Thomas Unterthiner
Aligning Machine and Human Visual Representations across Abstraction Levels
Sep 10, 2024



Abstract:Deep neural networks have achieved success across a wide range of applications, including as models of human behavior in vision tasks. However, neural network training and human learning differ in fundamental ways, and neural networks often fail to generalize as robustly as humans do, raising questions regarding the similarity of their underlying representations. What is missing for modern learning systems to exhibit more human-like behavior? We highlight a key misalignment between vision models and humans: whereas human conceptual knowledge is hierarchically organized from fine- to coarse-scale distinctions, model representations do not accurately capture all these levels of abstraction. To address this misalignment, we first train a teacher model to imitate human judgments, then transfer human-like structure from its representations into pretrained state-of-the-art vision foundation models. These human-aligned models more accurately approximate human behavior and uncertainty across a wide range of similarity tasks, including a new dataset of human judgments spanning multiple levels of semantic abstractions. They also perform better on a diverse set of machine learning tasks, increasing generalization and out-of-distribution robustness. Thus, infusing neural networks with additional human knowledge yields a best-of-both-worlds representation that is both more consistent with human cognition and more practically useful, thus paving the way toward more robust, interpretable, and human-like artificial intelligence systems.
PaliGemma: A versatile 3B VLM for transfer
Jul 10, 2024



Abstract:PaliGemma is an open Vision-Language Model (VLM) that is based on the SigLIP-So400m vision encoder and the Gemma-2B language model. It is trained to be a versatile and broadly knowledgeable base model that is effective to transfer. It achieves strong performance on a wide variety of open-world tasks. We evaluate PaliGemma on almost 40 diverse tasks including standard VLM benchmarks, but also more specialized tasks such as remote-sensing and segmentation.
Getting aligned on representational alignment
Nov 02, 2023



Abstract:Biological and artificial information processing systems form representations that they can use to categorize, reason, plan, navigate, and make decisions. How can we measure the extent to which the representations formed by these diverse systems agree? Do similarities in representations then translate into similar behavior? How can a system's representations be modified to better match those of another system? These questions pertaining to the study of representational alignment are at the heart of some of the most active research areas in cognitive science, neuroscience, and machine learning. For example, cognitive scientists measure the representational alignment of multiple individuals to identify shared cognitive priors, neuroscientists align fMRI responses from multiple individuals into a shared representational space for group-level analyses, and ML researchers distill knowledge from teacher models into student models by increasing their alignment. Unfortunately, there is limited knowledge transfer between research communities interested in representational alignment, so progress in one field often ends up being rediscovered independently in another. Thus, greater cross-field communication would be advantageous. To improve communication between these fields, we propose a unifying framework that can serve as a common language between researchers studying representational alignment. We survey the literature from all three fields and demonstrate how prior work fits into this framework. Finally, we lay out open problems in representational alignment where progress can benefit all three of these fields. We hope that our work can catalyze cross-disciplinary collaboration and accelerate progress for all communities studying and developing information processing systems. We note that this is a working paper and encourage readers to reach out with their suggestions for future revisions.
Set Learning for Accurate and Calibrated Models
Jul 10, 2023



Abstract:Model overconfidence and poor calibration are common in machine learning and difficult to account for when applying standard empirical risk minimization. In this work, we propose a novel method to alleviate these problems that we call odd-$k$-out learning (OKO), which minimizes the cross-entropy error for sets rather than for single examples. This naturally allows the model to capture correlations across data examples and achieves both better accuracy and calibration, especially in limited training data and class-imbalanced regimes. Perhaps surprisingly, OKO often yields better calibration even when training with hard labels and dropping any additional calibration parameter tuning, such as temperature scaling. We provide theoretical justification, establishing that OKO naturally yields better calibration, and provide extensive experimental analyses that corroborate our theoretical findings. We emphasize that OKO is a general framework that can be easily adapted to many settings and the trained model can be applied to single examples at inference time, without introducing significant run-time overhead or architecture changes.
Accurate Machine Learned Quantum-Mechanical Force Fields for Biomolecular Simulations
May 17, 2022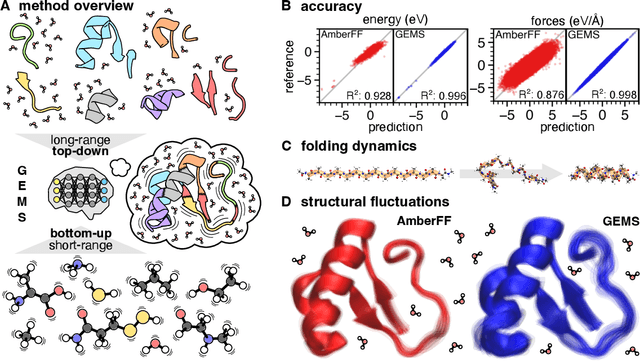
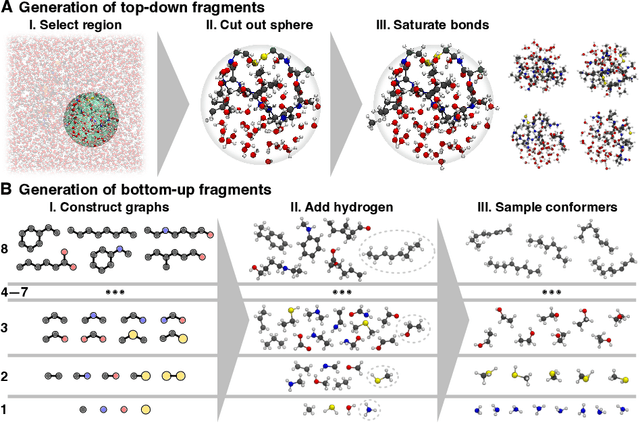
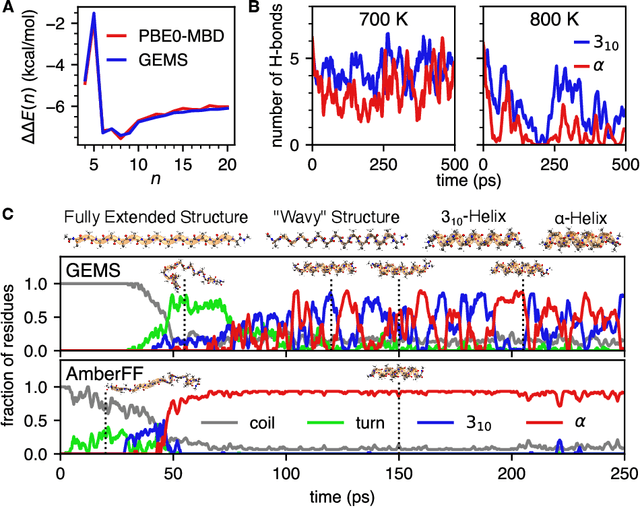
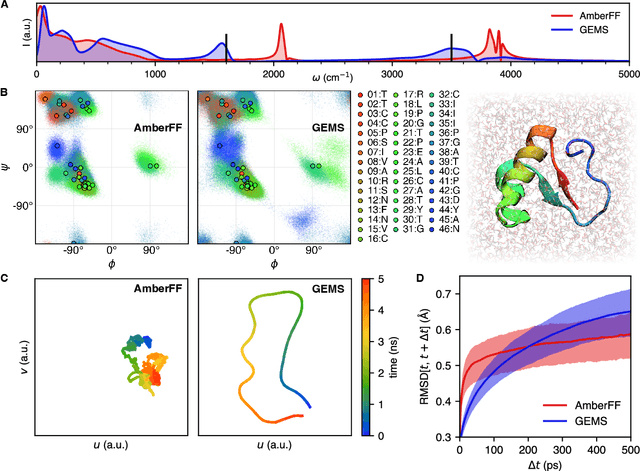
Abstract:Molecular dynamics (MD) simulations allow atomistic insights into chemical and biological processes. Accurate MD simulations require computationally demanding quantum-mechanical calculations, being practically limited to short timescales and few atoms. For larger systems, efficient, but much less reliable empirical force fields are used. Recently, machine learned force fields (MLFFs) emerged as an alternative means to execute MD simulations, offering similar accuracy as ab initio methods at orders-of-magnitude speedup. Until now, MLFFs mainly capture short-range interactions in small molecules or periodic materials, due to the increased complexity of constructing models and obtaining reliable reference data for large molecules, where long-ranged many-body effects become important. This work proposes a general approach to constructing accurate MLFFs for large-scale molecular simulations (GEMS) by training on "bottom-up" and "top-down" molecular fragments of varying size, from which the relevant physicochemical interactions can be learned. GEMS is applied to study the dynamics of alanine-based peptides and the 46-residue protein crambin in aqueous solution, allowing nanosecond-scale MD simulations of >25k atoms at essentially ab initio quality. Our findings suggest that structural motifs in peptides and proteins are more flexible than previously thought, indicating that simulations at ab initio accuracy might be necessary to understand dynamic biomolecular processes such as protein (mis)folding, drug-protein binding, or allosteric regulation.
GradMax: Growing Neural Networks using Gradient Information
Jan 13, 2022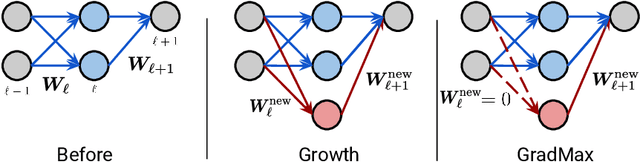
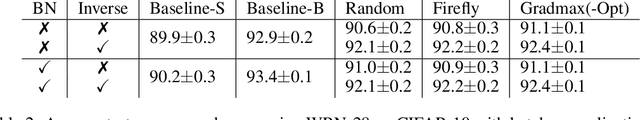

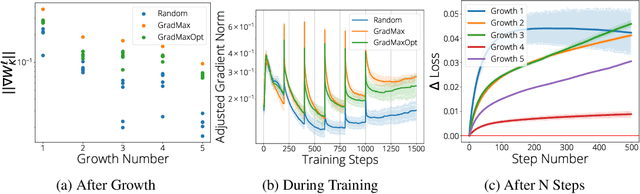

Abstract:The architecture and the parameters of neural networks are often optimized independently, which requires costly retraining of the parameters whenever the architecture is modified. In this work we instead focus on growing the architecture without requiring costly retraining. We present a method that adds new neurons during training without impacting what is already learned, while improving the training dynamics. We achieve the latter by maximizing the gradients of the new weights and find the optimal initialization efficiently by means of the singular value decomposition (SVD). We call this technique Gradient Maximizing Growth (GradMax) and demonstrate its effectiveness in variety of vision tasks and architectures.
Do Vision Transformers See Like Convolutional Neural Networks?
Aug 19, 2021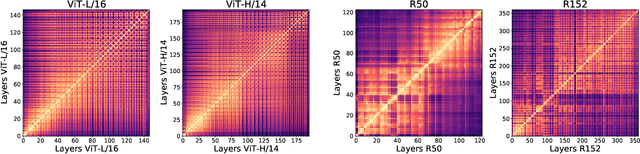
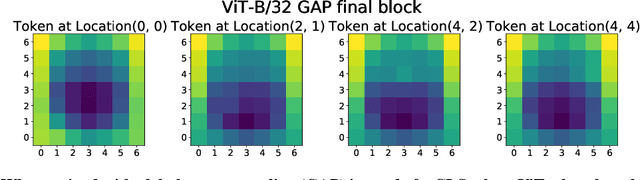
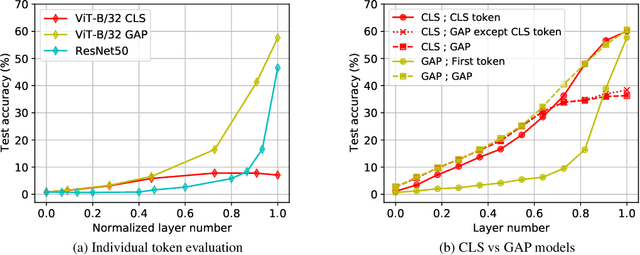
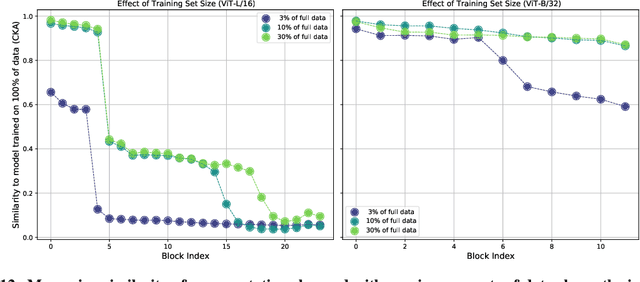
Abstract:Convolutional neural networks (CNNs) have so far been the de-facto model for visual data. Recent work has shown that (Vision) Transformer models (ViT) can achieve comparable or even superior performance on image classification tasks. This raises a central question: how are Vision Transformers solving these tasks? Are they acting like convolutional networks, or learning entirely different visual representations? Analyzing the internal representation structure of ViTs and CNNs on image classification benchmarks, we find striking differences between the two architectures, such as ViT having more uniform representations across all layers. We explore how these differences arise, finding crucial roles played by self-attention, which enables early aggregation of global information, and ViT residual connections, which strongly propagate features from lower to higher layers. We study the ramifications for spatial localization, demonstrating ViTs successfully preserve input spatial information, with noticeable effects from different classification methods. Finally, we study the effect of (pretraining) dataset scale on intermediate features and transfer learning, and conclude with a discussion on connections to new architectures such as the MLP-Mixer.
MLP-Mixer: An all-MLP Architecture for Vision
May 17, 2021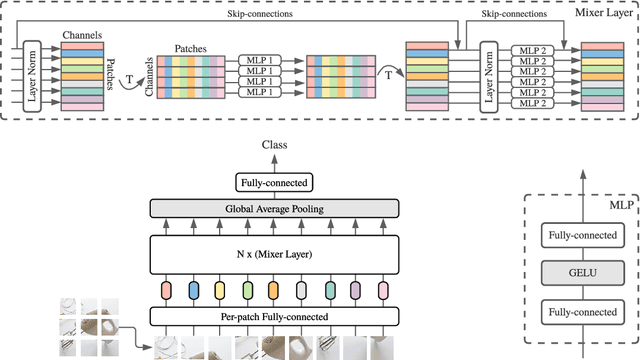
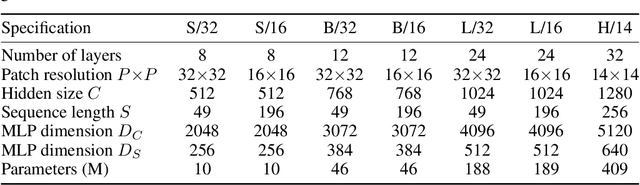
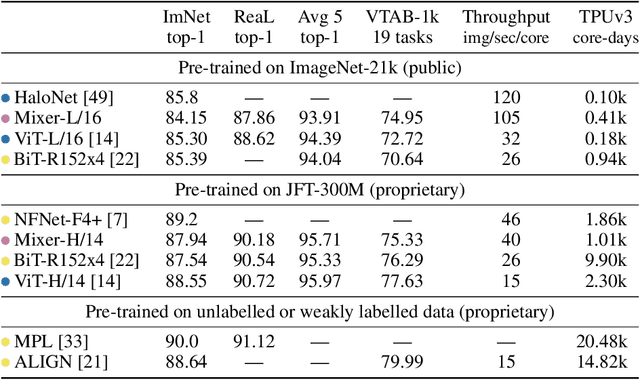
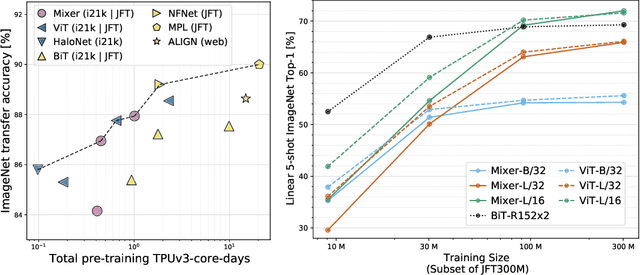
Abstract:Convolutional Neural Networks (CNNs) are the go-to model for computer vision. Recently, attention-based networks, such as the Vision Transformer, have also become popular. In this paper we show that while convolutions and attention are both sufficient for good performance, neither of them are necessary. We present MLP-Mixer, an architecture based exclusively on multi-layer perceptrons (MLPs). MLP-Mixer contains two types of layers: one with MLPs applied independently to image patches (i.e. "mixing" the per-location features), and one with MLPs applied across patches (i.e. "mixing" spatial information). When trained on large datasets, or with modern regularization schemes, MLP-Mixer attains competitive scores on image classification benchmarks, with pre-training and inference cost comparable to state-of-the-art models. We hope that these results spark further research beyond the realms of well established CNNs and Transformers.
Differentiable Patch Selection for Image Recognition
Apr 07, 2021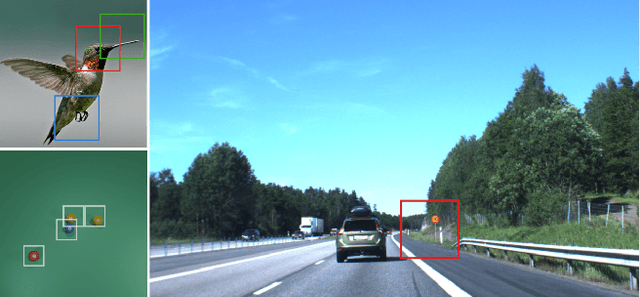
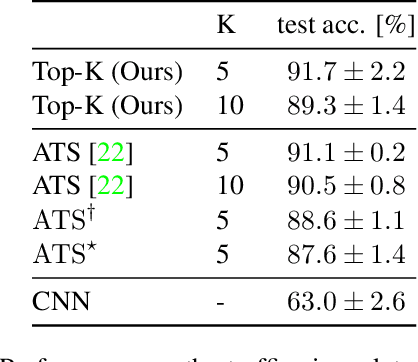
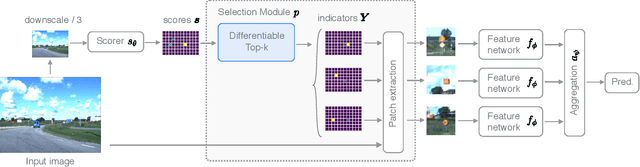
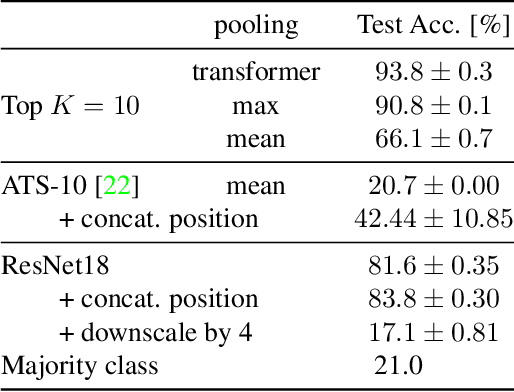
Abstract:Neural Networks require large amounts of memory and compute to process high resolution images, even when only a small part of the image is actually informative for the task at hand. We propose a method based on a differentiable Top-K operator to select the most relevant parts of the input to efficiently process high resolution images. Our method may be interfaced with any downstream neural network, is able to aggregate information from different patches in a flexible way, and allows the whole model to be trained end-to-end using backpropagation. We show results for traffic sign recognition, inter-patch relationship reasoning, and fine-grained recognition without using object/part bounding box annotations during training.
Understanding Robustness of Transformers for Image Classification
Mar 26, 2021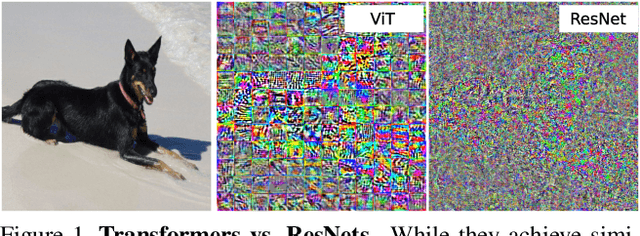
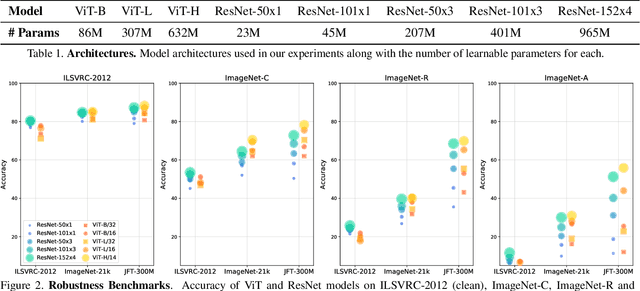
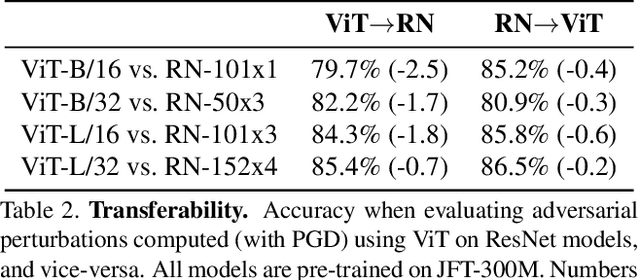

Abstract:Deep Convolutional Neural Networks (CNNs) have long been the architecture of choice for computer vision tasks. Recently, Transformer-based architectures like Vision Transformer (ViT) have matched or even surpassed ResNets for image classification. However, details of the Transformer architecture -- such as the use of non-overlapping patches -- lead one to wonder whether these networks are as robust. In this paper, we perform an extensive study of a variety of different measures of robustness of ViT models and compare the findings to ResNet baselines. We investigate robustness to input perturbations as well as robustness to model perturbations. We find that when pre-trained with a sufficient amount of data, ViT models are at least as robust as the ResNet counterparts on a broad range of perturbations. We also find that Transformers are robust to the removal of almost any single layer, and that while activations from later layers are highly correlated with each other, they nevertheless play an important role in classification.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge