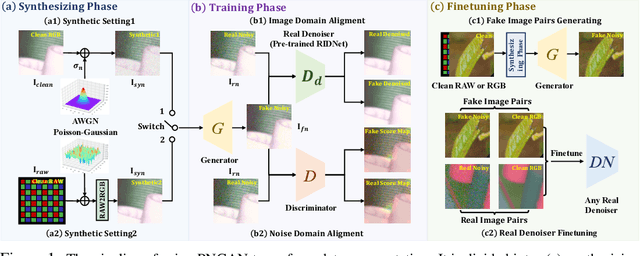
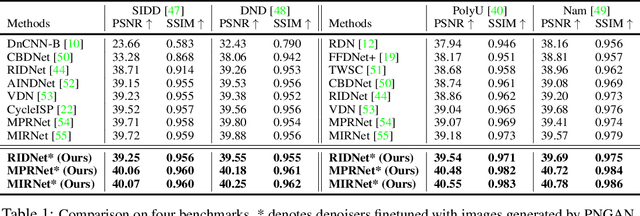
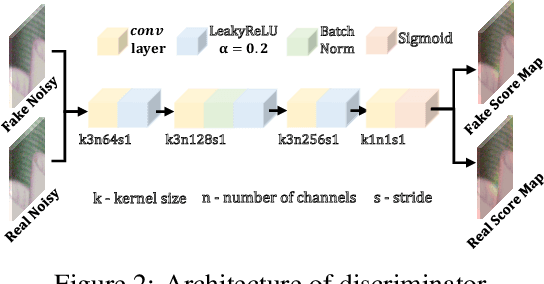
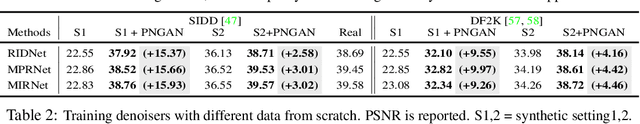
Abstract:Existing deep learning real denoising methods require a large amount of noisy-clean image pairs for supervision. Nonetheless, capturing a real noisy-clean dataset is an unacceptable expensive and cumbersome procedure. To alleviate this problem, this work investigates how to generate realistic noisy images. Firstly, we formulate a simple yet reasonable noise model that treats each real noisy pixel as a random variable. This model splits the noisy image generation problem into two sub-problems: image domain alignment and noise domain alignment. Subsequently, we propose a novel framework, namely Pixel-level Noise-aware Generative Adversarial Network (PNGAN). PNGAN employs a pre-trained real denoiser to map the fake and real noisy images into a nearly noise-free solution space to perform image domain alignment. Simultaneously, PNGAN establishes a pixel-level adversarial training to conduct noise domain alignment. Additionally, for better noise fitting, we present an efficient architecture Simple Multi-scale Network (SMNet) as the generator. Qualitative validation shows that noise generated by PNGAN is highly similar to real noise in terms of intensity and distribution. Quantitative experiments demonstrate that a series of denoisers trained with the generated noisy images achieve state-of-the-art (SOTA) results on four real denoising benchmarks.
Revisiting RCAN: Improved Training for Image Super-Resolution
Jan 27, 2022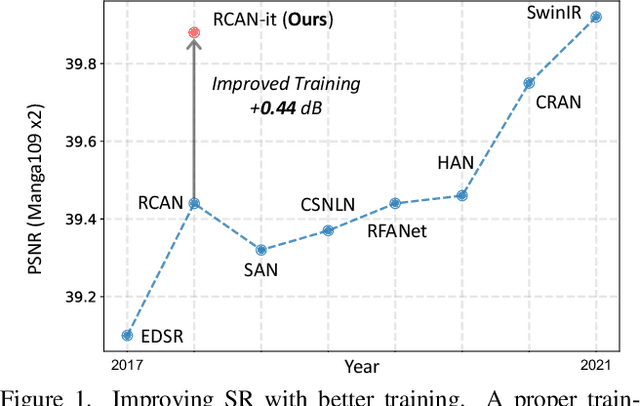
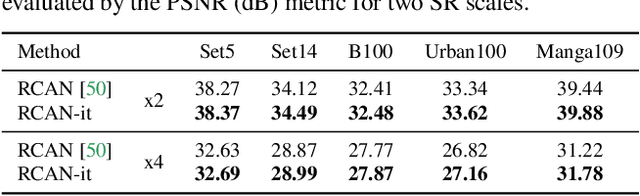
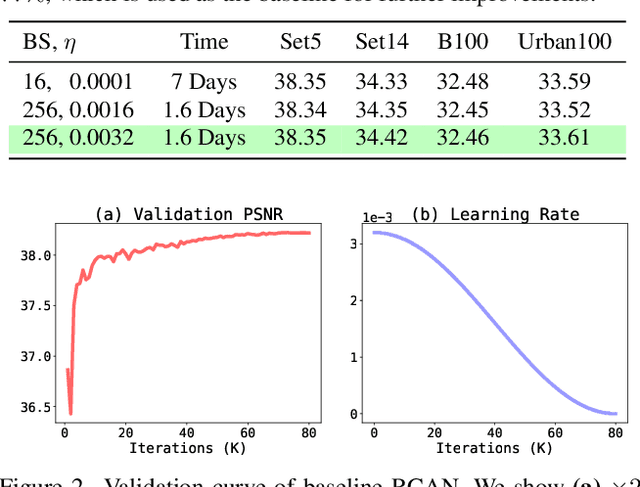
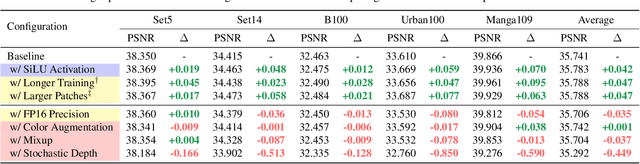
Abstract:Image super-resolution (SR) is a fast-moving field with novel architectures attracting the spotlight. However, most SR models were optimized with dated training strategies. In this work, we revisit the popular RCAN model and examine the effect of different training options in SR. Surprisingly (or perhaps as expected), we show that RCAN can outperform or match nearly all the CNN-based SR architectures published after RCAN on standard benchmarks with a proper training strategy and minimal architecture change. Besides, although RCAN is a very large SR architecture with more than four hundred convolutional layers, we draw a notable conclusion that underfitting is still the main problem restricting the model capability instead of overfitting. We observe supportive evidence that increasing training iterations clearly improves the model performance while applying regularization techniques generally degrades the predictions. We denote our simply revised RCAN as RCAN-it and recommend practitioners to use it as baselines for future research. Code is publicly available at https://github.com/zudi-lin/rcan-it.
PyTorch Connectomics: A Scalable and Flexible Segmentation Framework for EM Connectomics
Dec 10, 2021
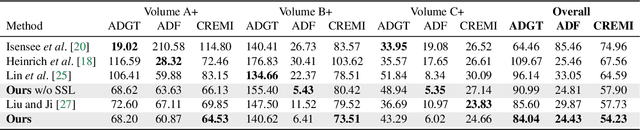
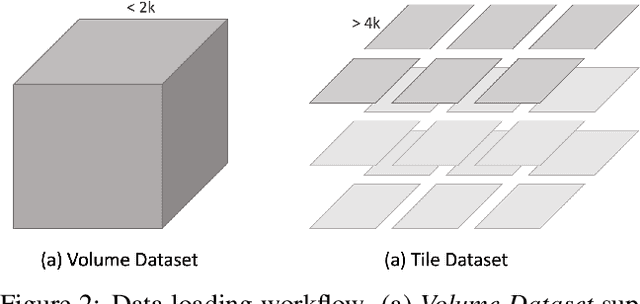
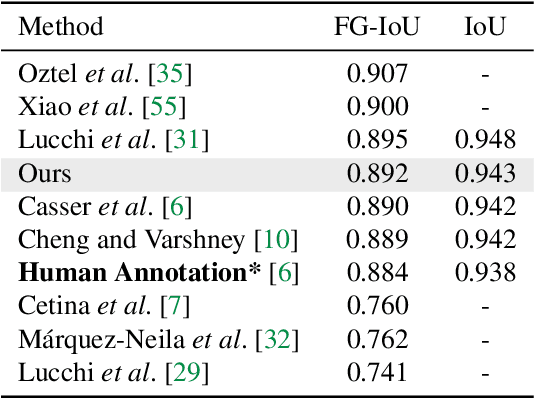
Abstract:We present PyTorch Connectomics (PyTC), an open-source deep-learning framework for the semantic and instance segmentation of volumetric microscopy images, built upon PyTorch. We demonstrate the effectiveness of PyTC in the field of connectomics, which aims to segment and reconstruct neurons, synapses, and other organelles like mitochondria at nanometer resolution for understanding neuronal communication, metabolism, and development in animal brains. PyTC is a scalable and flexible toolbox that tackles datasets at different scales and supports multi-task and semi-supervised learning to better exploit expensive expert annotations and the vast amount of unlabeled data during training. Those functionalities can be easily realized in PyTC by changing the configuration options without coding and adapted to other 2D and 3D segmentation tasks for different tissues and imaging modalities. Quantitatively, our framework achieves the best performance in the CREMI challenge for synaptic cleft segmentation (outperforms existing best result by relatively 6.1$\%$) and competitive performance on mitochondria and neuronal nuclei segmentation. Code and tutorials are publicly available at https://connectomics.readthedocs.io.
Object Propagation via Inter-Frame Attentions for Temporally Stable Video Instance Segmentation
Nov 15, 2021
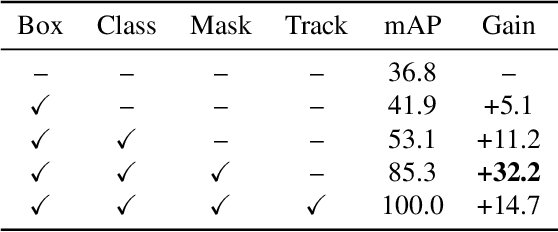
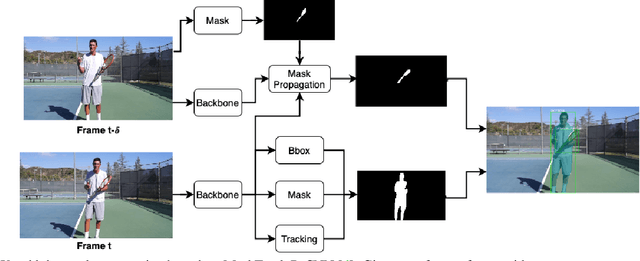
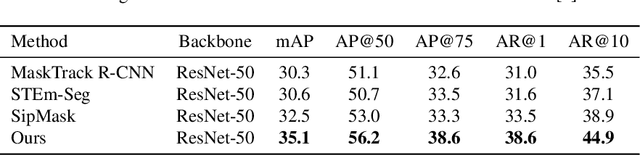
Abstract:Video instance segmentation aims to detect, segment, and track objects in a video. Current approaches extend image-level segmentation algorithms to the temporal domain. However, this results in temporally inconsistent masks. In this work, we identify the mask quality due to temporal stability as a performance bottleneck. Motivated by this, we propose a video instance segmentation method that alleviates the problem due to missing detections. Since this cannot be solved simply using spatial information, we leverage temporal context using inter-frame attentions. This allows our network to refocus on missing objects using box predictions from the neighbouring frame, thereby overcoming missing detections. Our method significantly outperforms previous state-of-the-art algorithms using the Mask R-CNN backbone, by achieving 35.1% mAP on the YouTube-VIS benchmark. Additionally, our method is completely online and requires no future frames. Our code is publicly available at https://github.com/anirudh-chakravarthy/ObjProp.
MedMNIST v2: A Large-Scale Lightweight Benchmark for 2D and 3D Biomedical Image Classification
Oct 27, 2021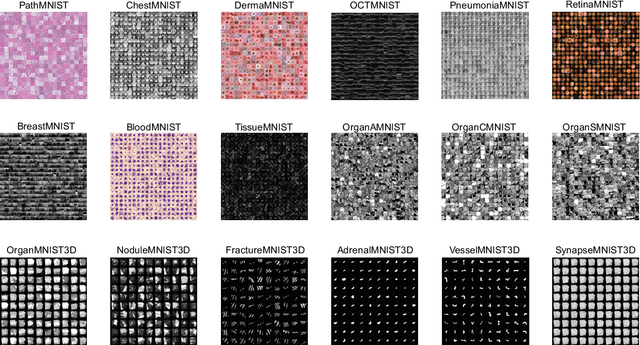
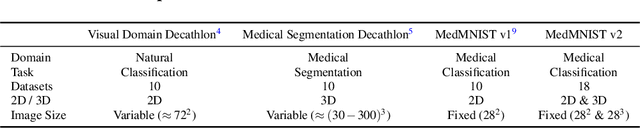

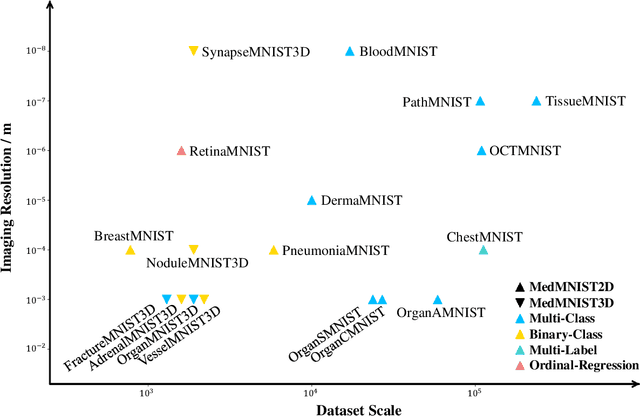
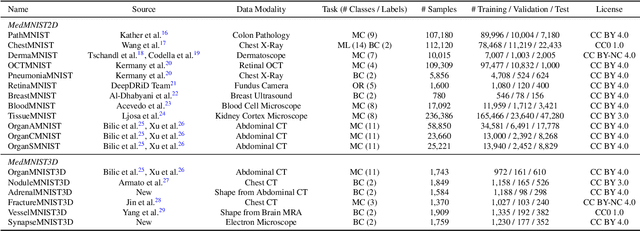
Abstract:We introduce MedMNIST v2, a large-scale MNIST-like dataset collection of standardized biomedical images, including 12 datasets for 2D and 6 datasets for 3D. All images are pre-processed into a small size of 28x28 (2D) or 28x28x28 (3D) with the corresponding classification labels so that no background knowledge is required for users. Covering primary data modalities in biomedical images, MedMNIST v2 is designed to perform classification on lightweight 2D and 3D images with various dataset scales (from 100 to 100,000) and diverse tasks (binary/multi-class, ordinal regression, and multi-label). The resulting dataset, consisting of 708,069 2D images and 10,214 3D images in total, could support numerous research / educational purposes in biomedical image analysis, computer vision, and machine learning. We benchmark several baseline methods on MedMNIST v2, including 2D / 3D neural networks and open-source / commercial AutoML tools. The data and code are publicly available at https://medmnist.com/.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge