Yi Guo
Defining Robust Ultrasound Quality Metrics via an Ultrasound Foundation Model
Apr 21, 2026Abstract:Clinicians lack a principled framework to quantify diagnostic utility in ultrasound reconstructions. Existing standards like PSNR and VGG-LPIPS are inadequate, failing to account for modality-specific physics or the structural nuances of acoustic imaging. We close this gap with a TinyUSFM-based evaluation framework featuring two distinct metrics: TinyUSFM-uLPIPS, a full-reference perceptual distance based on multi-layer token relations, and TinyUSFM-NRQ, a deployable no-reference quality score utilizing clean-manifold modeling and worst-region aggregation to detect localized harmful artifacts. We demonstrate that the presented metrics have four unique advantages: 1) Task-linked quality, where TinyUSFM-uLPIPS achieves superior calibration with semantic task damage, accurately reflecting Dice-score drops in segmentation where VGG-based metrics fail; 2) Cross-organ comparability, maintaining stable scoring scales and consistent severity rankings across diverse anatomical sites and domain-shifted data; 3) PSNR-consistent sensitivity, with TinyUSFM-NRQ providing a reliable quality score without ground-truth images that remains consistent with traditional fidelity benchmarks (i.e. PSNR); and 4) Clinical utility, improving the prediction of expert preference from 47.2$\%$ to 72.8$\%$ accuracy and producing super-resolution reconstructions preferred by sonographers. By integrating these advantages into a unified assessment and optimization loop, this work establishes a modality-aligned standard that finally bridges the gap between algorithmic performance and diagnostic utility. https://github.com/sextant-fable/US-Metrics
EchoAgent: Towards Reliable Echocardiography Interpretation with "Eyes","Hands" and "Minds"
Apr 07, 2026Abstract:Reliable interpretation of echocardiography (Echo) is crucial for assessing cardiac function, which demands clinicians to synchronously orchestrate multiple capabilities, including visual observation (eyes), manual measurement (hands), and expert knowledge learning and reasoning (minds). While current task-specific deep-learning approaches and multimodal large language models have demonstrated promise in assisting Echo analysis through automated segmentation or reasoning, they remain focused on restricted skills, i.e., eyes-hands or eyes-minds, thereby limiting clinical reliability and utility. To address these issues, we propose EchoAgent, an agentic system tailored for end-to-end Echo interpretation, which achieves a fully coordinated eyes-hands-minds workflow that learns, observes, operates, and reasons like a cardiac sonographer. First, we introduce an expertise-driven cognition engine where our agent can automatically assimilate credible Echo guidelines into a structured knowledge base, thus constructing an Echo-customized mind. Second, we devise a hierarchical collaboration toolkit to endow EchoAgent with eyes-hands, which can automatically parse Echo video streams, identify cardiac views, perform anatomical segmentation, and quantitative measurement. Third, we integrate the perceived multimodal evidence with the exclusive knowledge base into an orchestrated reasoning hub to conduct explainable inferences. We evaluate EchoAgent on CAMUS and MIMIC-EchoQA datasets, which cover 48 distinct echocardiographic views spanning 14 cardiac anatomical regions. Experimental results show that EchoAgent achieves optimal performance across diverse structure analyses, yielding overall accuracy of up to 80.00%. Importantly, EchoAgent empowers a single system with abilities to learn, observe, operate and reason like an echocardiologist, which holds great promise for reliable Echo interpretation.
PolyFormer: learning efficient reformulations for scalable optimization under complex physical constraints
Mar 09, 2026Abstract:Real-world optimization problems are often constrained by complex physical laws that limit computational scalability. These constraints are inherently tied to complex regions, and thus learning models that incorporate physical and geometric knowledge, i.e., physics-informed machine learning (PIML), offer a promising pathway for efficient solution. Here, we introduce PolyFormer, which opens a new direction for PIML in prescriptive optimization tasks, where physical and geometric knowledge is not merely used to regularize learning models, but to simplify the problems themselves. PolyFormer captures geometric structures behind constraints and transforms them into efficient polytopic reformulations, thereby decoupling problem complexity from solution difficulty and enabling off-the-shelf optimization solvers to efficiently produce feasible solutions with acceptable optimality loss. Through evaluations across three important problems (large-scale resource aggregation, network-constrained optimization, and optimization under uncertainty), PolyFormer achieves computational speedups up to 6,400-fold and memory reductions up to 99.87%, while maintaining solution quality competitive with or superior to state-of-the-art methods. These results demonstrate that PolyFormer provides an efficient and reliable solution for scalable constrained optimization, expanding the scope of PIML to prescriptive tasks in scientific discovery and engineering applications.
ColoDiff: Integrating Dynamic Consistency With Content Awareness for Colonoscopy Video Generation
Feb 26, 2026Abstract:Colonoscopy video generation delivers dynamic, information-rich data critical for diagnosing intestinal diseases, particularly in data-scarce scenarios. High-quality video generation demands temporal consistency and precise control over clinical attributes, but faces challenges from irregular intestinal structures, diverse disease representations, and various imaging modalities. To this end, we propose ColoDiff, a diffusion-based framework that generates dynamic-consistent and content-aware colonoscopy videos, aiming to alleviate data shortage and assist clinical analysis. At the inter-frame level, our TimeStream module decouples temporal dependency from video sequences through a cross-frame tokenization mechanism, enabling intricate dynamic modeling despite irregular intestinal structures. At the intra-frame level, our Content-Aware module incorporates noise-injected embeddings and learnable prototypes to realize precise control over clinical attributes, breaking through the coarse guidance of diffusion models. Additionally, ColoDiff employs a non-Markovian sampling strategy that cuts steps by over 90% for real-time generation. ColoDiff is evaluated across three public datasets and one hospital database, based on both generation metrics and downstream tasks including disease diagnosis, modality discrimination, bowel preparation scoring, and lesion segmentation. Extensive experiments show ColoDiff generates videos with smooth transitions and rich dynamics. ColoDiff presents an effort in controllable colonoscopy video generation, revealing the potential of synthetic videos in complementing authentic representation and mitigating data scarcity in clinical settings.
SHIELD: An Auto-Healing Agentic Defense Framework for LLM Resource Exhaustion Attacks
Jan 27, 2026Abstract:Sponge attacks increasingly threaten LLM systems by inducing excessive computation and DoS. Existing defenses either rely on statistical filters that fail on semantically meaningful attacks or use static LLM-based detectors that struggle to adapt as attack strategies evolve. We introduce SHIELD, a multi-agent, auto-healing defense framework centered on a three-stage Defense Agent that integrates semantic similarity retrieval, pattern matching, and LLM-based reasoning. Two auxiliary agents, a Knowledge Updating Agent and a Prompt Optimization Agent, form a closed self-healing loop, when an attack bypasses detection, the system updates an evolving knowledgebase, and refines defense instructions. Extensive experiments show that SHIELD consistently outperforms perplexity-based and standalone LLM defenses, achieving high F1 scores across both non-semantic and semantic sponge attacks, demonstrating the effectiveness of agentic self-healing against evolving resource-exhaustion threats.
Prompt-Induced Over-Generation as Denial-of-Service: A Black-Box Attack-Side Benchmark
Dec 29, 2025Abstract:Large language models (LLMs) can be driven into over-generation, emitting thousands of tokens before producing an end-of-sequence (EOS) token. This degrades answer quality, inflates latency and cost, and can be weaponized as a denial-of-service (DoS) attack. Recent work has begun to study DoS-style prompt attacks, but typically focuses on a single attack algorithm or assumes white-box access, without an attack-side benchmark that compares prompt-based attackers in a black-box, query-only regime with a known tokenizer. We introduce such a benchmark and study two prompt-only attackers. The first is Evolutionary Over-Generation Prompt Search (EOGen), which searches the token space for prefixes that suppress EOS and induce long continuations. The second is a goal-conditioned reinforcement learning attacker (RL-GOAL) that trains a network to generate prefixes conditioned on a target length. To characterize behavior, we introduce Over-Generation Factor (OGF), the ratio of produced tokens to a model's context window, along with stall and latency summaries. Our evolutionary attacker achieves mean OGF = 1.38 +/- 1.15 and Success@OGF >= 2 of 24.5 percent on Phi-3. RL-GOAL is stronger: across victims it achieves higher mean OGF (up to 2.81 +/- 1.38).
TinyUSFM: Towards Compact and Efficient Ultrasound Foundation Models
Oct 22, 2025Abstract:Foundation models for medical imaging demonstrate superior generalization capabilities across diverse anatomical structures and clinical applications. Their outstanding performance relies on substantial computational resources, limiting deployment in resource-constrained clinical environments. This paper presents TinyUSFM, the first lightweight ultrasound foundation model that maintains superior organ versatility and task adaptability of our large-scale Ultrasound Foundation Model (USFM) through knowledge distillation with strategically curated small datasets, delivering significant computational efficiency without sacrificing performance. Considering the limited capacity and representation ability of lightweight models, we propose a feature-gradient driven coreset selection strategy to curate high-quality compact training data, avoiding training degradation from low-quality redundant images. To preserve the essential spatial and frequency domain characteristics during knowledge transfer, we develop domain-separated masked image modeling assisted consistency-driven dynamic distillation. This novel framework adaptively transfers knowledge from large foundation models by leveraging teacher model consistency across different domain masks, specifically tailored for ultrasound interpretation. For evaluation, we establish the UniUS-Bench, the largest publicly available ultrasound benchmark comprising 8 classification and 10 segmentation datasets across 15 organs. Using only 200K images in distillation, TinyUSFM matches USFM's performance with just 6.36% of parameters and 6.40% of GFLOPs. TinyUSFM significantly outperforms the vanilla model by 9.45% in classification and 7.72% in segmentation, surpassing all state-of-the-art lightweight models, and achieving 84.91% average classification accuracy and 85.78% average segmentation Dice score across diverse medical devices and centers.
SplitMeanFlow: Interval Splitting Consistency in Few-Step Generative Modeling
Jul 22, 2025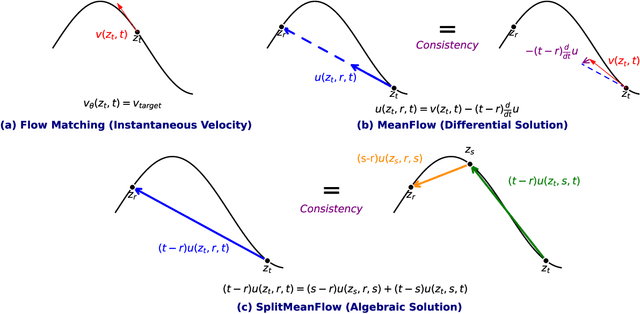
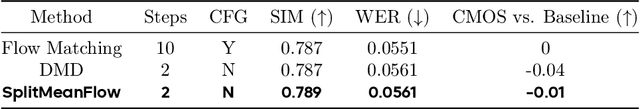
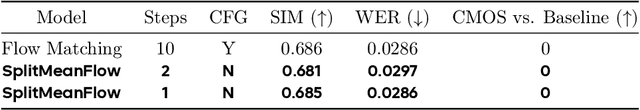
Abstract:Generative models like Flow Matching have achieved state-of-the-art performance but are often hindered by a computationally expensive iterative sampling process. To address this, recent work has focused on few-step or one-step generation by learning the average velocity field, which directly maps noise to data. MeanFlow, a leading method in this area, learns this field by enforcing a differential identity that connects the average and instantaneous velocities. In this work, we argue that this differential formulation is a limiting special case of a more fundamental principle. We return to the first principles of average velocity and leverage the additivity property of definite integrals. This leads us to derive a novel, purely algebraic identity we term Interval Splitting Consistency. This identity establishes a self-referential relationship for the average velocity field across different time intervals without resorting to any differential operators. Based on this principle, we introduce SplitMeanFlow, a new training framework that enforces this algebraic consistency directly as a learning objective. We formally prove that the differential identity at the core of MeanFlow is recovered by taking the limit of our algebraic consistency as the interval split becomes infinitesimal. This establishes SplitMeanFlow as a direct and more general foundation for learning average velocity fields. From a practical standpoint, our algebraic approach is significantly more efficient, as it eliminates the need for JVP computations, resulting in simpler implementation, more stable training, and broader hardware compatibility. One-step and two-step SplitMeanFlow models have been successfully deployed in large-scale speech synthesis products (such as Doubao), achieving speedups of 20x.
Scale-Invariance Drives Convergence in AI and Brain Representations
Jun 13, 2025Abstract:Despite variations in architecture and pretraining strategies, recent studies indicate that large-scale AI models often converge toward similar internal representations that also align with neural activity. We propose that scale-invariance, a fundamental structural principle in natural systems, is a key driver of this convergence. In this work, we propose a multi-scale analytical framework to quantify two core aspects of scale-invariance in AI representations: dimensional stability and structural similarity across scales. We further investigate whether these properties can predict alignment performance with functional Magnetic Resonance Imaging (fMRI) responses in the visual cortex. Our analysis reveals that embeddings with more consistent dimension and higher structural similarity across scales align better with fMRI data. Furthermore, we find that the manifold structure of fMRI data is more concentrated, with most features dissipating at smaller scales. Embeddings with similar scale patterns align more closely with fMRI data. We also show that larger pretraining datasets and the inclusion of language modalities enhance the scale-invariance properties of embeddings, further improving neural alignment. Our findings indicate that scale-invariance is a fundamental structural principle that bridges artificial and biological representations, providing a new framework for evaluating the structural quality of human-like AI systems.
A Topic Modeling Analysis of Stigma Dimensions, Social, and Related Behavioral Circumstances in Clinical Notes Among Patients with HIV
Jun 10, 2025



Abstract:Objective: To characterize stigma dimensions, social, and related behavioral circumstances in people living with HIV (PLWHs) seeking care, using natural language processing methods applied to a large collection of electronic health record (EHR) clinical notes from a large integrated health system in the southeast United States. Methods: We identified 9,140 cohort of PLWHs from the UF Health IDR and performed topic modeling analysis using Latent Dirichlet Allocation (LDA) to uncover stigma dimensions, social, and related behavioral circumstances. Domain experts created a seed list of HIV-related stigma keywords, then applied a snowball strategy to iteratively review notes for additional terms until saturation was reached. To identify more target topics, we tested three keyword-based filtering strategies. Domain experts manually reviewed the detected topics using the prevalent terms and key discussion topics. Word frequency analysis was used to highlight the prevalent terms associated with each topic. In addition, we conducted topic variation analysis among subgroups to examine differences across age and sex-specific demographics. Results and Conclusion: Topic modeling on sentences containing at least one keyword uncovered a wide range of topic themes associated with HIV-related stigma, social, and related behaviors circumstances, including "Mental Health Concern and Stigma", "Social Support and Engagement", "Limited Healthcare Access and Severe Illness", "Treatment Refusal and Isolation" and so on. Topic variation analysis across age subgroups revealed differences. Extracting and understanding the HIV-related stigma dimensions, social, and related behavioral circumstances from EHR clinical notes enables scalable, time-efficient assessment, overcoming the limitations of traditional questionnaires and improving patient outcomes.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge