Wen Zhang
Zhejiang University
Prompting Disentangled Embeddings for Knowledge Graph Completion with Pre-trained Language Model
Dec 04, 2023



Abstract:Both graph structures and textual information play a critical role in Knowledge Graph Completion (KGC). With the success of Pre-trained Language Models (PLMs) such as BERT, they have been applied for text encoding for KGC. However, the current methods mostly prefer to fine-tune PLMs, leading to huge training costs and limited scalability to larger PLMs. In contrast, we propose to utilize prompts and perform KGC on a frozen PLM with only the prompts trained. Accordingly, we propose a new KGC method named PDKGC with two prompts -- a hard task prompt which is to adapt the KGC task to the PLM pre-training task of token prediction, and a disentangled structure prompt which learns disentangled graph representation so as to enable the PLM to combine more relevant structure knowledge with the text information. With the two prompts, PDKGC builds a textual predictor and a structural predictor, respectively, and their combination leads to more comprehensive entity prediction. Solid evaluation on two widely used KGC datasets has shown that PDKGC often outperforms the baselines including the state-of-the-art, and its components are all effective. Our codes and data are available at https://github.com/genggengcss/PDKGC.
Knowledgeable Preference Alignment for LLMs in Domain-specific Question Answering
Nov 11, 2023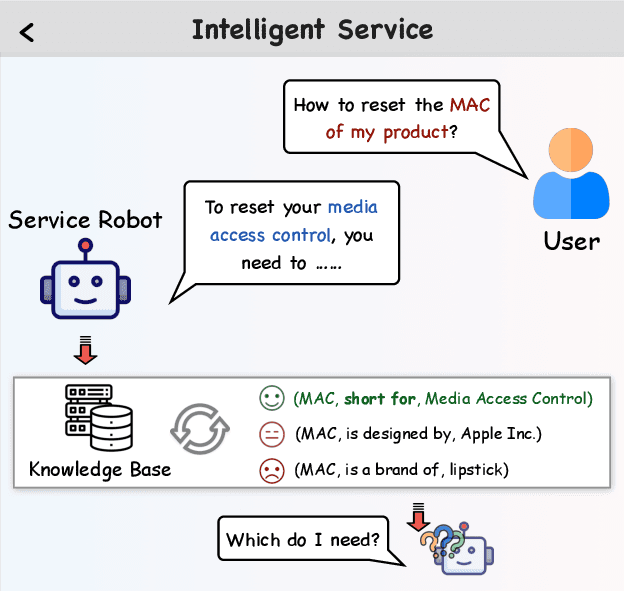
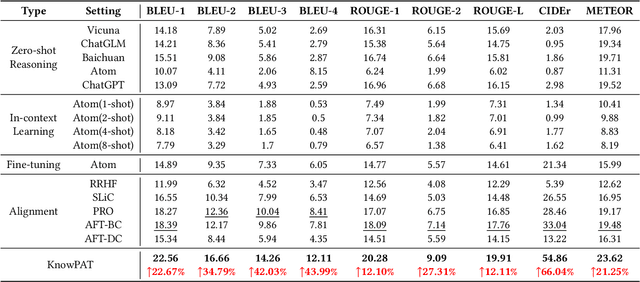
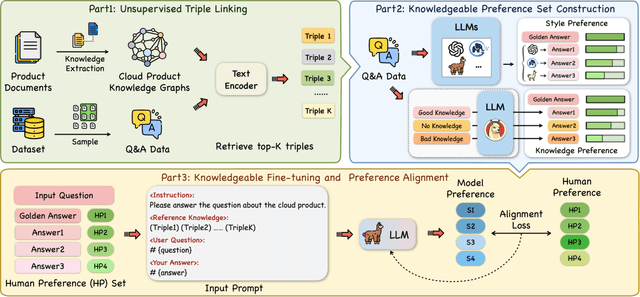
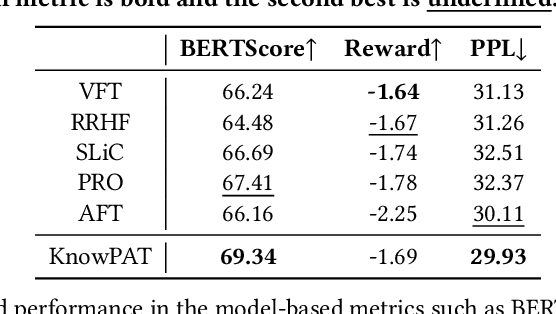
Abstract:Recently, the development of large language models (LLMs) has attracted wide attention in academia and industry. Deploying LLMs to real scenarios is one of the key directions in the current Internet industry. In this paper, we present a novel pipeline to apply LLMs for domain-specific question answering (QA) that incorporates domain knowledge graphs (KGs), addressing an important direction of LLM application. As a real-world application, the content generated by LLMs should be user-friendly to serve the customers. Additionally, the model needs to utilize domain knowledge properly to generate reliable answers. These two issues are the two major difficulties in the LLM application as vanilla fine-tuning can not adequately address them. We think both requirements can be unified as the model preference problem that needs to align with humans to achieve practical application. Thus, we introduce Knowledgeable Preference AlignmenT (KnowPAT), which constructs two kinds of preference set called style preference set and knowledge preference set respectively to tackle the two issues. Besides, we design a new alignment objective to align the LLM preference with human preference, aiming to train a better LLM for real-scenario domain-specific QA to generate reliable and user-friendly answers. Adequate experiments and comprehensive with 15 baseline methods demonstrate that our KnowPAT is an outperforming pipeline for real-scenario domain-specific QA with LLMs. Our code is open-source at https://github.com/zjukg/KnowPAT.
Sound field reconstruction using neural processes with dynamic kernels
Nov 09, 2023Abstract:Accurately representing the sound field with the high spatial resolution is critical for immersive and interactive sound field reproduction technology. To minimize experimental effort, data-driven methods have been proposed to estimate sound fields from a small number of discrete observations. In particular, kernel-based methods using Gaussian Processes (GPs) with a covariance function to model spatial correlations have been used for sound field reconstruction. However, these methods have limitations due to the fixed kernels having limited expressiveness, requiring manual identification of optimal kernels for different sound fields. In this work, we propose a new approach that parameterizes GPs using a deep neural network based on Neural Processes (NPs) to reconstruct the magnitude of the sound field. This method has the advantage of dynamically learning kernels from simulated data using an attention mechanism, allowing for greater flexibility and adaptability to the acoustic properties of the sound field. Numerical experiments demonstrate that our proposed approach outperforms current methods in reconstructing accuracy, providing a promising alternative for sound field reconstruction.
CBSiMT: Mitigating Hallucination in Simultaneous Machine Translation with Weighted Prefix-to-Prefix Training
Nov 07, 2023



Abstract:Simultaneous machine translation (SiMT) is a challenging task that requires starting translation before the full source sentence is available. Prefix-to-prefix framework is often applied to SiMT, which learns to predict target tokens using only a partial source prefix. However, due to the word order difference between languages, misaligned prefix pairs would make SiMT models suffer from serious hallucination problems, i.e. target outputs that are unfaithful to source inputs. Such problems can not only produce target tokens that are not supported by the source prefix, but also hinder generating the correct translation by receiving more source words. In this work, we propose a Confidence-Based Simultaneous Machine Translation (CBSiMT) framework, which uses model confidence to perceive hallucination tokens and mitigates their negative impact with weighted prefix-to-prefix training. Specifically, token-level and sentence-level weights are calculated based on model confidence and acted on the loss function. We explicitly quantify the faithfulness of the generated target tokens using the token-level weight, and employ the sentence-level weight to alleviate the disturbance of sentence pairs with serious word order differences on the model. Experimental results on MuST-C English-to-Chinese and WMT15 German-to-English SiMT tasks demonstrate that our method can consistently improve translation quality at most latency regimes, with up to 2 BLEU scores improvement at low latency.
Reliable Academic Conference Question Answering: A Study Based on Large Language Model
Oct 19, 2023



Abstract:The rapid growth of computer science has led to a proliferation of research presented at academic conferences, fostering global scholarly communication. Researchers consistently seek accurate, current information about these events at all stages. This data surge necessitates an intelligent question-answering system to efficiently address researchers' queries and ensure awareness of the latest advancements. The information of conferences is usually published on their official website, organized in a semi-structured way with a lot of text. To address this need, we have developed the ConferenceQA dataset for 7 diverse academic conferences with human annotations. Firstly, we employ a combination of manual and automated methods to organize academic conference data in a semi-structured JSON format. Subsequently, we annotate nearly 100 question-answer pairs for each conference. Each pair is classified into four different dimensions. To ensure the reliability of the data, we manually annotate the source of each answer. In light of recent advancements, Large Language Models (LLMs) have demonstrated impressive performance in various NLP tasks. They have demonstrated impressive capabilities in information-seeking question answering after instruction fine-tuning, and as such, we present our conference QA study based on LLM. Due to hallucination and outdated knowledge of LLMs, we adopt retrieval based methods to enhance LLMs' question-answering abilities. We have proposed a structure-aware retrieval method, specifically designed to leverage inherent structural information during the retrieval process. Empirical validation on the ConferenceQA dataset has demonstrated the effectiveness of this method. The dataset and code are readily accessible on https://github.com/zjukg/ConferenceQA.
Negative Sampling with Adaptive Denoising Mixup for Knowledge Graph Embedding
Oct 15, 2023



Abstract:Knowledge graph embedding (KGE) aims to map entities and relations of a knowledge graph (KG) into a low-dimensional and dense vector space via contrasting the positive and negative triples. In the training process of KGEs, negative sampling is essential to find high-quality negative triples since KGs only contain positive triples. Most existing negative sampling methods assume that non-existent triples with high scores are high-quality negative triples. However, negative triples sampled by these methods are likely to contain noise. Specifically, they ignore that non-existent triples with high scores might also be true facts due to the incompleteness of KGs, which are usually called false negative triples. To alleviate the above issue, we propose an easily pluggable denoising mixup method called DeMix, which generates high-quality triples by refining sampled negative triples in a self-supervised manner. Given a sampled unlabeled triple, DeMix firstly classifies it into a marginal pseudo-negative triple or a negative triple based on the judgment of the KGE model itself. Secondly, it selects an appropriate mixup partner for the current triple to synthesize a partially positive or a harder negative triple. Experimental results on the knowledge graph completion task show that the proposed DeMix is superior to other negative sampling techniques, ensuring corresponding KGEs a faster convergence and better link prediction results.
Making Large Language Models Perform Better in Knowledge Graph Completion
Oct 10, 2023Abstract:Large language model (LLM) based knowledge graph completion (KGC) aims to predict the missing triples in the KGs with LLMs and enrich the KGs to become better web infrastructure, which can benefit a lot of web-based automatic services. However, research about LLM-based KGC is limited and lacks effective utilization of LLM's inference capabilities, which ignores the important structural information in KGs and prevents LLMs from acquiring accurate factual knowledge. In this paper, we discuss how to incorporate the helpful KG structural information into the LLMs, aiming to achieve structrual-aware reasoning in the LLMs. We first transfer the existing LLM paradigms to structural-aware settings and further propose a knowledge prefix adapter (KoPA) to fulfill this stated goal. KoPA employs structural embedding pre-training to capture the structural information of entities and relations in the KG. Then KoPA informs the LLMs of the knowledge prefix adapter which projects the structural embeddings into the textual space and obtains virtual knowledge tokens as a prefix of the input prompt. We conduct comprehensive experiments on these structural-aware LLM-based KGC methods and provide an in-depth analysis comparing how the introduction of structural information would be better for LLM's knowledge reasoning ability. Our code is released at https://github.com/zjukg/KoPA.
A Comprehensive Study on Knowledge Graph Embedding over Relational Patterns Based on Rule Learning
Aug 15, 2023Abstract:Knowledge Graph Embedding (KGE) has proven to be an effective approach to solving the Knowledge Graph Completion (KGC) task. Relational patterns which refer to relations with specific semantics exhibiting graph patterns are an important factor in the performance of KGE models. Though KGE models' capabilities are analyzed over different relational patterns in theory and a rough connection between better relational patterns modeling and better performance of KGC has been built, a comprehensive quantitative analysis on KGE models over relational patterns remains absent so it is uncertain how the theoretical support of KGE to a relational pattern contributes to the performance of triples associated to such a relational pattern. To address this challenge, we evaluate the performance of 7 KGE models over 4 common relational patterns on 2 benchmarks, then conduct an analysis in theory, entity frequency, and part-to-whole three aspects and get some counterintuitive conclusions. Finally, we introduce a training-free method Score-based Patterns Adaptation (SPA) to enhance KGE models' performance over various relational patterns. This approach is simple yet effective and can be applied to KGE models without additional training. Our experimental results demonstrate that our method generally enhances performance over specific relational patterns. Our source code is available from GitHub at https://github.com/zjukg/Comprehensive-Study-over-Relational-Patterns.
MACO: A Modality Adversarial and Contrastive Framework for Modality-missing Multi-modal Knowledge Graph Completion
Aug 13, 2023Abstract:Recent years have seen significant advancements in multi-modal knowledge graph completion (MMKGC). MMKGC enhances knowledge graph completion (KGC) by integrating multi-modal entity information, thereby facilitating the discovery of unobserved triples in the large-scale knowledge graphs (KGs). Nevertheless, existing methods emphasize the design of elegant KGC models to facilitate modality interaction, neglecting the real-life problem of missing modalities in KGs. The missing modality information impedes modal interaction, consequently undermining the model's performance. In this paper, we propose a modality adversarial and contrastive framework (MACO) to solve the modality-missing problem in MMKGC. MACO trains a generator and discriminator adversarially to generate missing modality features that can be incorporated into the MMKGC model. Meanwhile, we design a cross-modal contrastive loss to improve the performance of the generator. Experiments on public benchmarks with further explorations demonstrate that MACO could achieve state-of-the-art results and serve as a versatile framework to bolster various MMKGC models. Our code and benchmark data are available at https://github.com/zjukg/MACO.
Large Language Models and Knowledge Graphs: Opportunities and Challenges
Aug 11, 2023Abstract:Large Language Models (LLMs) have taken Knowledge Representation -- and the world -- by storm. This inflection point marks a shift from explicit knowledge representation to a renewed focus on the hybrid representation of both explicit knowledge and parametric knowledge. In this position paper, we will discuss some of the common debate points within the community on LLMs (parametric knowledge) and Knowledge Graphs (explicit knowledge) and speculate on opportunities and visions that the renewed focus brings, as well as related research topics and challenges.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge