Bilian Ke
Learning Unsupervised Gaze Representation via Eye Mask Driven Information Bottleneck
Jun 29, 2024



Abstract:Appearance-based supervised methods with full-face image input have made tremendous advances in recent gaze estimation tasks. However, intensive human annotation requirement inhibits current methods from achieving industrial level accuracy and robustness. Although current unsupervised pre-training frameworks have achieved success in many image recognition tasks, due to the deep coupling between facial and eye features, such frameworks are still deficient in extracting useful gaze features from full-face. To alleviate above limitations, this work proposes a novel unsupervised/self-supervised gaze pre-training framework, which forces the full-face branch to learn a low dimensional gaze embedding without gaze annotations, through collaborative feature contrast and squeeze modules. In the heart of this framework is an alternating eye-attended/unattended masking training scheme, which squeezes gaze-related information from full-face branch into an eye-masked auto-encoder through an injection bottleneck design that successfully encourages the model to pays more attention to gaze direction rather than facial textures only, while still adopting the eye self-reconstruction objective. In the same time, a novel eye/gaze-related information contrastive loss has been designed to further boost the learned representation by forcing the model to focus on eye-centered regions. Extensive experimental results on several gaze benchmarks demonstrate that the proposed scheme achieves superior performances over unsupervised state-of-the-art.
Differentiable Projection from Optical Coherence Tomography B-Scan without Retinal Layer Segmentation Supervision
Jun 11, 2022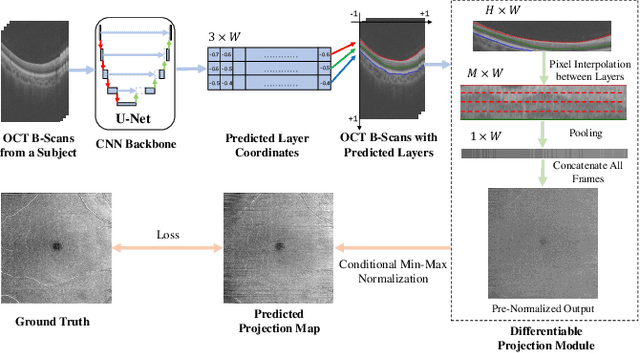
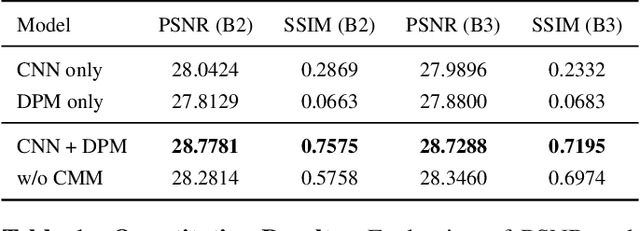
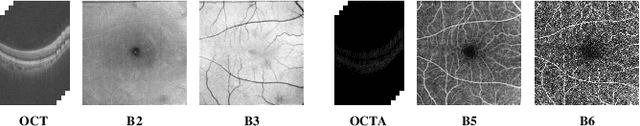
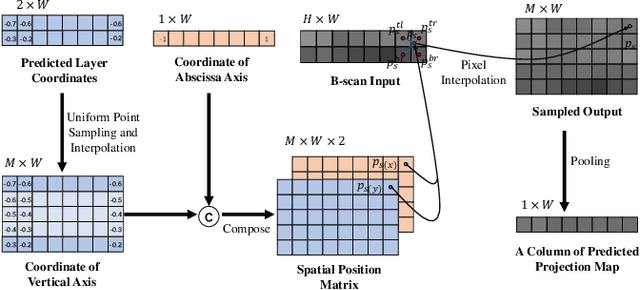
Abstract:Projection map (PM) from optical coherence tomography (OCT) B-scan is an important tool to diagnose retinal diseases, which typically requires retinal layer segmentation. In this study, we present a novel end-to-end framework to predict PMs from B-scans. Instead of segmenting retinal layers explicitly, we represent them implicitly as predicted coordinates. By pixel interpolation on uniformly sampled coordinates between retinal layers, the corresponding PMs could be easily obtained with pooling. Notably, all the operators are differentiable; therefore, this Differentiable Projection Module (DPM) enables end-to-end training with the ground truth of PMs rather than retinal layer segmentation. Our framework produces high-quality PMs, significantly outperforming baselines, including a vanilla CNN without DPM and an optimization-based DPM without a deep prior. Furthermore, the proposed DPM, as a novel neural representation of areas/volumes between curves/surfaces, could be of independent interest for geometric deep learning.
MedMNIST v2: A Large-Scale Lightweight Benchmark for 2D and 3D Biomedical Image Classification
Oct 27, 2021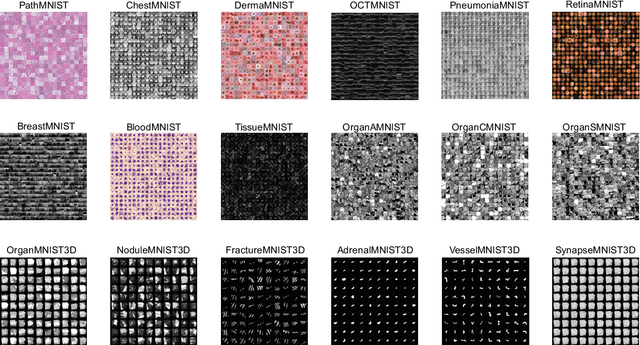
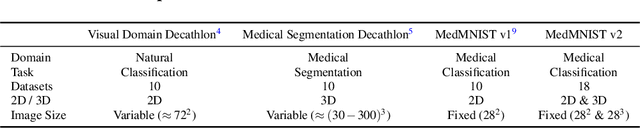

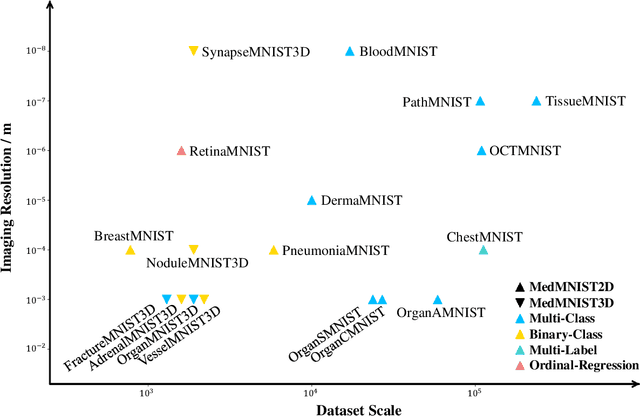
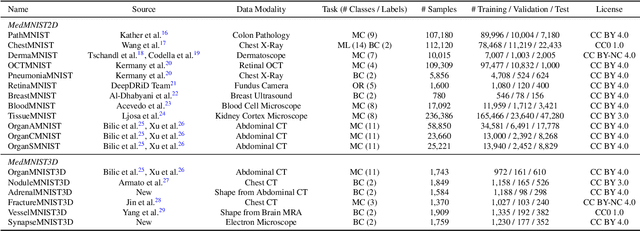
Abstract:We introduce MedMNIST v2, a large-scale MNIST-like dataset collection of standardized biomedical images, including 12 datasets for 2D and 6 datasets for 3D. All images are pre-processed into a small size of 28x28 (2D) or 28x28x28 (3D) with the corresponding classification labels so that no background knowledge is required for users. Covering primary data modalities in biomedical images, MedMNIST v2 is designed to perform classification on lightweight 2D and 3D images with various dataset scales (from 100 to 100,000) and diverse tasks (binary/multi-class, ordinal regression, and multi-label). The resulting dataset, consisting of 708,069 2D images and 10,214 3D images in total, could support numerous research / educational purposes in biomedical image analysis, computer vision, and machine learning. We benchmark several baseline methods on MedMNIST v2, including 2D / 3D neural networks and open-source / commercial AutoML tools. The data and code are publicly available at https://medmnist.com/.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge