Zhen Qian
Decadal sink-source shifts of forest aboveground carbon since 1988
Jun 13, 2025Abstract:As enduring carbon sinks, forest ecosystems are vital to the terrestrial carbon cycle and help moderate global warming. However, the long-term dynamics of aboveground carbon (AGC) in forests and their sink-source transitions remain highly uncertain, owing to changing disturbance regimes and inconsistencies in observations, data processing, and analysis methods. Here, we derive reliable, harmonized AGC stocks and fluxes in global forests from 1988 to 2021 at high spatial resolution by integrating multi-source satellite observations with probabilistic deep learning models. Our approach simultaneously estimates AGC and associated uncertainties, showing high reliability across space and time. We find that, although global forests remained an AGC sink of 6.2 PgC over 30 years, moist tropical forests shifted to a substantial AGC source between 2001 and 2010 and, together with boreal forests, transitioned toward a source in the 2011-2021 period. Temperate, dry tropical and subtropical forests generally exhibited increasing AGC stocks, although Europe and Australia became sources after 2011. Regionally, pronounced sink-to-source transitions occurred in tropical forests over the past three decades. The interannual relationship between global atmospheric CO2 growth rates and tropical AGC flux variability became increasingly negative, reaching Pearson's r = -0.63 (p < 0.05) in the most recent decade. In the Brazilian Amazon, the contribution of deforested regions to AGC losses declined from 60% in 1989-2000 to 13% in 2011-2021, while the share from untouched areas increased from 33% to 76%. Our findings suggest a growing role of tropical forest AGC in modulating variability in the terrestrial carbon cycle, with anthropogenic climate change potentially contributing increasingly to AGC changes, particularly in previously untouched areas.
Teaching Large Language Models Number-Focused Headline Generation With Key Element Rationales
Feb 05, 2025Abstract:Number-focused headline generation is a summarization task requiring both high textual quality and precise numerical accuracy, which poses a unique challenge for Large Language Models (LLMs). Existing studies in the literature focus only on either textual quality or numerical reasoning and thus are inadequate to address this challenge. In this paper, we propose a novel chain-of-thought framework for using rationales comprising key elements of the Topic, Entities, and Numerical reasoning (TEN) in news articles to enhance the capability for LLMs to generate topic-aligned high-quality texts with precise numerical accuracy. Specifically, a teacher LLM is employed to generate TEN rationales as supervision data, which are then used to teach and fine-tune a student LLM. Our approach teaches the student LLM automatic generation of rationales with enhanced capability for numerical reasoning and topic-aligned numerical headline generation. Experiments show that our approach achieves superior performance in both textual quality and numerical accuracy.
Reconstruct Spine CT from Biplanar X-Rays via Diffusion Learning
Aug 21, 2024


Abstract:Intraoperative CT imaging serves as a crucial resource for surgical guidance; however, it may not always be readily accessible or practical to implement. In scenarios where CT imaging is not an option, reconstructing CT scans from X-rays can offer a viable alternative. In this paper, we introduce an innovative method for 3D CT reconstruction utilizing biplanar X-rays. Distinct from previous research that relies on conventional image generation techniques, our approach leverages a conditional diffusion process to tackle the task of reconstruction. More precisely, we employ a diffusion-based probabilistic model trained to produce 3D CT images based on orthogonal biplanar X-rays. To improve the structural integrity of the reconstructed images, we incorporate a novel projection loss function. Experimental results validate that our proposed method surpasses existing state-of-the-art benchmarks in both visual image quality and multiple evaluative metrics. Specifically, our technique achieves a higher Structural Similarity Index (SSIM) of 0.83, a relative increase of 10\%, and a lower Fr\'echet Inception Distance (FID) of 83.43, which represents a relative decrease of 25\%.
HYDEN: Hyperbolic Density Representations for Medical Images and Reports
Aug 20, 2024Abstract:In light of the inherent entailment relations between images and text, hyperbolic point vector embeddings, leveraging the hierarchical modeling advantages of hyperbolic space, have been utilized for visual semantic representation learning. However, point vector embedding approaches fail to address the issue of semantic uncertainty, where an image may have multiple interpretations, and text may refer to different images, a phenomenon particularly prevalent in the medical domain. Therefor, we propose \textbf{HYDEN}, a novel hyperbolic density embedding based image-text representation learning approach tailored for specific medical domain data. This method integrates text-aware local features alongside global features from images, mapping image-text features to density features in hyperbolic space via using hyperbolic pseudo-Gaussian distributions. An encapsulation loss function is employed to model the partial order relations between image-text density distributions. Experimental results demonstrate the interpretability of our approach and its superior performance compared to the baseline methods across various zero-shot tasks and different datasets.
Coarse-Fine View Attention Alignment-Based GAN for CT Reconstruction from Biplanar X-Rays
Aug 19, 2024


Abstract:For surgical planning and intra-operation imaging, CT reconstruction using X-ray images can potentially be an important alternative when CT imaging is not available or not feasible. In this paper, we aim to use biplanar X-rays to reconstruct a 3D CT image, because biplanar X-rays convey richer information than single-view X-rays and are more commonly used by surgeons. Different from previous studies in which the two X-ray views were treated indifferently when fusing the cross-view data, we propose a novel attention-informed coarse-to-fine cross-view fusion method to combine the features extracted from the orthogonal biplanar views. This method consists of a view attention alignment sub-module and a fine-distillation sub-module that are designed to work together to highlight the unique or complementary information from each of the views. Experiments have demonstrated the superiority of our proposed method over the SOTA methods.
DiffuX2CT: Diffusion Learning to Reconstruct CT Images from Biplanar X-Rays
Jul 18, 2024

Abstract:Computed tomography (CT) is widely utilized in clinical settings because it delivers detailed 3D images of the human body. However, performing CT scans is not always feasible due to radiation exposure and limitations in certain surgical environments. As an alternative, reconstructing CT images from ultra-sparse X-rays offers a valuable solution and has gained significant interest in scientific research and medical applications. However, it presents great challenges as it is inherently an ill-posed problem, often compromised by artifacts resulting from overlapping structures in X-ray images. In this paper, we propose DiffuX2CT, which models CT reconstruction from orthogonal biplanar X-rays as a conditional diffusion process. DiffuX2CT is established with a 3D global coherence denoising model with a new, implicit conditioning mechanism. We realize the conditioning mechanism by a newly designed tri-plane decoupling generator and an implicit neural decoder. By doing so, DiffuX2CT achieves structure-controllable reconstruction, which enables 3D structural information to be recovered from 2D X-rays, therefore producing faithful textures in CT images. As an extra contribution, we collect a real-world lumbar CT dataset, called LumbarV, as a new benchmark to verify the clinical significance and performance of CT reconstruction from X-rays. Extensive experiments on this dataset and three more publicly available datasets demonstrate the effectiveness of our proposal.
Multi-task deep learning for large-scale building detail extraction from high-resolution satellite imagery
Oct 29, 2023



Abstract:Understanding urban dynamics and promoting sustainable development requires comprehensive insights about buildings. While geospatial artificial intelligence has advanced the extraction of such details from Earth observational data, existing methods often suffer from computational inefficiencies and inconsistencies when compiling unified building-related datasets for practical applications. To bridge this gap, we introduce the Multi-task Building Refiner (MT-BR), an adaptable neural network tailored for simultaneous extraction of spatial and attributional building details from high-resolution satellite imagery, exemplified by building rooftops, urban functional types, and roof architectural types. Notably, MT-BR can be fine-tuned to incorporate additional building details, extending its applicability. For large-scale applications, we devise a novel spatial sampling scheme that strategically selects limited but representative image samples. This process optimizes both the spatial distribution of samples and the urban environmental characteristics they contain, thus enhancing extraction effectiveness while curtailing data preparation expenditures. We further enhance MT-BR's predictive performance and generalization capabilities through the integration of advanced augmentation techniques. Our quantitative results highlight the efficacy of the proposed methods. Specifically, networks trained with datasets curated via our sampling method demonstrate improved predictive accuracy relative to those using alternative sampling approaches, with no alterations to network architecture. Moreover, MT-BR consistently outperforms other state-of-the-art methods in extracting building details across various metrics. The real-world practicality is also demonstrated in an application across Shanghai, generating a unified dataset that encompasses both the spatial and attributional details of buildings.
Computation Offloading for Edge Computing in RIS-Assisted Symbiotic Radio Systems
May 29, 2023



Abstract:In the paper, we investigate the coordination process of sensing and computation offloading in a reconfigurable intelligent surface (RIS)-aided base station (BS)-centric symbiotic radio (SR) systems. Specifically, the Internet-of-Things (IoT) devices first sense data from environment and then tackle the data locally or offload the data to BS for remote computing, while RISs are leveraged to enhance the quality of blocked channels and also act as IoT devices to transmit its sensed data. To explore the mechanism of cooperative sensing and computation offloading in this system, we aim at maximizing the total completed sensed bits of all users and RISs by jointly optimizing the time allocation parameter, the passive beamforming at each RIS, the transmit beamforming at BS, and the energy partition parameters for all users subject to the size of sensed data, energy supply and given time cycle. The formulated nonconvex problem is tightly coupled by the time allocation parameter and involves the mathematical expectations, which cannot be solved straightly. We use Monte Carlo and fractional programming methods to transform the nonconvex objective function and then propose an alternating optimization-based algorithm to find an approximate solution with guaranteed convergence. Numerical results show that the RIS-aided SR system outperforms other benchmarks in sensing. Furthermore, with the aid of RIS, the channel and system performance can be significantly improved.
Test-Time Training for Deformable Multi-Scale Image Registration
Mar 25, 2021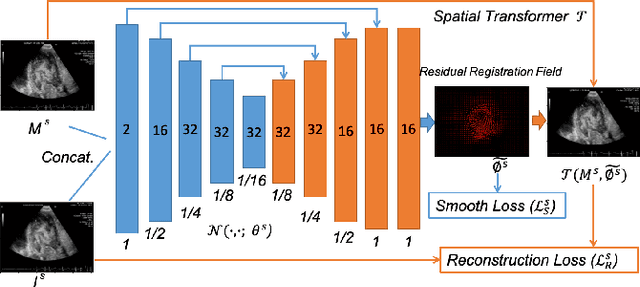
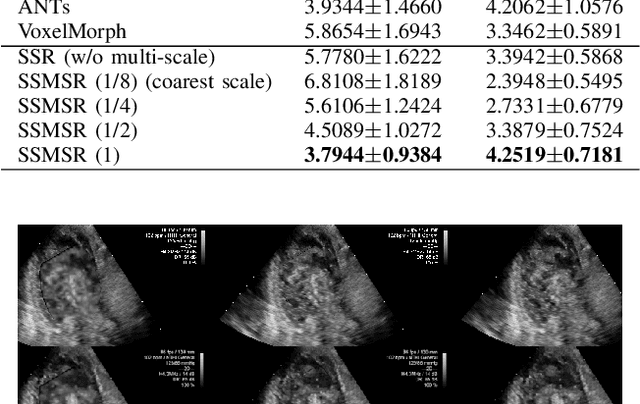
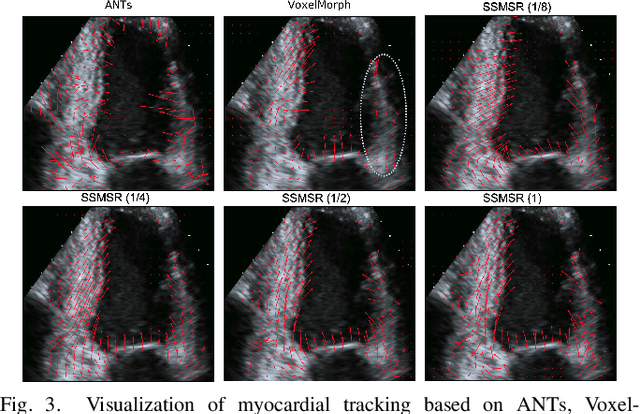
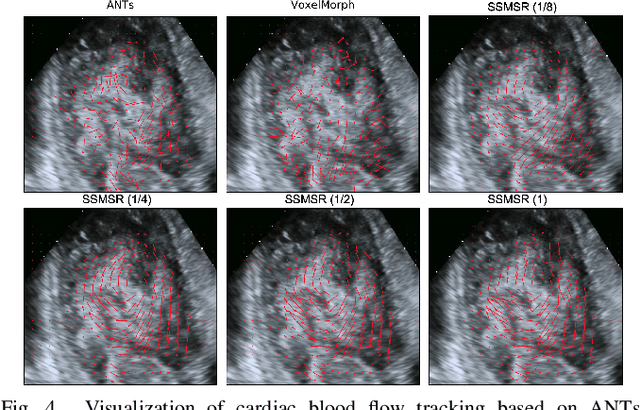
Abstract:Registration is a fundamental task in medical robotics and is often a crucial step for many downstream tasks such as motion analysis, intra-operative tracking and image segmentation. Popular registration methods such as ANTs and NiftyReg optimize objective functions for each pair of images from scratch, which are time-consuming for 3D and sequential images with complex deformations. Recently, deep learning-based registration approaches such as VoxelMorph have been emerging and achieve competitive performance. In this work, we construct a test-time training for deep deformable image registration to improve the generalization ability of conventional learning-based registration model. We design multi-scale deep networks to consecutively model the residual deformations, which is effective for high variational deformations. Extensive experiments validate the effectiveness of multi-scale deep registration with test-time training based on Dice coefficient for image segmentation and mean square error (MSE), normalized local cross-correlation (NLCC) for tissue dense tracking tasks. Two videos are in https://www.youtube.com/watch?v=NvLrCaqCiAE and https://www.youtube.com/watch?v=pEA6ZmtTNuQ
* ICRA 2021; 8 pages, 4 figures, 2 big tables
Bipartite Distance for Shape-Aware Landmark Detection in Spinal X-Ray Images
May 28, 2020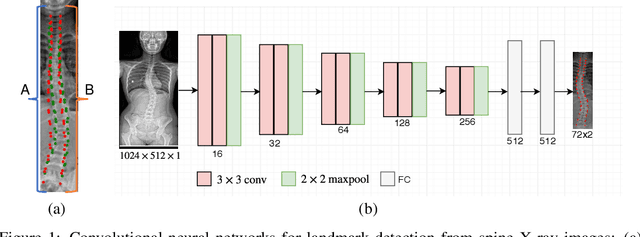
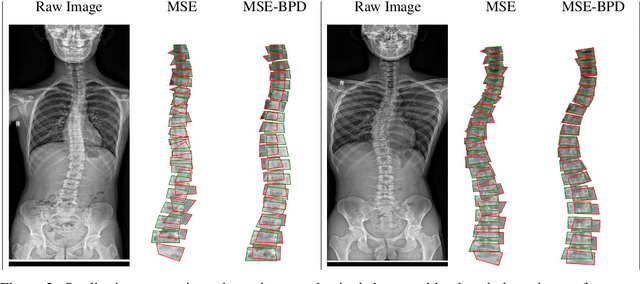
Abstract:Scoliosis is a congenital disease that causes lateral curvature in the spine. Its assessment relies on the identification and localization of vertebrae in spinal X-ray images, conventionally via tedious and time-consuming manual radiographic procedures that are prone to subjectivity and observational variability. Reliability can be improved through the automatic detection and localization of spinal landmarks. To guide a CNN in the learning of spinal shape while detecting landmarks in X-ray images, we propose a novel loss based on a bipartite distance (BPD) measure, and show that it consistently improves landmark detection performance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge