Yufeng Liu
Edge-based Monocular Thermal-Inertial Odometry in Visually Degraded Environments
Oct 18, 2022
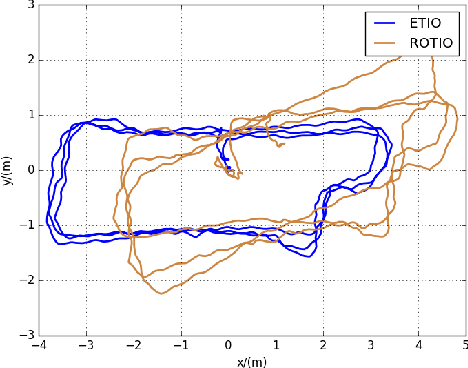
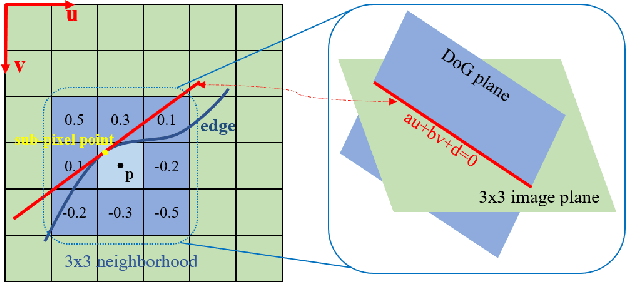
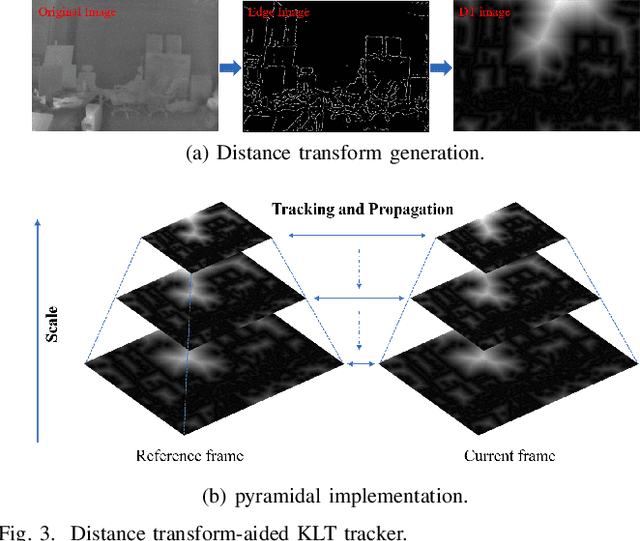
Abstract:State estimation in complex illumination environments based on conventional visual-inertial odometry is a challenging task due to the severe visual degradation of the visual camera. The thermal infrared camera is capable of all-day time and is less affected by illumination variation. However, most existing visual data association algorithms are incompatible because the thermal infrared data contains large noise and low contrast. Motivated by the phenomenon that thermal radiation varies most significantly at the edges of objects, the study proposes an ETIO, which is the first edge-based monocular thermal-inertial odometry for robust localization in visually degraded environments. Instead of the raw image, we utilize the binarized image from edge extraction for pose estimation to overcome the poor thermal infrared image quality. Then, an adaptive feature tracking strategy ADT-KLT is developed for robust data association based on limited edge information and its distance distribution. Finally, a pose graph optimization performs real-time estimation over a sliding window of recent states by combining IMU pre-integration with reprojection error of all edge feature observations. We evaluated the performance of the proposed system on public datasets and real-world experiments and compared it against state-of-the-art methods. The proposed ETIO was verified with the ability to enable accurate and robust localization all-day time.
MobRecon: Mobile-Friendly Hand Mesh Reconstruction from Monocular Image
Dec 06, 2021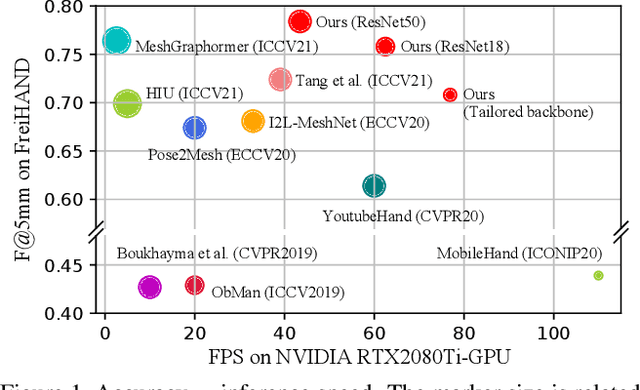
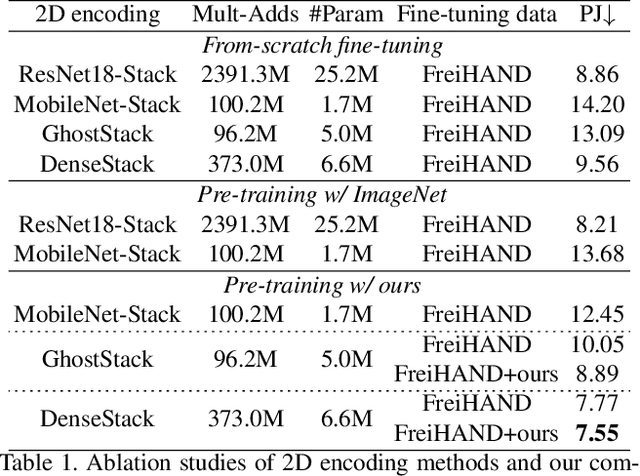
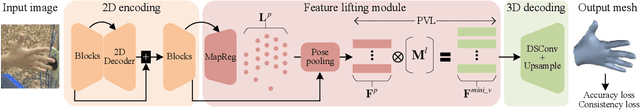
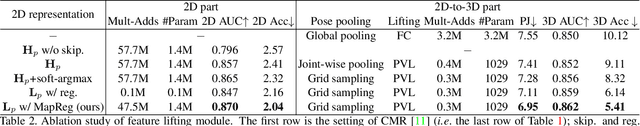
Abstract:In this work, we propose a framework for single-view hand mesh reconstruction, which can simultaneously achieve high reconstruction accuracy, fast inference speed, and temporal coherence. Specifically, for 2D encoding, we propose lightweight yet effective stacked structures. Regarding 3D decoding, we provide an efficient graph operator, namely depth-separable spiral convolution. Moreover, we present a novel feature lifting module for bridging the gap between 2D and 3D representations. This module starts with a map-based position regression (MapReg) block to integrate the merits of both heatmap encoding and position regression paradigms to improve 2D accuracy and temporal coherence. Furthermore, MapReg is followed by pose pooling and pose-to-vertex lifting approaches, which transform 2D pose encodings to semantic features of 3D vertices. Overall, our hand reconstruction framework, called MobRecon, comprises affordable computational costs and miniature model size, which reaches a high inference speed of 83FPS on Apple A14 CPU. Extensive experiments on popular datasets such as FreiHAND, RHD, and HO3Dv2 demonstrate that our MobRecon achieves superior performance on reconstruction accuracy and temporal coherence. Our code is publicly available at https://github.com/SeanChenxy/HandMesh.
Rejoinder: Learning Optimal Distributionally Robust Individualized Treatment Rules
Oct 17, 2021Abstract:We thank the opportunity offered by editors for this discussion and the discussants for their insightful comments and thoughtful contributions. We also want to congratulate Kallus (2020) for his inspiring work in improving the efficiency of policy learning by retargeting. Motivated from the discussion in Dukes and Vansteelandt (2020), we first point out interesting connections and distinctions between our work and Kallus (2020) in Section 1. In particular, the assumptions and sources of variation for consideration in these two papers lead to different research problems with different scopes and focuses. In Section 2, following the discussions in Li et al. (2020); Liang and Zhao (2020), we also consider the efficient policy evaluation problem when we have some data from the testing distribution available at the training stage. We show that under the assumption that the sample sizes from training and testing are growing in the same order, efficient value function estimates can deliver competitive performance. We further show some connections of these estimates with existing literature. However, when the growth of testing sample size available for training is in a slower order, efficient value function estimates may not perform well anymore. In contrast, the requirement of the testing sample size for DRITR is not as strong as that of efficient policy evaluation using the combined data. Finally, we highlight the general applicability and usefulness of DRITR in Section 3.
Efficient Learning of Optimal Individualized Treatment Rules for Heteroscedastic or Misspecified Treatment-Free Effect Models
Sep 06, 2021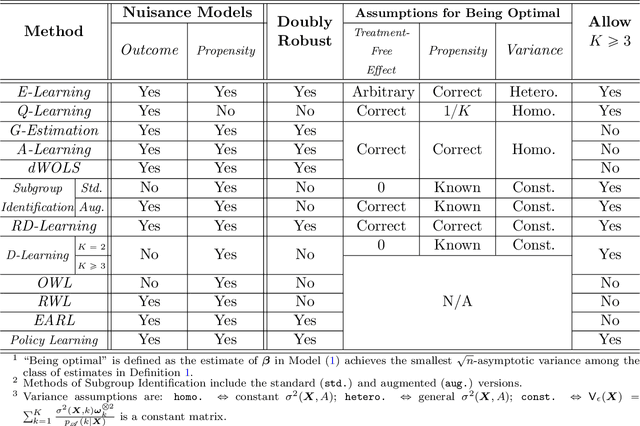
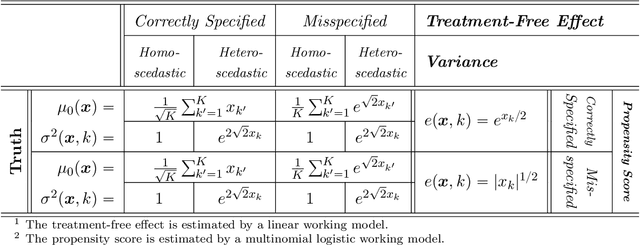
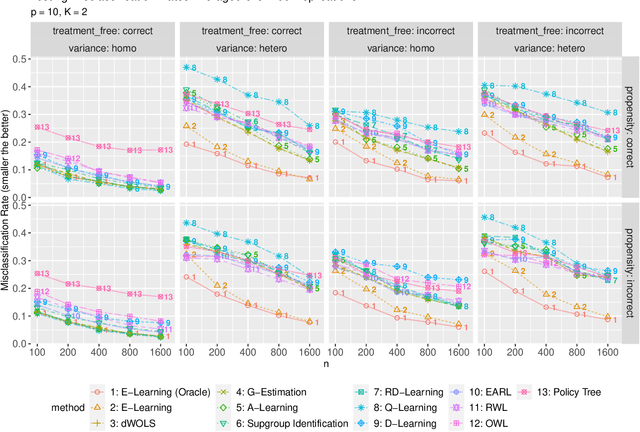

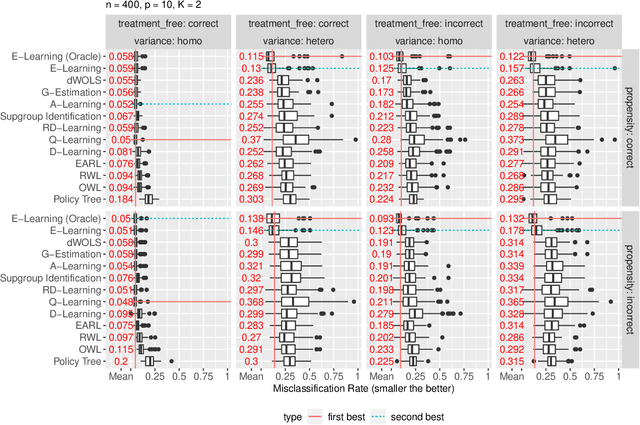
Abstract:Recent development in data-driven decision science has seen great advances in individualized decision making. Given data with individual covariates, treatment assignments and outcomes, researchers can search for the optimal individualized treatment rule (ITR) that maximizes the expected outcome. Existing methods typically require initial estimation of some nuisance models. The double robustness property that can protect from misspecification of either the treatment-free effect or the propensity score has been widely advocated. However, when model misspecification exists, a doubly robust estimate can be consistent but may suffer from downgraded efficiency. Other than potential misspecified nuisance models, most existing methods do not account for the potential problem when the variance of outcome is heterogeneous among covariates and treatment. We observe that such heteroscedasticity can greatly affect the estimation efficiency of the optimal ITR. In this paper, we demonstrate that the consequences of misspecified treatment-free effect and heteroscedasticity can be unified as a covariate-treatment dependent variance of residuals. To improve efficiency of the estimated ITR, we propose an Efficient Learning (E-Learning) framework for finding an optimal ITR in the multi-armed treatment setting. We show that the proposed E-Learning is optimal among a regular class of semiparametric estimates that can allow treatment-free effect misspecification. In our simulation study, E-Learning demonstrates its effectiveness if one of or both misspecified treatment-free effect and heteroscedasticity exist. Our analysis of a Type 2 Diabetes Mellitus (T2DM) observational study also suggests the improved efficiency of E-Learning.
Improving Robustness for Pose Estimation via Stable Heatmap Regression
May 08, 2021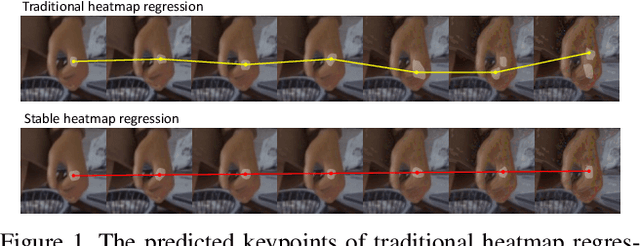

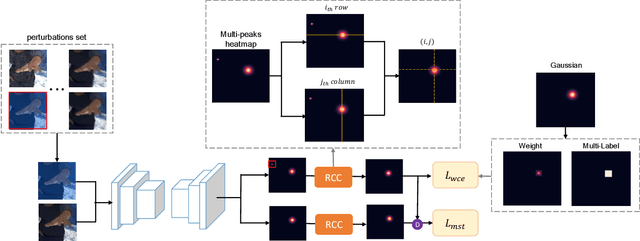

Abstract:Deep learning methods have achieved excellent performance in pose estimation, but the lack of robustness causes the keypoints to change drastically between similar images. In view of this problem, a stable heatmap regression method is proposed to alleviate network vulnerability to small perturbations. We utilize the correlation between different rows and columns in a heatmap to alleviate the multi-peaks problem, and design a highly differentiated heatmap regression to make a keypoint discriminative from surrounding points. A maximum stability training loss is used to simplify the optimization difficulty when minimizing the prediction gap of two similar images. The proposed method achieves a significant advance in robustness over state-of-the-art approaches on two benchmark datasets and maintains high performance.
Camera-Space Hand Mesh Recovery via Semantic Aggregation and Adaptive 2D-1D Registration
Mar 31, 2021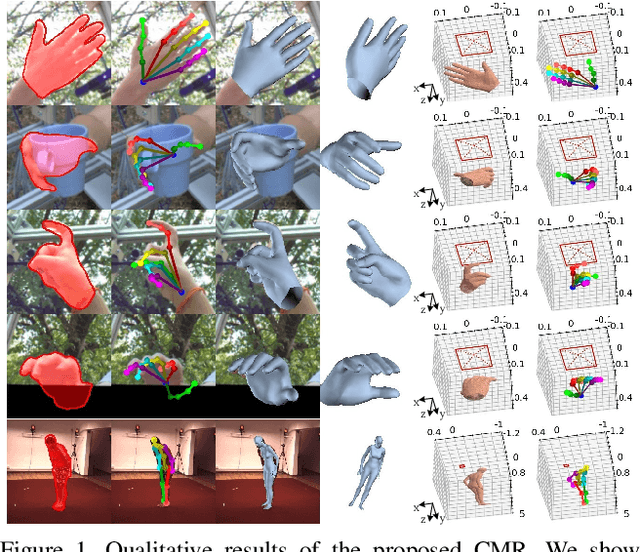
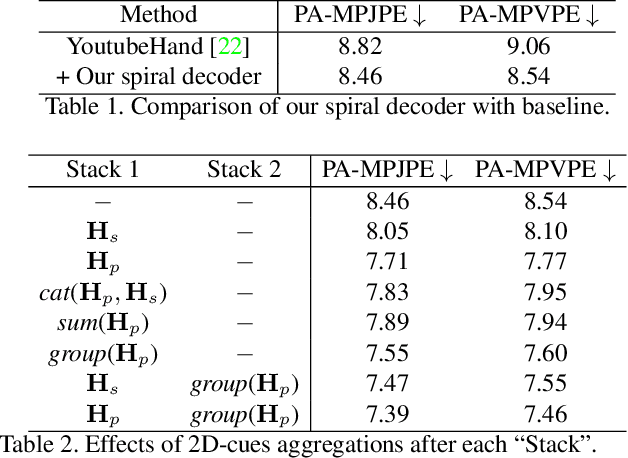
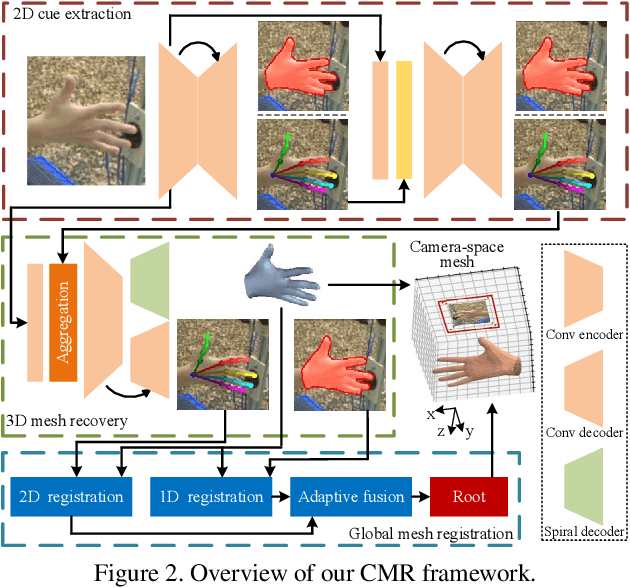
Abstract:Recent years have witnessed significant progress in 3D hand mesh recovery. Nevertheless, because of the intrinsic 2D-to-3D ambiguity, recovering camera-space 3D information from a single RGB image remains challenging. To tackle this problem, we divide camera-space mesh recovery into two sub-tasks, i.e., root-relative mesh recovery and root recovery. First, joint landmarks and silhouette are extracted from a single input image to provide 2D cues for the 3D tasks. In the root-relative mesh recovery task, we exploit semantic relations among joints to generate a 3D mesh from the extracted 2D cues. Such generated 3D mesh coordinates are expressed relative to a root position, i.e., wrist of the hand. In the root recovery task, the root position is registered to the camera space by aligning the generated 3D mesh back to 2D cues, thereby completing cameraspace 3D mesh recovery. Our pipeline is novel in that (1) it explicitly makes use of known semantic relations among joints and (2) it exploits 1D projections of the silhouette and mesh to achieve robust registration. Extensive experiments on popular datasets such as FreiHAND, RHD, and Human3.6M demonstrate that our approach achieves stateof-the-art performance on both root-relative mesh recovery and root recovery. Our code is publicly available at https://github.com/SeanChenxy/HandMesh.
Regressive Domain Adaptation for Unsupervised Keypoint Detection
Mar 10, 2021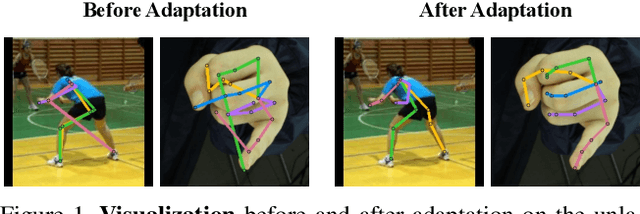
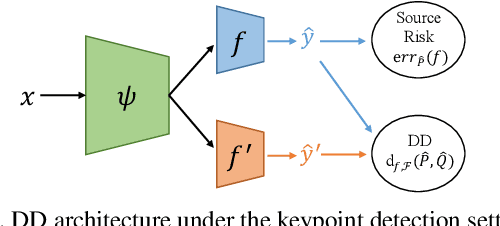
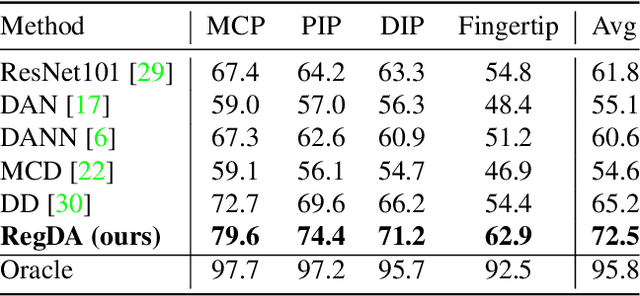
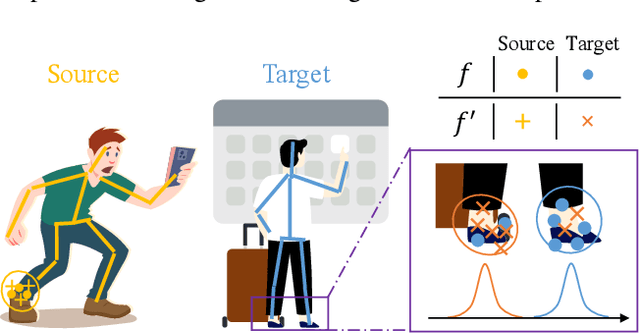
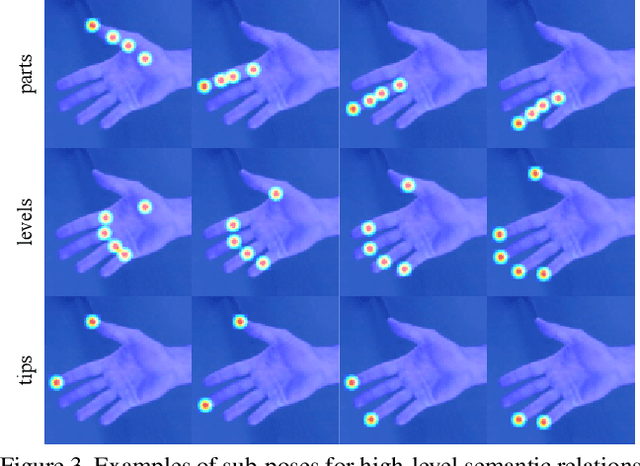
Abstract:Domain adaptation (DA) aims at transferring knowledge from a labeled source domain to an unlabeled target domain. Though many DA theories and algorithms have been proposed, most of them are tailored into classification settings and may fail in regression tasks, especially in the practical keypoint detection task. To tackle this difficult but significant task, we present a method of regressive domain adaptation (RegDA) for unsupervised keypoint detection. Inspired by the latest theoretical work, we first utilize an adversarial regressor to maximize the disparity on the target domain and train a feature generator to minimize this disparity. However, due to the high dimension of the output space, this regressor fails to detect samples that deviate from the support of the source. To overcome this problem, we propose two important ideas. First, based on our observation that the probability density of the output space is sparse, we introduce a spatial probability distribution to describe this sparsity and then use it to guide the learning of the adversarial regressor. Second, to alleviate the optimization difficulty in the high-dimensional space, we innovatively convert the minimax game in the adversarial training to the minimization of two opposite goals. Extensive experiments show that our method brings large improvement by 8% to 11% in terms of PCK on different datasets.
Learning Optimal Distributionally Robust Individualized Treatment Rules
Jun 26, 2020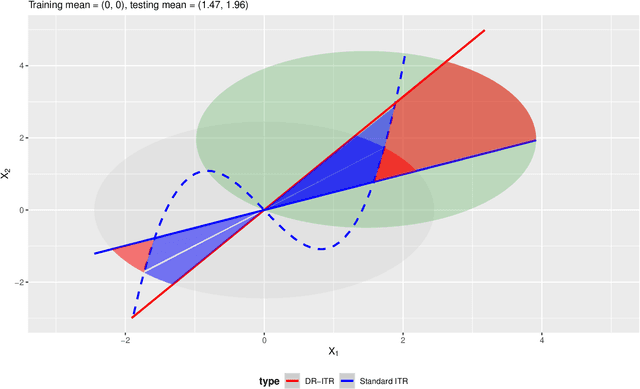
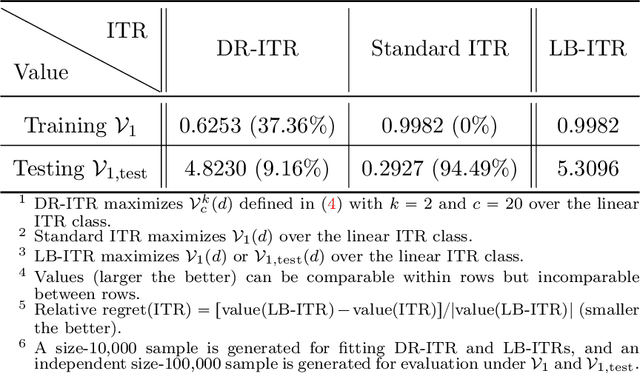
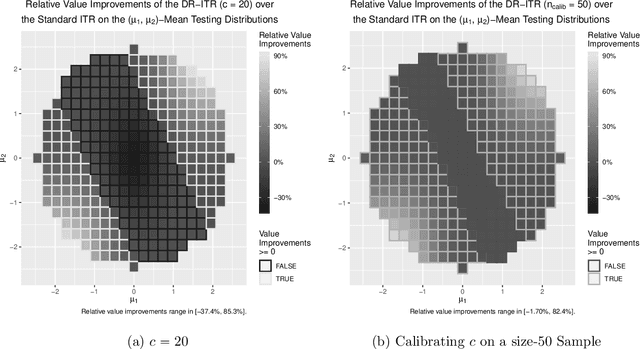
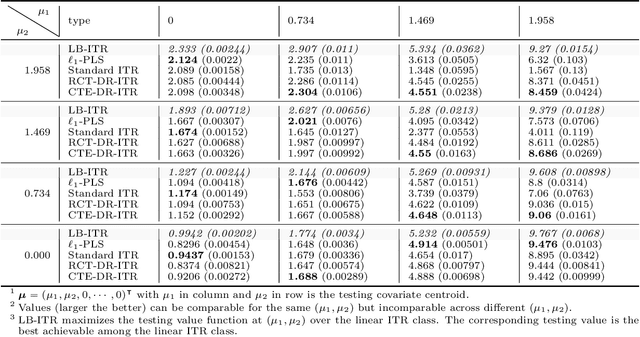
Abstract:Recent development in the data-driven decision science has seen great advances in individualized decision making. Given data with individual covariates, treatment assignments and outcomes, policy makers best individualized treatment rule (ITR) that maximizes the expected outcome, known as the value function. Many existing methods assume that the training and testing distributions are the same. However, the estimated optimal ITR may have poor generalizability when the training and testing distributions are not identical. In this paper, we consider the problem of finding an optimal ITR from a restricted ITR class where there is some unknown covariate changes between the training and testing distributions. We propose a novel distributionally robust ITR (DR-ITR) framework that maximizes the worst-case value function across the values under a set of underlying distributions that are "close" to the training distribution. The resulting DR-ITR can guarantee the performance among all such distributions reasonably well. We further propose a calibrating procedure that tunes the DR-ITR adaptively to a small amount of calibration data from a target population. In this way, the calibrated DR-ITR can be shown to enjoy better generalizability than the standard ITR based on our numerical studies.
High dimensional precision medicine from patient-derived xenografts
Dec 13, 2019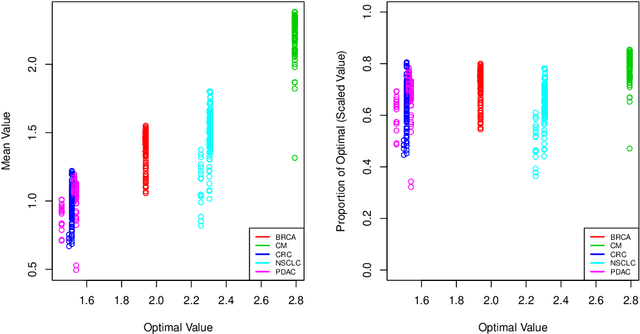

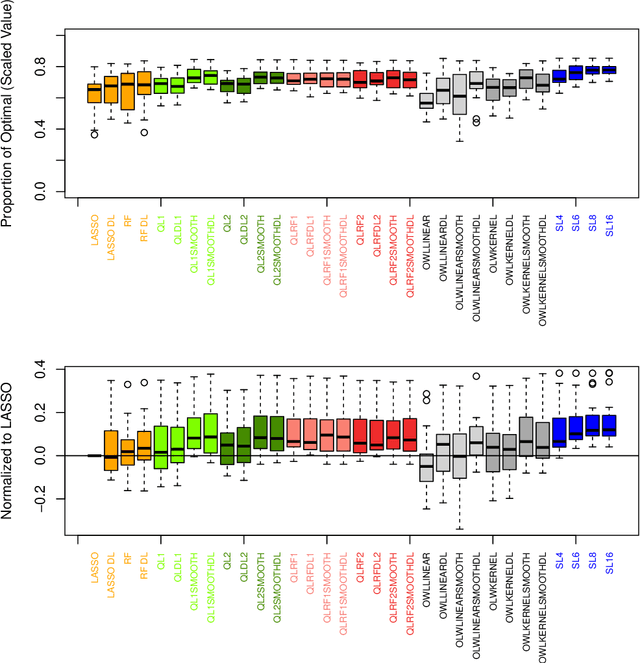
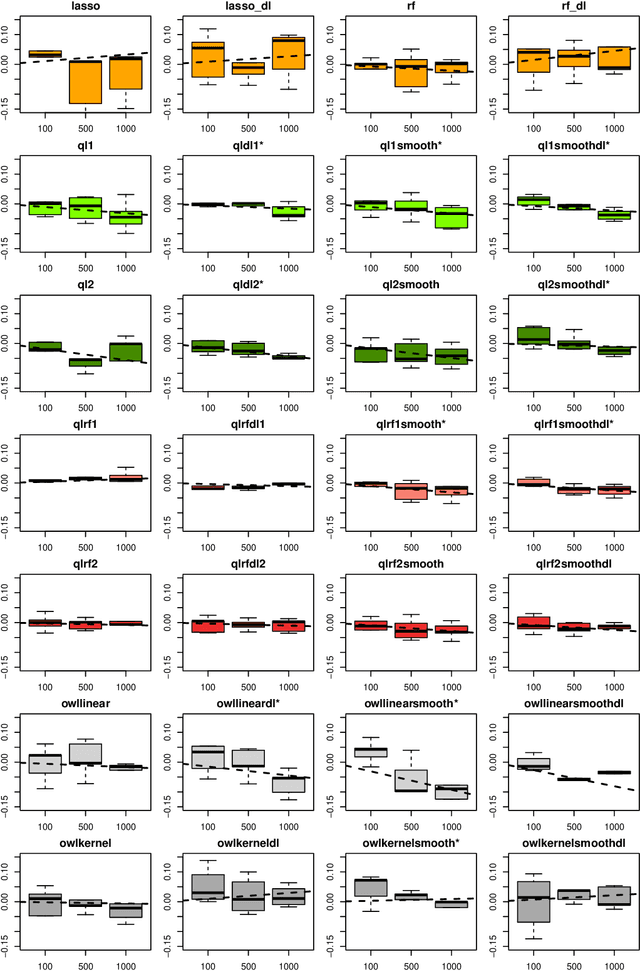
Abstract:The complexity of human cancer often results in significant heterogeneity in response to treatment. Precision medicine offers potential to improve patient outcomes by leveraging this heterogeneity. Individualized treatment rules (ITRs) formalize precision medicine as maps from the patient covariate space into the space of allowable treatments. The optimal ITR is that which maximizes the mean of a clinical outcome in a population of interest. Patient-derived xenograft (PDX) studies permit the evaluation of multiple treatments within a single tumor and thus are ideally suited for estimating optimal ITRs. PDX data are characterized by correlated outcomes, a high-dimensional feature space, and a large number of treatments. Existing methods for estimating optimal ITRs do not take advantage of the unique structure of PDX data or handle the associated challenges well. In this paper, we explore machine learning methods for estimating optimal ITRs from PDX data. We analyze data from a large PDX study to identify biomarkers that are informative for developing personalized treatment recommendations in multiple cancers. We estimate optimal ITRs using regression-based approaches such as Q-learning and direct search methods such as outcome weighted learning. Finally, we implement a superlearner approach to combine a set of estimated ITRs and show that the resulting ITR performs better than any of the input ITRs, mitigating uncertainty regarding user choice of any particular ITR estimation methodology. Our results indicate that PDX data are a valuable resource for developing individualized treatment strategies in oncology.
Statistical Analysis of Stationary Solutions of Coupled Nonconvex Nonsmooth Empirical Risk Minimization
Oct 06, 2019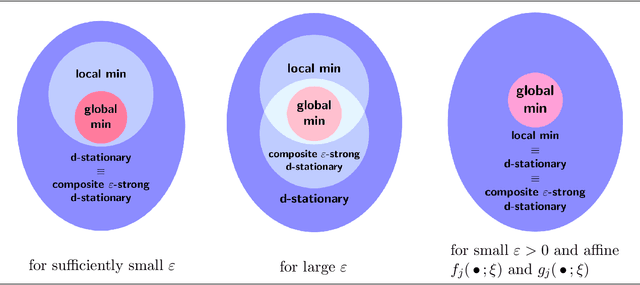
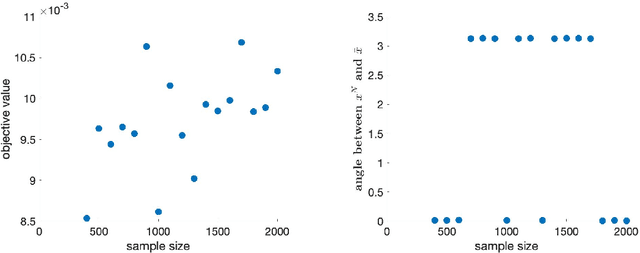
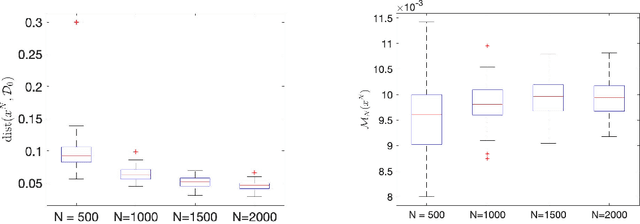
Abstract:This paper has two main goals: (a) establish several statistical properties---consistency, asymptotic distributions, and convergence rates---of stationary solutions and values of a class of coupled nonconvex and nonsmoothempirical risk minimization problems, and (b) validate these properties by a noisy amplitude-based phase retrieval problem, the latter being of much topical interest.Derived from available data via sampling, these empirical risk minimization problems are the computational workhorse of a population risk model which involves the minimization of an expected value of a random functional. When these minimization problems are nonconvex, the computation of their globally optimal solutions is elusive. Together with the fact that the expectation operator cannot be evaluated for general probability distributions, it becomes necessary to justify whether the stationary solutions of the empirical problems are practical approximations of the stationary solution of the population problem. When these two features, general distribution and nonconvexity, are coupled with nondifferentiability that often renders the problems "non-Clarke regular", the task of the justification becomes challenging. Our work aims to address such a challenge within an algorithm-free setting. The resulting analysis is therefore different from the much of the analysis in the recent literature that is based on local search algorithms. Furthermore, supplementing the classical minimizer-centric analysis, our results offer a first step to close the gap between computational optimization and asymptotic analysis of coupled nonconvex nonsmooth statistical estimation problems, expanding the former with statistical properties of the practically obtained solution and providing the latter with a more practical focus pertaining to computational tractability.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge