"Image": models, code, and papers
MonoScene: Monocular 3D Semantic Scene Completion
Dec 01, 2021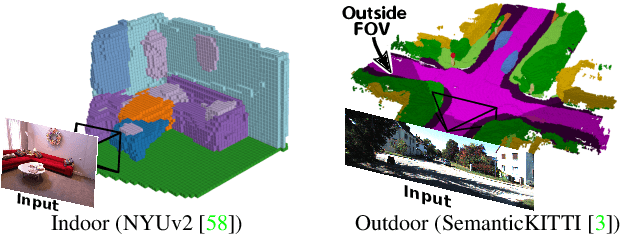
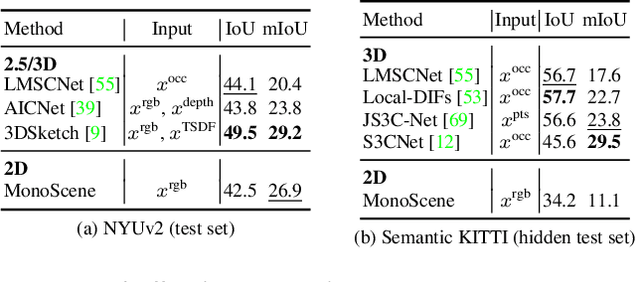
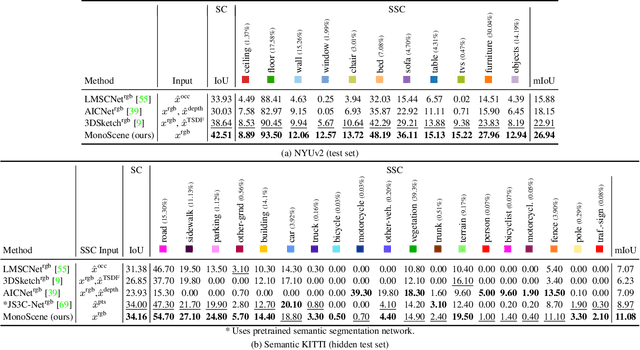
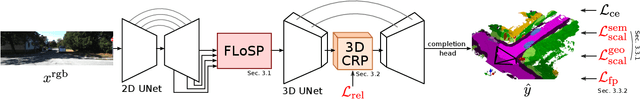

MonoScene proposes a 3D Semantic Scene Completion (SSC) framework, where the dense geometry and semantics of a scene are inferred from a single monocular RGB image. Different from the SSC literature, relying on 2.5 or 3D input, we solve the complex problem of 2D to 3D scene reconstruction while jointly inferring its semantics. Our framework relies on successive 2D and 3D UNets bridged by a novel 2D-3D features projection inspiring from optics and introduces a 3D context relation prior to enforce spatio-semantic consistency. Along with architectural contributions, we introduce novel global scene and local frustums losses. Experiments show we outperform the literature on all metrics and datasets while hallucinating plausible scenery even beyond the camera field of view. Our code and trained models are available at https://github.com/cv-rits/MonoScene
Assessing learned features of Deep Learning applied to EEG
Nov 08, 2021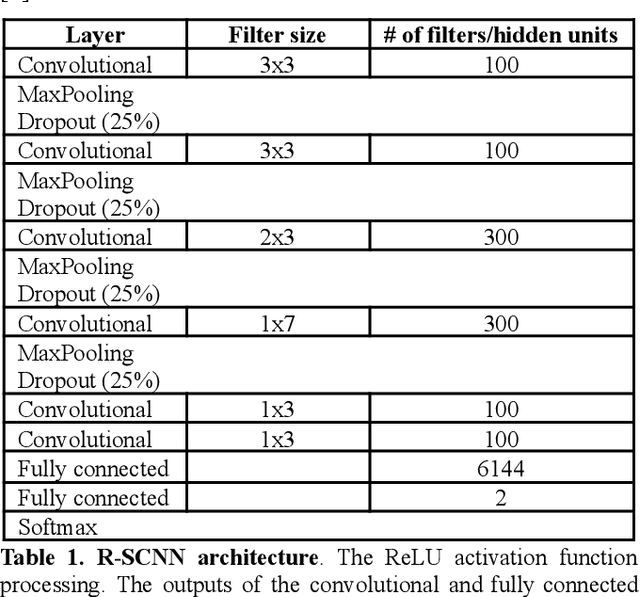
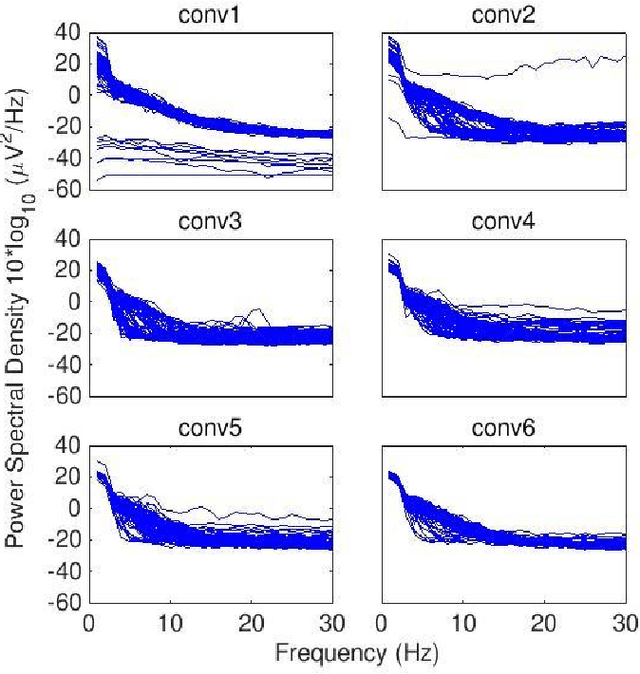
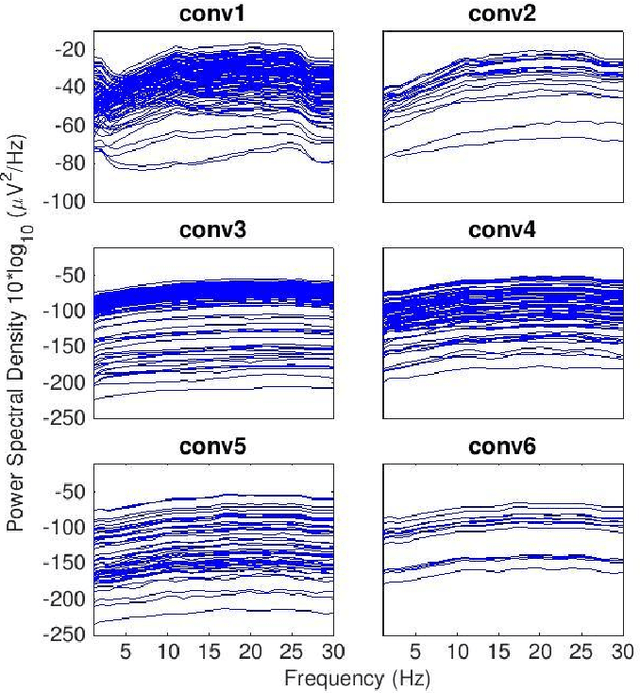
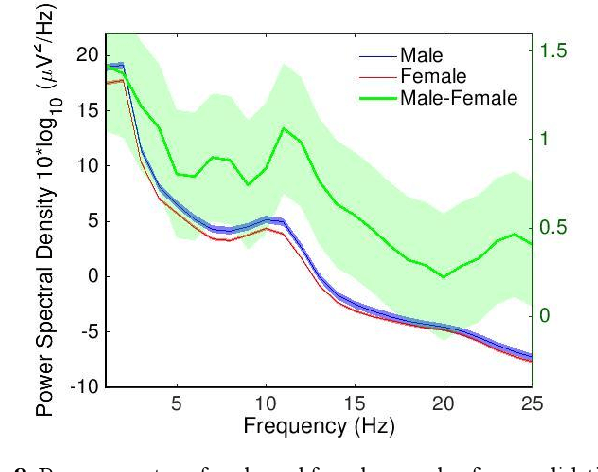
Convolutional Neural Networks (CNNs) have achieved impressive performance on many computer vision related tasks, such as object detection, image recognition, image retrieval, etc. These achievements benefit from the CNNs' outstanding capability to learn discriminative features with deep layers of neuron structures and iterative training process. This has inspired the EEG research community to adopt CNN in performing EEG classification tasks. However, CNNs learned features are not immediately interpretable, causing a lack of understanding of the CNNs' internal working mechanism. To improve CNN interpretability, CNN visualization methods are applied to translate the internal features into visually perceptible patterns for qualitative analysis of CNN layers. Many CNN visualization methods have been proposed in the Computer Vision literature to interpret the CNN network structure, operation, and semantic concept, yet applications to EEG data analysis have been limited. In this work we use 3 different methods to extract EEG-relevant features from a CNN trained on raw EEG data: optimal samples for each classification category, activation maximization, and reverse convolution. We applied these methods to a high-performing Deep Learning model with state-of-the-art performance for an EEG sex classification task, and show that the model features a difference in the theta frequency band. We show that visualization of a CNN model can reveal interesting EEG results. Using these tools, EEG researchers using Deep Learning can better identify the learned EEG features, possibly identifying new class relevant biomarkers.
SIR: Self-supervised Image Rectification via Seeing the Same Scene from Multiple Different Lenses
Nov 30, 2020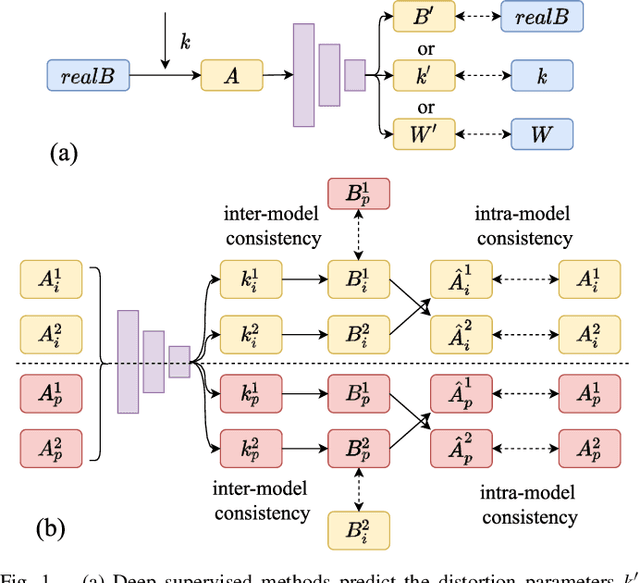
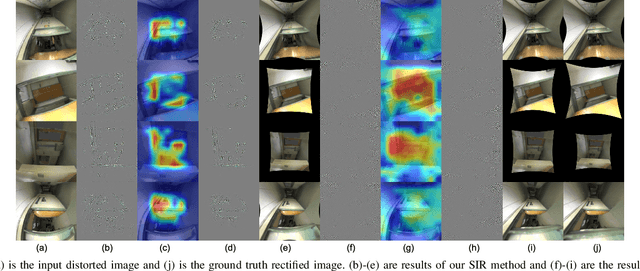
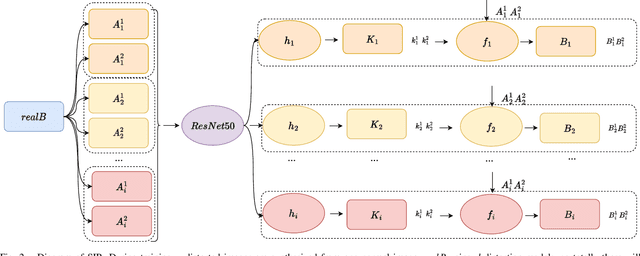
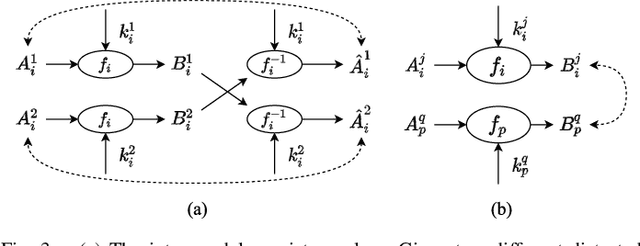
Deep learning has demonstrated its power in image rectification by leveraging the representation capacity of deep neural networks via supervised training based on a large-scale synthetic dataset. However, the model may overfit the synthetic images and generalize not well on real-world fisheye images due to the limited universality of a specific distortion model and the lack of explicitly modeling the distortion and rectification process. In this paper, we propose a novel self-supervised image rectification (SIR) method based on an important insight that the rectified results of distorted images of the same scene from different lens should be the same. Specifically, we devise a new network architecture with a shared encoder and several prediction heads, each of which predicts the distortion parameter of a specific distortion model. We further leverage a differentiable warping module to generate the rectified images and re-distorted images from the distortion parameters and exploit the intra- and inter-model consistency between them during training, thereby leading to a self-supervised learning scheme without the need for ground-truth distortion parameters or normal images. Experiments on synthetic dataset and real-world fisheye images demonstrate that our method achieves comparable or even better performance than the supervised baseline method and representative state-of-the-art methods. Self-supervised learning also improves the universality of distortion models while keeping their self-consistency.
Robust and Resource-Efficient Data-Free Knowledge Distillation by Generative Pseudo Replay
Jan 09, 2022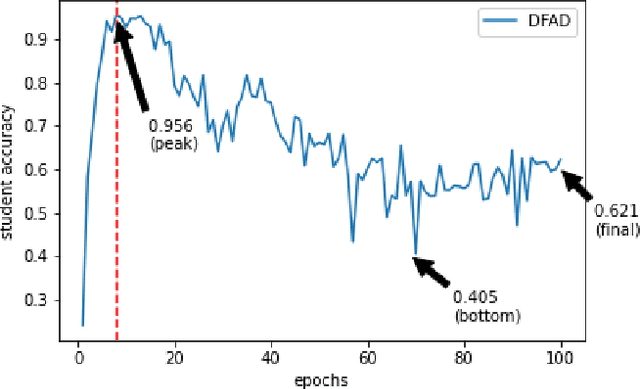
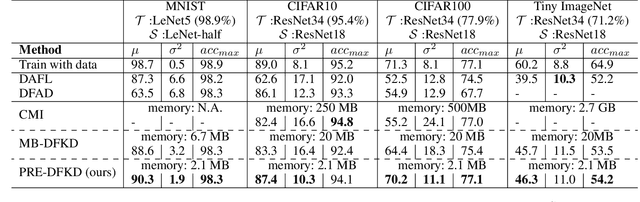

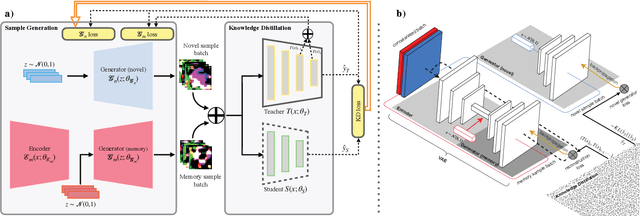

Data-Free Knowledge Distillation (KD) allows knowledge transfer from a trained neural network (teacher) to a more compact one (student) in the absence of original training data. Existing works use a validation set to monitor the accuracy of the student over real data and report the highest performance throughout the entire process. However, validation data may not be available at distillation time either, making it infeasible to record the student snapshot that achieved the peak accuracy. Therefore, a practical data-free KD method should be robust and ideally provide monotonically increasing student accuracy during distillation. This is challenging because the student experiences knowledge degradation due to the distribution shift of the synthetic data. A straightforward approach to overcome this issue is to store and rehearse the generated samples periodically, which increases the memory footprint and creates privacy concerns. We propose to model the distribution of the previously observed synthetic samples with a generative network. In particular, we design a Variational Autoencoder (VAE) with a training objective that is customized to learn the synthetic data representations optimally. The student is rehearsed by the generative pseudo replay technique, with samples produced by the VAE. Hence knowledge degradation can be prevented without storing any samples. Experiments on image classification benchmarks show that our method optimizes the expected value of the distilled model accuracy while eliminating the large memory overhead incurred by the sample-storing methods.
Learning Hierarchical Graph Neural Networks for Image Clustering
Jul 17, 2021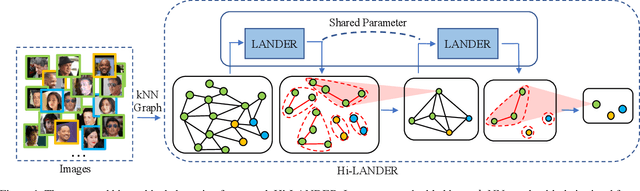
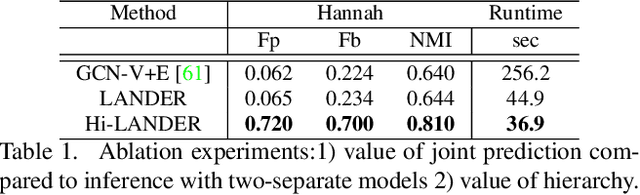
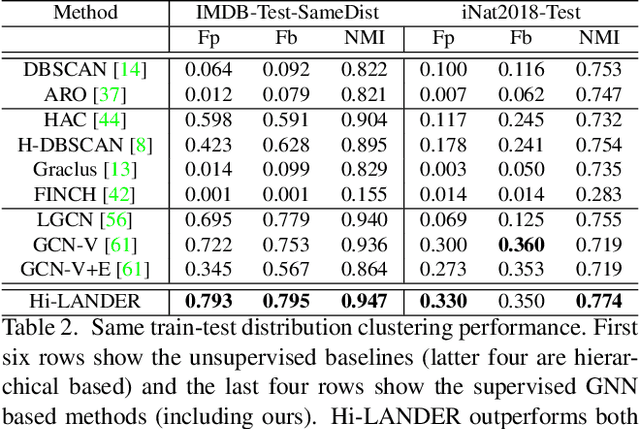
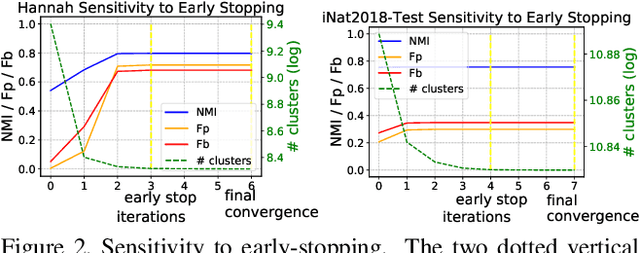
We propose a hierarchical graph neural network (GNN) model that learns how to cluster a set of images into an unknown number of identities using a training set of images annotated with labels belonging to a disjoint set of identities. Our hierarchical GNN uses a novel approach to merge connected components predicted at each level of the hierarchy to form a new graph at the next level. Unlike fully unsupervised hierarchical clustering, the choice of grouping and complexity criteria stems naturally from supervision in the training set. The resulting method, Hi-LANDER, achieves an average of 54% improvement in F-score and 8% increase in Normalized Mutual Information (NMI) relative to current GNN-based clustering algorithms. Additionally, state-of-the-art GNN-based methods rely on separate models to predict linkage probabilities and node densities as intermediate steps of the clustering process. In contrast, our unified framework achieves a seven-fold decrease in computational cost. We release our training and inference code at https://github.com/dmlc/dgl/tree/master/examples/pytorch/hilander.
Extending nn-UNet for brain tumor segmentation
Dec 09, 2021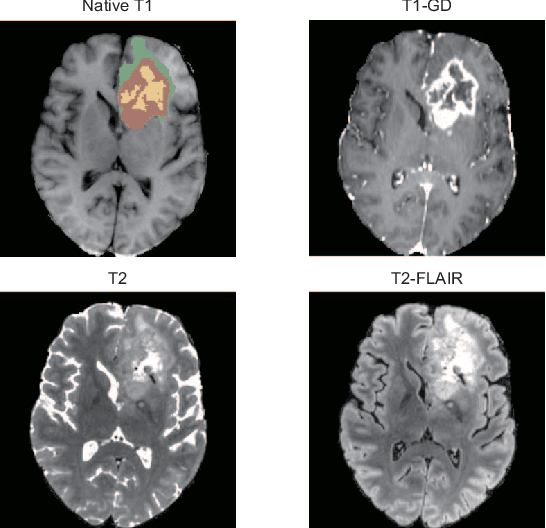
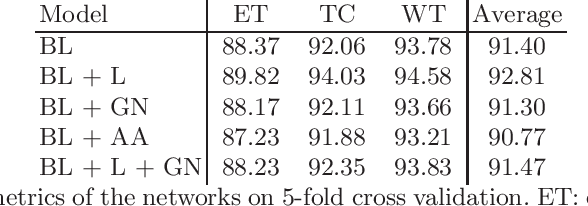
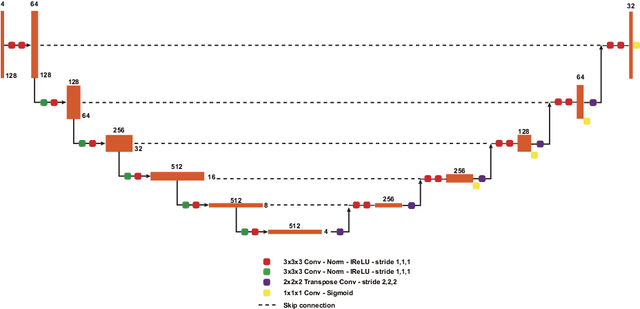
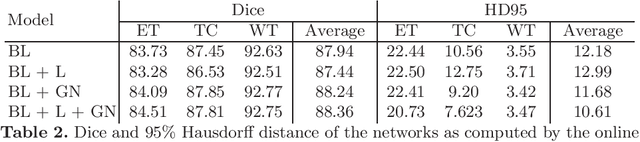
Brain tumor segmentation is essential for the diagnosis and prognosis of patients with gliomas. The brain tumor segmentation challenge has continued to provide a great source of data to develop automatic algorithms to perform the task. This paper describes our contribution to the 2021 competition. We developed our methods based on nn-UNet, the winning entry of last year competition. We experimented with several modifications, including using a larger network, replacing batch normalization with group normalization, and utilizing axial attention in the decoder. Internal 5-fold cross validation as well as online evaluation from the organizers showed the effectiveness of our approach, with minor improvement in quantitative metrics when compared to the baseline. The proposed models won first place in the final ranking on unseen test data. The codes, pretrained weights, and docker image for the winning submission are publicly available at https://github.com/rixez/Brats21_KAIST_MRI_Lab
Tensor Networks for Medical Image Classification
Apr 21, 2020
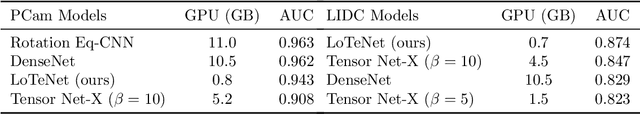


With the increasing adoption of machine learning tools like neural networks across several domains, interesting connections and comparisons to concepts from other domains are coming to light. In this work, we focus on the class of Tensor Networks, which has been a work horse for physicists in the last two decades to analyse quantum many-body systems. Building on the recent interest in tensor networks for machine learning, we extend the Matrix Product State tensor networks (which can be interpreted as linear classifiers operating in exponentially high dimensional spaces) to be useful in medical image analysis tasks. We focus on classification problems as a first step where we motivate the use of tensor networks and propose adaptions for 2D images using classical image domain concepts such as local orderlessness of images. With the proposed locally orderless tensor network model (LoTeNet), we show that tensor networks are capable of attaining performance that is comparable to state-of-the-art deep learning methods. We evaluate the model on two publicly available medical imaging datasets and show performance improvements with fewer model hyperparameters and lesser computational resources compared to relevant baseline methods.
ZstGAN: An Adversarial Approach for Unsupervised Zero-Shot Image-to-Image Translation
Jun 01, 2019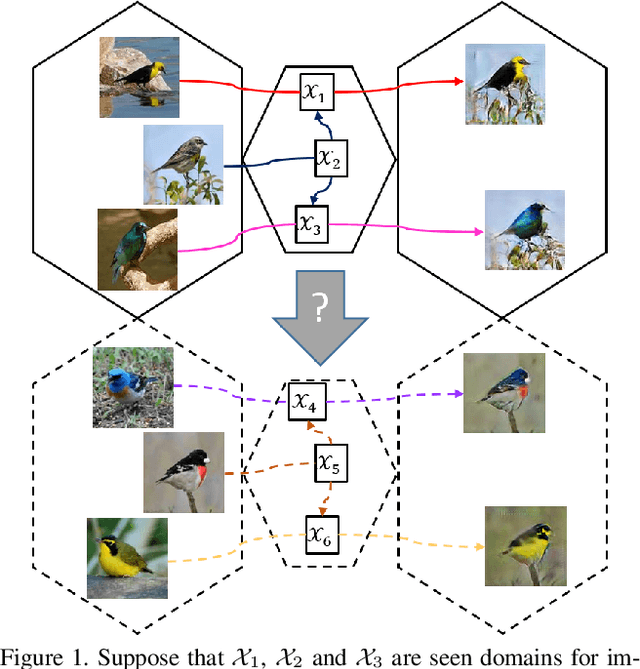
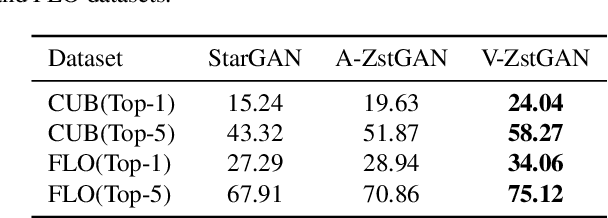
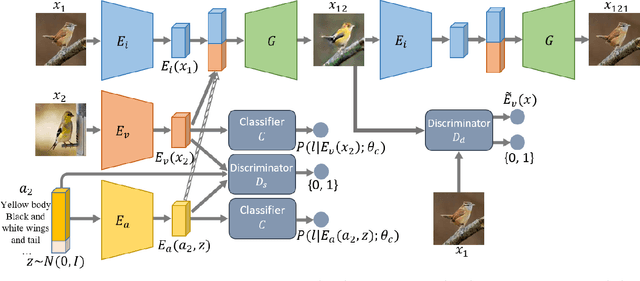
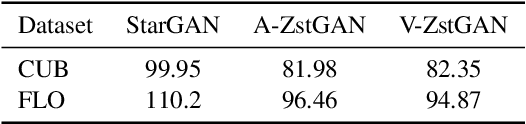
Image-to-image translation models have shown remarkable ability on transferring images among different domains. Most of existing work follows the setting that the source domain and target domain keep the same at training and inference phases, which cannot be generalized to the scenarios for translating an image from an unseen domain to an another unseen domain. In this work, we propose the Unsupervised Zero-Shot Image-to-image Translation (UZSIT) problem, which aims to learn a model that can transfer translation knowledge from seen domains to unseen domains. Accordingly, we propose a framework called ZstGAN: By introducing an adversarial training scheme, ZstGAN learns to model each domain with domain-specific feature distribution that is semantically consistent on vision and attribute modalities. Then the domain-invariant features are disentangled with an shared encoder for image generation. We carry out extensive experiments on CUB and FLO datasets, and the results demonstrate the effectiveness of proposed method on UZSIT task. Moreover, ZstGAN shows significant accuracy improvements over state-of-the-art zero-shot learning methods on CUB and FLO.
SLCRF: Subspace Learning with Conditional Random Field for Hyperspectral Image Classification
Oct 07, 2020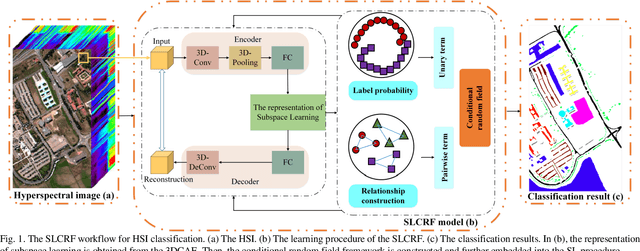

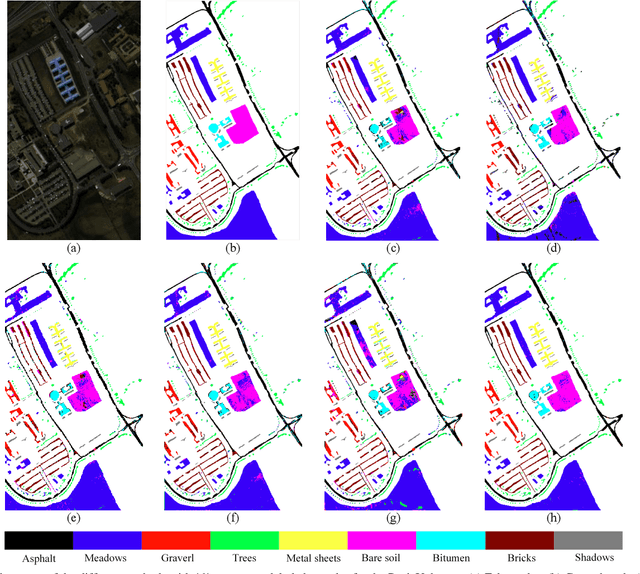
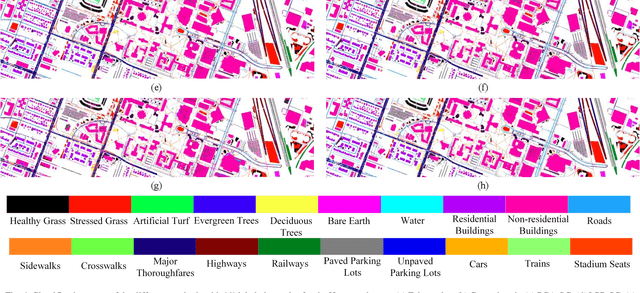
Subspace learning (SL) plays an important role in hyperspectral image (HSI) classification, since it can provide an effective solution to reduce the redundant information in the image pixels of HSIs. Previous works about SL aim to improve the accuracy of HSI recognition. Using a large number of labeled samples, related methods can train the parameters of the proposed solutions to obtain better representations of HSI pixels. However, the data instances may not be sufficient enough to learn a precise model for HSI classification in real applications. Moreover, it is well-known that it takes much time, labor and human expertise to label HSI images. To avoid the aforementioned problems, a novel SL method that includes the probability assumption called subspace learning with conditional random field (SLCRF) is developed. In SLCRF, first, the 3D convolutional autoencoder (3DCAE) is introduced to remove the redundant information in HSI pixels. In addition, the relationships are also constructed using the spectral-spatial information among the adjacent pixels. Then, the conditional random field (CRF) framework can be constructed and further embedded into the HSI SL procedure with the semi-supervised approach. Through the linearized alternating direction method termed LADMAP, the objective function of SLCRF is optimized using a defined iterative algorithm. The proposed method is comprehensively evaluated using the challenging public HSI datasets. We can achieve stateof-the-art performance using these HSI sets.
HRINet: Alternative Supervision Network for High-resolution CT image Interpolation
Feb 11, 2020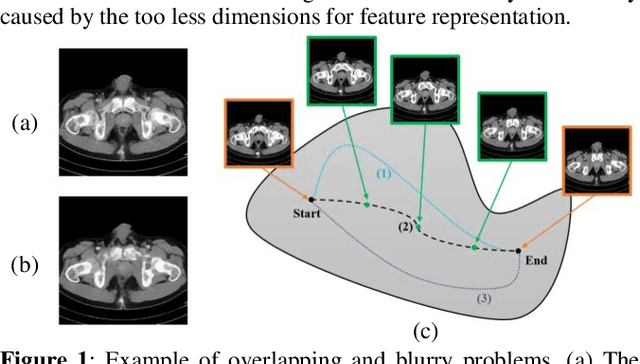
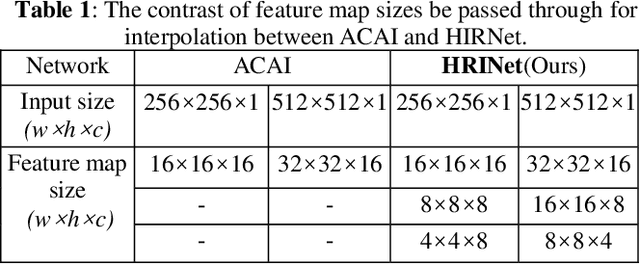

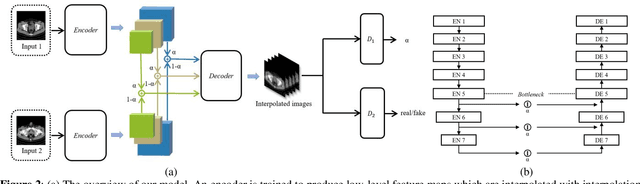
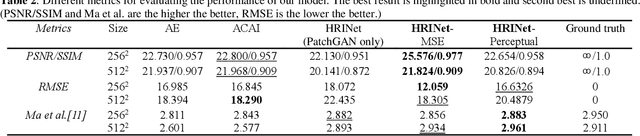
Image interpolation in medical area is of high importance as most 3D biomedical volume images are sampled where the distance between consecutive slices significantly greater than the in-plane pixel size due to radiation dose or scanning time. Image interpolation creates a number of new slices between known slices in order to obtain an isotropic volume image. The results can be used for the higher quality of 3D reconstruction and visualization of human body structures. Semantic interpolation on the manifold has been proved to be very useful for smoothing image interpolation. Nevertheless, all previous methods focused on low-resolution image interpolation, and most of them work poorly on high-resolution image. We propose a novel network, High Resolution Interpolation Network (HRINet), aiming at producing high-resolution CT image interpolations. We combine the idea of ACAI and GANs, and propose a novel idea of alternative supervision method by applying supervised and unsupervised training alternatively to raise the accuracy of human organ structures in CT while keeping high quality. We compare an MSE based and a perceptual based loss optimizing methods for high quality interpolation, and show the tradeoff between the structural correctness and sharpness. Our experiments show the great improvement on 256 2 and 5122 images quantitatively and qualitatively.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge