Zhi Yang
Cascade Bagging for Accuracy Prediction with Few Training Samples
Aug 12, 2021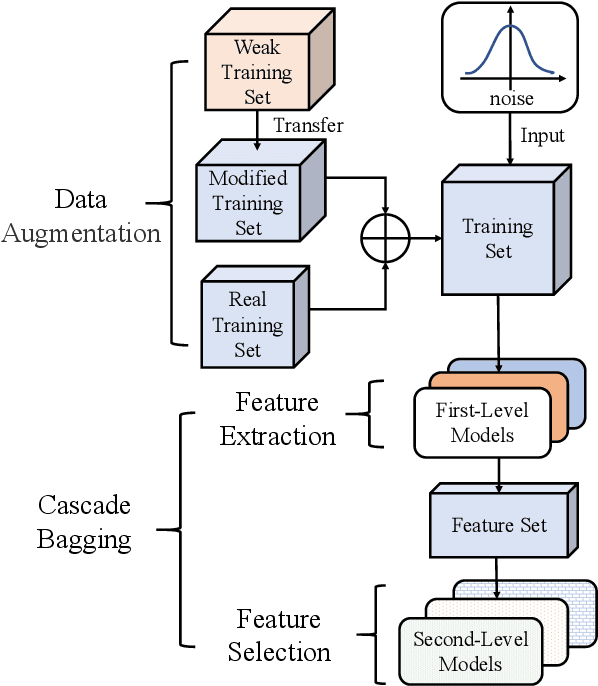
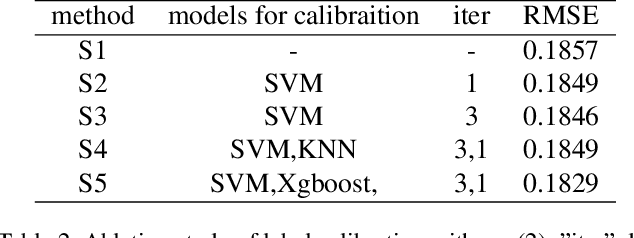
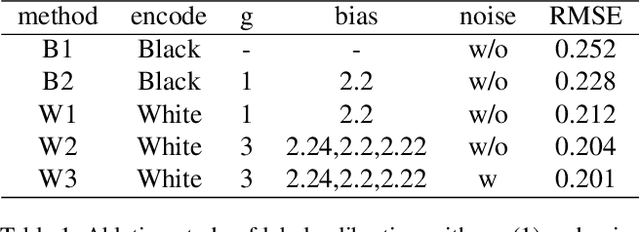
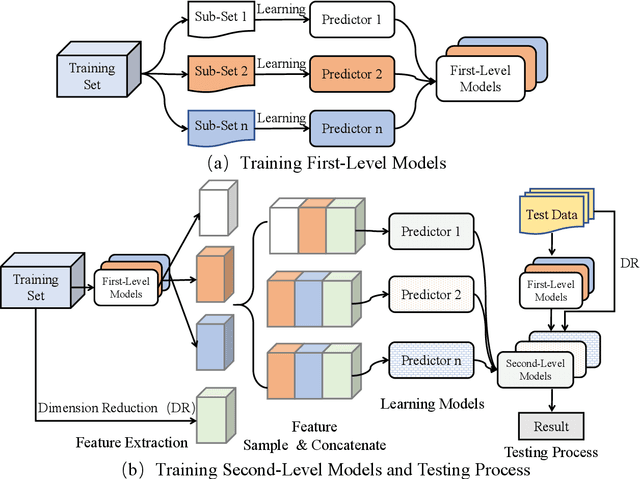

Abstract:Accuracy predictor is trained to predict the validation accuracy of an network from its architecture encoding. It can effectively assist in designing networks and improving Neural Architecture Search(NAS) efficiency. However, a high-performance predictor depends on adequate trainning samples, which requires unaffordable computation overhead. To alleviate this problem, we propose a novel framework to train an accuracy predictor under few training samples. The framework consists ofdata augmentation methods and an ensemble learning algorithm. The data augmentation methods calibrate weak labels and inject noise to feature space. The ensemble learning algorithm, termed cascade bagging, trains two-level models by sampling data and features. In the end, the advantages of above methods are proved in the Performance Prediciton Track of CVPR2021 1st Lightweight NAS Challenge. Our code is made public at: https://github.com/dlongry/Solutionto-CVPR2021-NAS-Track2.
Evaluating Deep Graph Neural Networks
Aug 02, 2021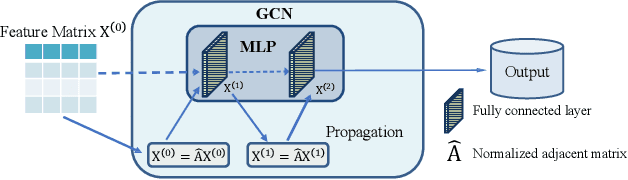
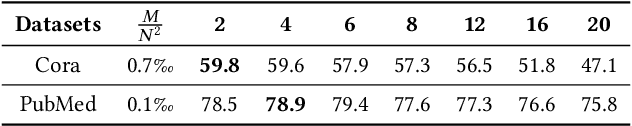
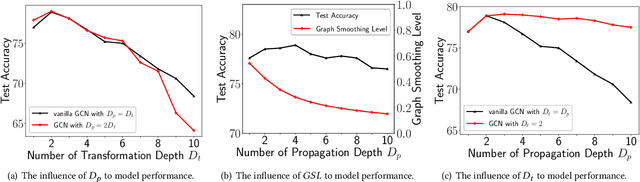
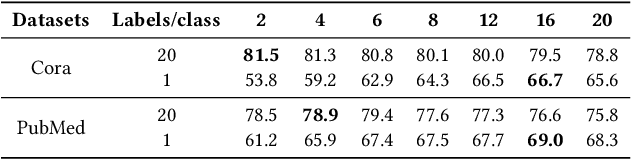
Abstract:Graph Neural Networks (GNNs) have already been widely applied in various graph mining tasks. However, they suffer from the shallow architecture issue, which is the key impediment that hinders the model performance improvement. Although several relevant approaches have been proposed, none of the existing studies provides an in-depth understanding of the root causes of performance degradation in deep GNNs. In this paper, we conduct the first systematic experimental evaluation to present the fundamental limitations of shallow architectures. Based on the experimental results, we answer the following two essential questions: (1) what actually leads to the compromised performance of deep GNNs; (2) when we need and how to build deep GNNs. The answers to the above questions provide empirical insights and guidelines for researchers to design deep and well-performed GNNs. To show the effectiveness of our proposed guidelines, we present Deep Graph Multi-Layer Perceptron (DGMLP), a powerful approach (a paradigm in its own right) that helps guide deep GNN designs. Experimental results demonstrate three advantages of DGMLP: 1) high accuracy -- it achieves state-of-the-art node classification performance on various datasets; 2) high flexibility -- it can flexibly choose different propagation and transformation depths according to graph size and sparsity; 3) high scalability and efficiency -- it supports fast training on large-scale graphs. Our code is available in https://github.com/zwt233/DGMLP.
Grain: Improving Data Efficiency of Graph Neural Networks via Diversified Influence Maximization
Jul 31, 2021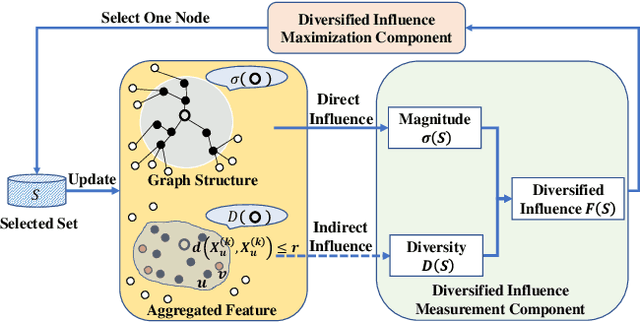
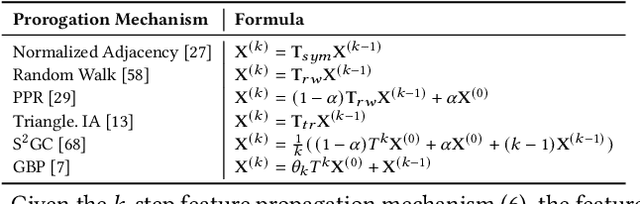

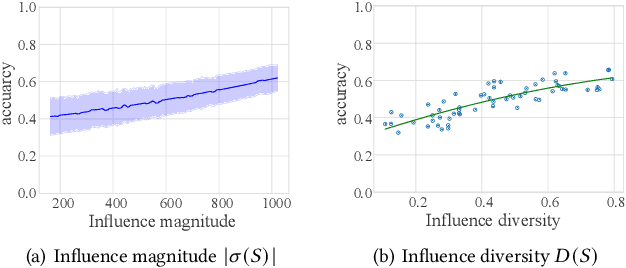
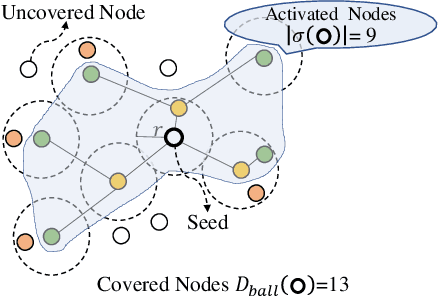
Abstract:Data selection methods, such as active learning and core-set selection, are useful tools for improving the data efficiency of deep learning models on large-scale datasets. However, recent deep learning models have moved forward from independent and identically distributed data to graph-structured data, such as social networks, e-commerce user-item graphs, and knowledge graphs. This evolution has led to the emergence of Graph Neural Networks (GNNs) that go beyond the models existing data selection methods are designed for. Therefore, we present Grain, an efficient framework that opens up a new perspective through connecting data selection in GNNs with social influence maximization. By exploiting the common patterns of GNNs, Grain introduces a novel feature propagation concept, a diversified influence maximization objective with novel influence and diversity functions, and a greedy algorithm with an approximation guarantee into a unified framework. Empirical studies on public datasets demonstrate that Grain significantly improves both the performance and efficiency of data selection (including active learning and core-set selection) for GNNs. To the best of our knowledge, this is the first attempt to bridge two largely parallel threads of research, data selection, and social influence maximization, in the setting of GNNs, paving new ways for improving data efficiency.
ROD: Reception-aware Online Distillation for Sparse Graphs
Jul 25, 2021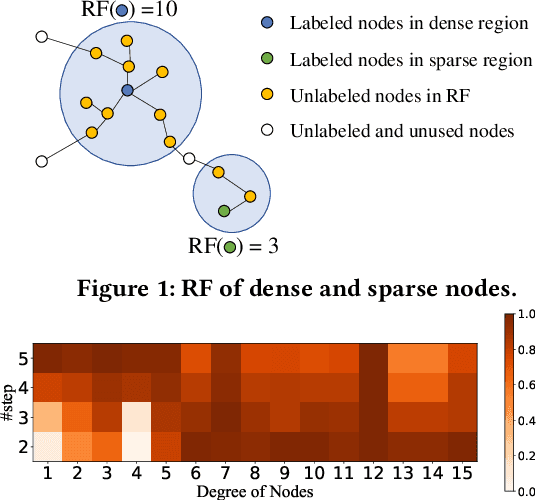
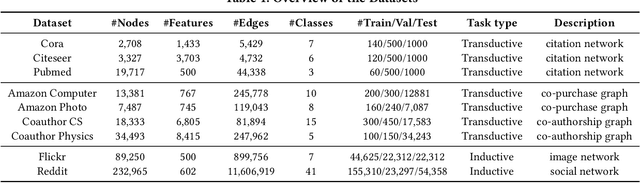
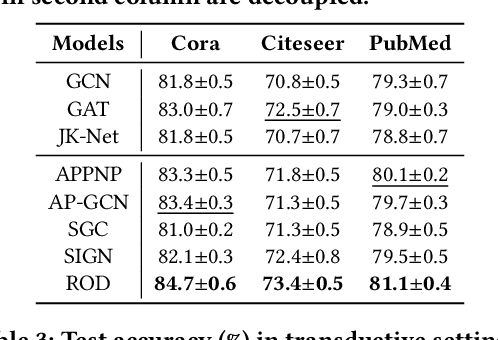
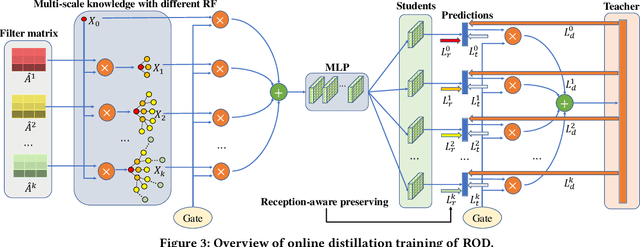
Abstract:Graph neural networks (GNNs) have been widely used in many graph-based tasks such as node classification, link prediction, and node clustering. However, GNNs gain their performance benefits mainly from performing the feature propagation and smoothing across the edges of the graph, thus requiring sufficient connectivity and label information for effective propagation. Unfortunately, many real-world networks are sparse in terms of both edges and labels, leading to sub-optimal performance of GNNs. Recent interest in this sparse problem has focused on the self-training approach, which expands supervised signals with pseudo labels. Nevertheless, the self-training approach inherently cannot realize the full potential of refining the learning performance on sparse graphs due to the unsatisfactory quality and quantity of pseudo labels. In this paper, we propose ROD, a novel reception-aware online knowledge distillation approach for sparse graph learning. We design three supervision signals for ROD: multi-scale reception-aware graph knowledge, task-based supervision, and rich distilled knowledge, allowing online knowledge transfer in a peer-teaching manner. To extract knowledge concealed in the multi-scale reception fields, ROD explicitly requires individual student models to preserve different levels of locality information. For a given task, each student would predict based on its reception-scale knowledge, while simultaneously a strong teacher is established on-the-fly by combining multi-scale knowledge. Our approach has been extensively evaluated on 9 datasets and a variety of graph-based tasks, including node classification, link prediction, and node clustering. The result demonstrates that ROD achieves state-of-art performance and is more robust for the graph sparsity.
VolcanoML: Speeding up End-to-End AutoML via Scalable Search Space Decomposition
Jul 20, 2021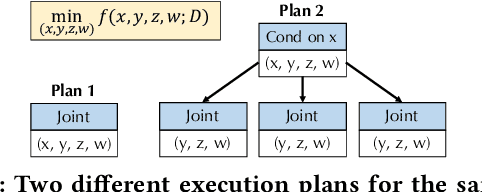
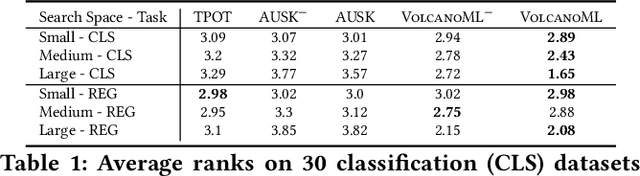
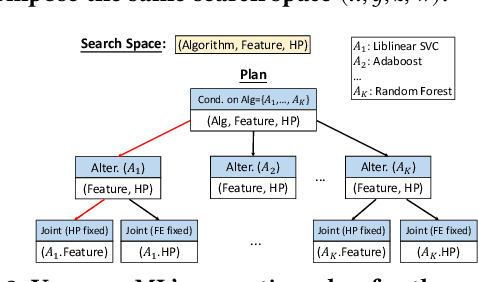
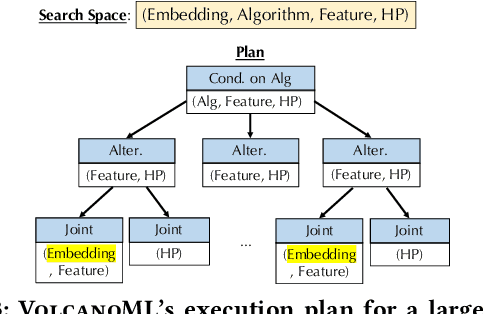
Abstract:End-to-end AutoML has attracted intensive interests from both academia and industry, which automatically searches for ML pipelines in a space induced by feature engineering, algorithm/model selection, and hyper-parameter tuning. Existing AutoML systems, however, suffer from scalability issues when applying to application domains with large, high-dimensional search spaces. We present VolcanoML, a scalable and extensible framework that facilitates systematic exploration of large AutoML search spaces. VolcanoML introduces and implements basic building blocks that decompose a large search space into smaller ones, and allows users to utilize these building blocks to compose an execution plan for the AutoML problem at hand. VolcanoML further supports a Volcano-style execution model - akin to the one supported by modern database systems - to execute the plan constructed. Our evaluation demonstrates that, not only does VolcanoML raise the level of expressiveness for search space decomposition in AutoML, it also leads to actual findings of decomposition strategies that are significantly more efficient than the ones employed by state-of-the-art AutoML systems such as auto-sklearn.
OpenBox: A Generalized Black-box Optimization Service
Jun 06, 2021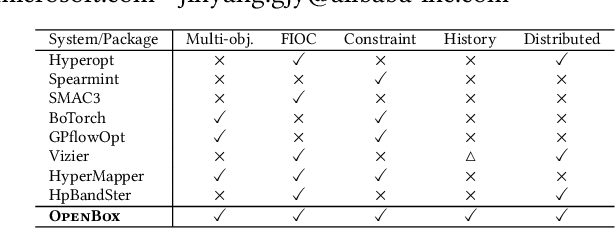
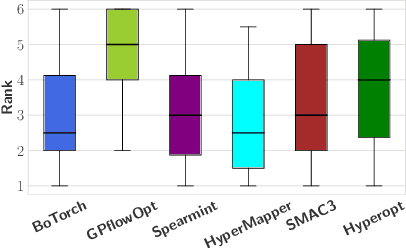
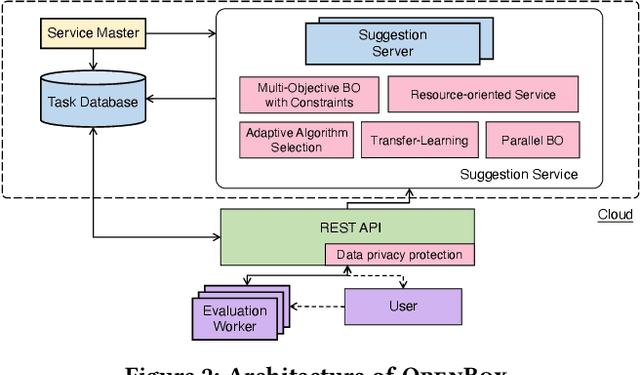

Abstract:Black-box optimization (BBO) has a broad range of applications, including automatic machine learning, engineering, physics, and experimental design. However, it remains a challenge for users to apply BBO methods to their problems at hand with existing software packages, in terms of applicability, performance, and efficiency. In this paper, we build OpenBox, an open-source and general-purpose BBO service with improved usability. The modular design behind OpenBox also facilitates flexible abstraction and optimization of basic BBO components that are common in other existing systems. OpenBox is distributed, fault-tolerant, and scalable. To improve efficiency, OpenBox further utilizes "algorithm agnostic" parallelization and transfer learning. Our experimental results demonstrate the effectiveness and efficiency of OpenBox compared to existing systems.
GMLP: Building Scalable and Flexible Graph Neural Networks with Feature-Message Passing
Apr 20, 2021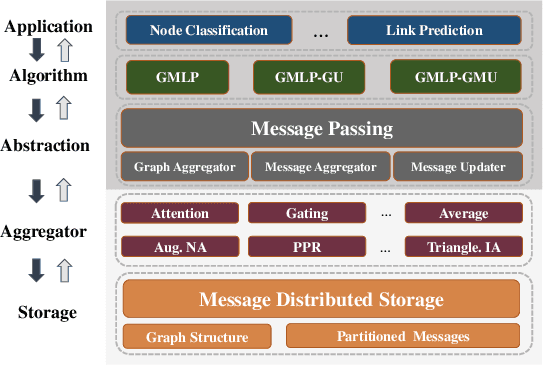
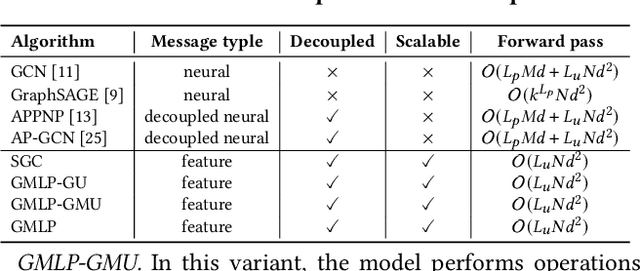
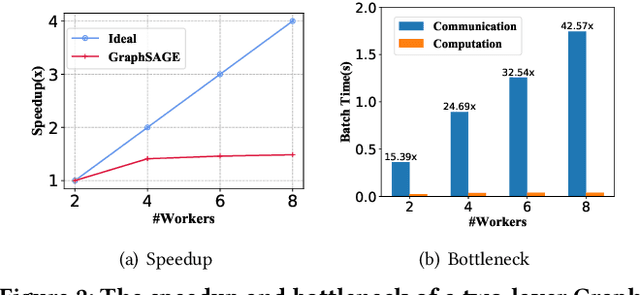
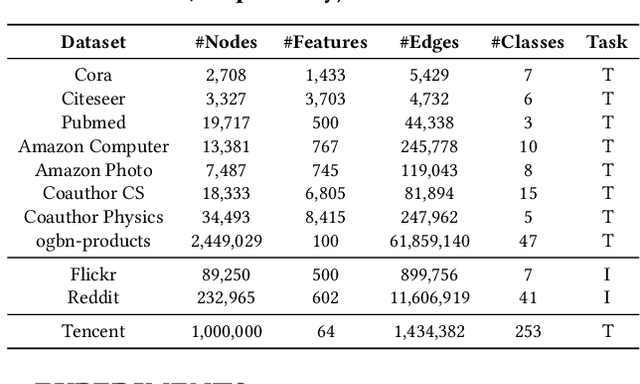
Abstract:In recent studies, neural message passing has proved to be an effective way to design graph neural networks (GNNs), which have achieved state-of-the-art performance in many graph-based tasks. However, current neural-message passing architectures typically need to perform an expensive recursive neighborhood expansion in multiple rounds and consequently suffer from a scalability issue. Moreover, most existing neural-message passing schemes are inflexible since they are restricted to fixed-hop neighborhoods and insensitive to the actual demands of different nodes. We circumvent these limitations by a novel feature-message passing framework, called Graph Multi-layer Perceptron (GMLP), which separates the neural update from the message passing. With such separation, GMLP significantly improves the scalability and efficiency by performing the message passing procedure in a pre-compute manner, and is flexible and adaptive in leveraging node feature messages over various levels of localities. We further derive novel variants of scalable GNNs under this framework to achieve the best of both worlds in terms of performance and efficiency. We conduct extensive evaluations on 11 benchmark datasets, including large-scale datasets like ogbn-products and an industrial dataset, demonstrating that GMLP achieves not only the state-of-art performance, but also high training scalability and efficiency.
A Portable, Self-Contained Neuroprosthetic Hand with Deep Learning-Based Finger Control
Mar 24, 2021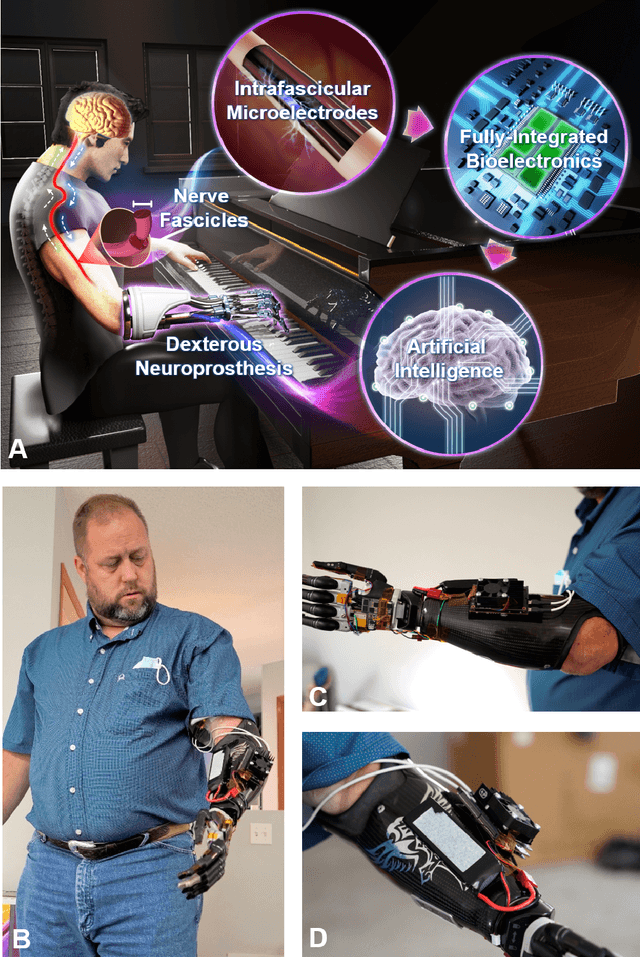
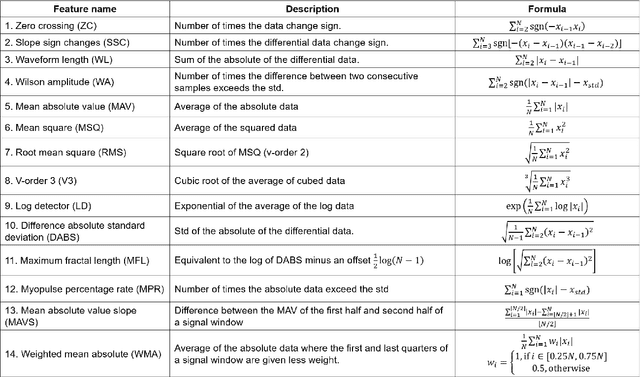

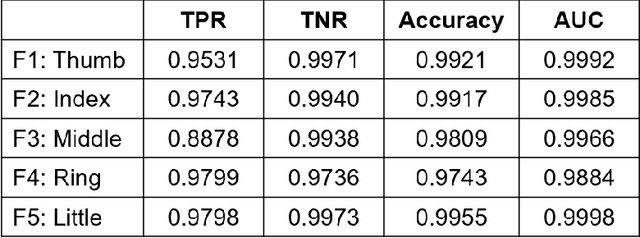
Abstract:Objective: Deep learning-based neural decoders have emerged as the prominent approach to enable dexterous and intuitive control of neuroprosthetic hands. Yet few studies have materialized the use of deep learning in clinical settings due to its high computational requirements. Methods: Recent advancements of edge computing devices bring the potential to alleviate this problem. Here we present the implementation of a neuroprosthetic hand with embedded deep learning-based control. The neural decoder is designed based on the recurrent neural network (RNN) architecture and deployed on the NVIDIA Jetson Nano - a compacted yet powerful edge computing platform for deep learning inference. This enables the implementation of the neuroprosthetic hand as a portable and self-contained unit with real-time control of individual finger movements. Results: The proposed system is evaluated on a transradial amputee using peripheral nerve signals (ENG) with implanted intrafascicular microelectrodes. The experiment results demonstrate the system's capabilities of providing robust, high-accuracy (95-99%) and low-latency (50-120 msec) control of individual finger movements in various laboratory and real-world environments. Conclusion: Modern edge computing platforms enable the effective use of deep learning-based neural decoders for neuroprosthesis control as an autonomous system. Significance: This work helps pioneer the deployment of deep neural networks in clinical applications underlying a new class of wearable biomedical devices with embedded artificial intelligence.
Fine-Grained Image Captioning with Global-Local Discriminative Objective
Jul 21, 2020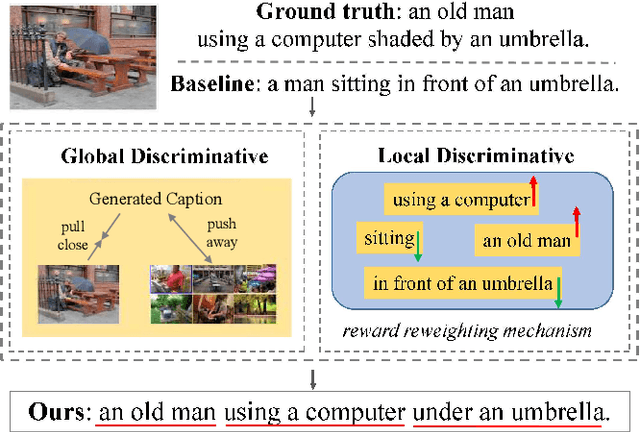
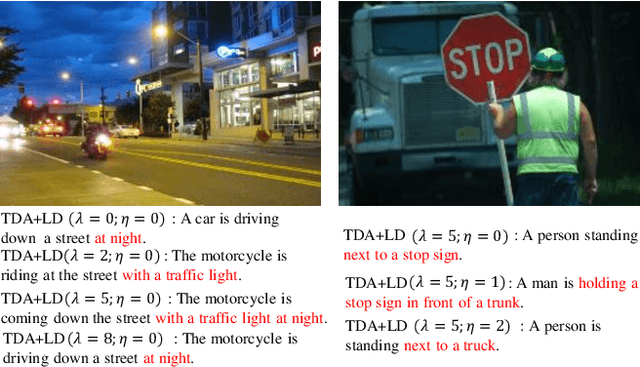

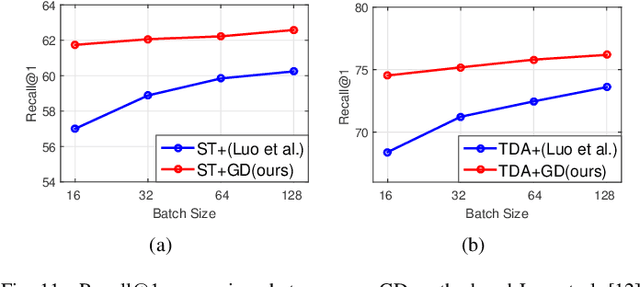
Abstract:Significant progress has been made in recent years in image captioning, an active topic in the fields of vision and language. However, existing methods tend to yield overly general captions and consist of some of the most frequent words/phrases, resulting in inaccurate and indistinguishable descriptions (see Figure 1). This is primarily due to (i) the conservative characteristic of traditional training objectives that drives the model to generate correct but hardly discriminative captions for similar images and (ii) the uneven word distribution of the ground-truth captions, which encourages generating highly frequent words/phrases while suppressing the less frequent but more concrete ones. In this work, we propose a novel global-local discriminative objective that is formulated on top of a reference model to facilitate generating fine-grained descriptive captions. Specifically, from a global perspective, we design a novel global discriminative constraint that pulls the generated sentence to better discern the corresponding image from all others in the entire dataset. From the local perspective, a local discriminative constraint is proposed to increase attention such that it emphasizes the less frequent but more concrete words/phrases, thus facilitating the generation of captions that better describe the visual details of the given images. We evaluate the proposed method on the widely used MS-COCO dataset, where it outperforms the baseline methods by a sizable margin and achieves competitive performance over existing leading approaches. We also conduct self-retrieval experiments to demonstrate the discriminability of the proposed method.
MSD: Multi-Self-Distillation Learning via Multi-classifiers within Deep Neural Networks
Dec 02, 2019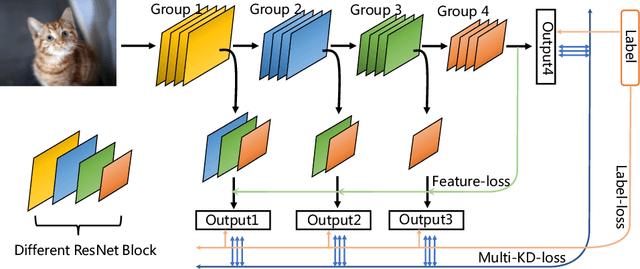
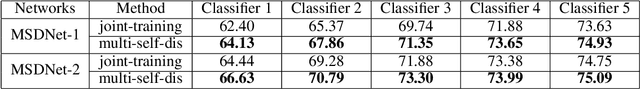
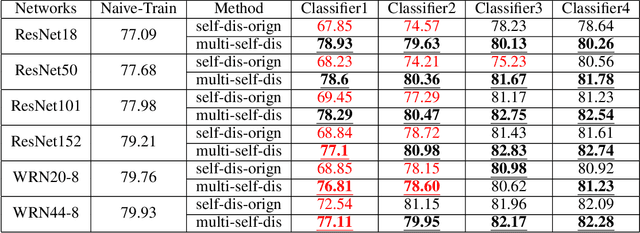
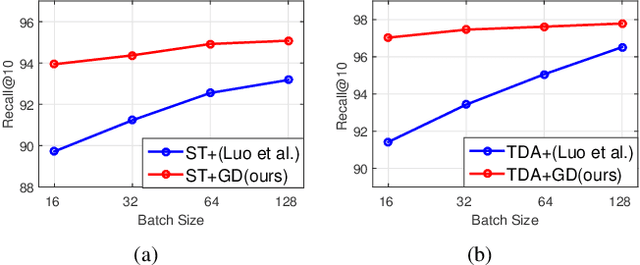
Abstract:As the development of neural networks, more and more deep neural networks are adopted in various tasks, such as image classification. However, as the huge computational overhead, these networks could not be applied on mobile devices or other low latency scenes. To address this dilemma, multi-classifier convolutional network is proposed to allow faster inference via early classifiers with the corresponding classifiers. These networks utilize sophisticated designing to increase the early classifier accuracy. However, naively training the multi-classifier network could hurt the performance (accuracy) of deep neural networks as early classifiers throughout interfere with the feature generation process. In this paper, we propose a general training framework named multi-self-distillation learning (MSD), which mining knowledge of different classifiers within the same network and increase every classifier accuracy. Our approach can be applied not only to multi-classifier networks, but also modern CNNs (e.g., ResNet Series) augmented with additional side branch classifiers. We use sampling-based branch augmentation technique to transform a single-classifier network into a multi-classifier network. This reduces the gap of capacity between different classifiers, and improves the effectiveness of applying MSD. Our experiments show that MSD improves the accuracy of various networks: enhancing the accuracy of every classifier significantly for existing multi-classifier network (MSDNet), improving vanilla single-classifier networks with internal classifiers with high accuracy, while also improving the final accuracy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge