Shan E Ahmed Raza
An Automated Pipeline for Tumour-Infiltrating Lymphocyte Scoring in Breast Cancer
Nov 21, 2023Abstract:Tumour-infiltrating lymphocytes (TILs) are considered as a valuable prognostic markers in both triple-negative and human epidermal growth factor receptor 2 (HER2) positive breast cancer. In this study, we introduce an innovative deep learning pipeline based on the Efficient-UNet architecture to predict the TILs score for breast cancer whole-slide images (WSIs). We first segment tumour and stromal regions in order to compute a tumour bulk mask. We then detect TILs within the tumour-associated stroma, generating a TILs score by closely mirroring the pathologist's workflow. Our method exhibits state-of-the-art performance in segmenting tumour/stroma areas and TILs detection, as demonstrated by internal cross-validation on the TiGER Challenge training dataset and evaluation on the final leaderboards. Additionally, our TILs score proves competitive in predicting survival outcomes within the same challenge, underscoring the clinical relevance and potential of our automated TILs scoring pipeline as a breast cancer prognostic tool.
Transformer-based Model for Oral Epithelial Dysplasia Segmentation
Nov 09, 2023



Abstract:Oral epithelial dysplasia (OED) is a premalignant histopathological diagnosis given to lesions of the oral cavity. OED grading is subject to large inter/intra-rater variability, resulting in the under/over-treatment of patients. We developed a new Transformer-based pipeline to improve detection and segmentation of OED in haematoxylin and eosin (H&E) stained whole slide images (WSIs). Our model was trained on OED cases (n = 260) and controls (n = 105) collected using three different scanners, and validated on test data from three external centres in the United Kingdom and Brazil (n = 78). Our internal experiments yield a mean F1-score of 0.81 for OED segmentation, which reduced slightly to 0.71 on external testing, showing good generalisability, and gaining state-of-the-art results. This is the first externally validated study to use Transformers for segmentation in precancerous histology images. Our publicly available model shows great promise to be the first step of a fully-integrated pipeline, allowing earlier and more efficient OED diagnosis, ultimately benefiting patient outcomes.
Domain Generalization in Computational Pathology: Survey and Guidelines
Oct 30, 2023



Abstract:Deep learning models have exhibited exceptional effectiveness in Computational Pathology (CPath) by tackling intricate tasks across an array of histology image analysis applications. Nevertheless, the presence of out-of-distribution data (stemming from a multitude of sources such as disparate imaging devices and diverse tissue preparation methods) can cause \emph{domain shift} (DS). DS decreases the generalization of trained models to unseen datasets with slightly different data distributions, prompting the need for innovative \emph{domain generalization} (DG) solutions. Recognizing the potential of DG methods to significantly influence diagnostic and prognostic models in cancer studies and clinical practice, we present this survey along with guidelines on achieving DG in CPath. We rigorously define various DS types, systematically review and categorize existing DG approaches and resources in CPath, and provide insights into their advantages, limitations, and applicability. We also conduct thorough benchmarking experiments with 28 cutting-edge DG algorithms to address a complex DG problem. Our findings suggest that careful experiment design and CPath-specific Stain Augmentation technique can be very effective. However, there is no one-size-fits-all solution for DG in CPath. Therefore, we establish clear guidelines for detecting and managing DS depending on different scenarios. While most of the concepts, guidelines, and recommendations are given for applications in CPath, we believe that they are applicable to most medical image analysis tasks as well.
A Fully Automated and Explainable Algorithm for the Prediction of Malignant Transformation in Oral Epithelial Dysplasia
Jul 06, 2023



Abstract:Oral epithelial dysplasia (OED) is a premalignant histopathological diagnosis given to lesions of the oral cavity. Its grading suffers from significant inter-/intra- observer variability, and does not reliably predict malignancy progression, potentially leading to suboptimal treatment decisions. To address this, we developed a novel artificial intelligence algorithm that can assign an Oral Malignant Transformation (OMT) risk score, based on histological patterns in the in Haematoxylin and Eosin stained whole slide images, to quantify the risk of OED progression. The algorithm is based on the detection and segmentation of nuclei within (and around) the epithelium using an in-house segmentation model. We then employed a shallow neural network fed with interpretable morphological/spatial features, emulating histological markers. We conducted internal cross-validation on our development cohort (Sheffield; n = 193 cases) followed by independent validation on two external cohorts (Birmingham and Belfast; n = 92 cases). The proposed OMTscore yields an AUROC = 0.74 in predicting whether an OED progresses to malignancy or not. Survival analyses showed the prognostic value of our OMTscore for predicting malignancy transformation, when compared to the manually-assigned WHO and binary grades. Analysis of the correctly predicted cases elucidated the presence of peri-epithelial and epithelium-infiltrating lymphocytes in the most predictive patches of cases that transformed (p < 0.0001). This is the first study to propose a completely automated algorithm for predicting OED transformation based on interpretable nuclear features, whilst being validated on external datasets. The algorithm shows better-than-human-level performance for prediction of OED malignant transformation and offers a promising solution to the challenges of grading OED in routine clinical practice.
CoNIC Challenge: Pushing the Frontiers of Nuclear Detection, Segmentation, Classification and Counting
Mar 14, 2023



Abstract:Nuclear detection, segmentation and morphometric profiling are essential in helping us further understand the relationship between histology and patient outcome. To drive innovation in this area, we setup a community-wide challenge using the largest available dataset of its kind to assess nuclear segmentation and cellular composition. Our challenge, named CoNIC, stimulated the development of reproducible algorithms for cellular recognition with real-time result inspection on public leaderboards. We conducted an extensive post-challenge analysis based on the top-performing models using 1,658 whole-slide images of colon tissue. With around 700 million detected nuclei per model, associated features were used for dysplasia grading and survival analysis, where we demonstrated that the challenge's improvement over the previous state-of-the-art led to significant boosts in downstream performance. Our findings also suggest that eosinophils and neutrophils play an important role in the tumour microevironment. We release challenge models and WSI-level results to foster the development of further methods for biomarker discovery.
Consistency Regularisation in Varying Contexts and Feature Perturbations for Semi-Supervised Semantic Segmentation of Histology Images
Feb 11, 2023



Abstract:Semantic segmentation of various tissue and nuclei types in histology images is fundamental to many downstream tasks in the area of computational pathology (CPath). In recent years, Deep Learning (DL) methods have been shown to perform well on segmentation tasks but DL methods generally require a large amount of pixel-wise annotated data. Pixel-wise annotation sometimes requires expert's knowledge and time which is laborious and costly to obtain. In this paper, we present a consistency based semi-supervised learning (SSL) approach that can help mitigate this challenge by exploiting a large amount of unlabelled data for model training thus alleviating the need for a large annotated dataset. However, SSL models might also be susceptible to changing context and features perturbations exhibiting poor generalisation due to the limited training data. We propose an SSL method that learns robust features from both labelled and unlabelled images by enforcing consistency against varying contexts and feature perturbations. The proposed method incorporates context-aware consistency by contrasting pairs of overlapping images in a pixel-wise manner from changing contexts resulting in robust and context invariant features. We show that cross-consistency training makes the encoder features invariant to different perturbations and improves the prediction confidence. Finally, entropy minimisation is employed to further boost the confidence of the final prediction maps from unlabelled data. We conduct an extensive set of experiments on two publicly available large datasets (BCSS and MoNuSeg) and show superior performance compared to the state-of-the-art methods.
LYSTO: The Lymphocyte Assessment Hackathon and Benchmark Dataset
Jan 16, 2023



Abstract:We introduce LYSTO, the Lymphocyte Assessment Hackathon, which was held in conjunction with the MICCAI 2019 Conference in Shenzen (China). The competition required participants to automatically assess the number of lymphocytes, in particular T-cells, in histopathological images of colon, breast, and prostate cancer stained with CD3 and CD8 immunohistochemistry. Differently from other challenges setup in medical image analysis, LYSTO participants were solely given a few hours to address this problem. In this paper, we describe the goal and the multi-phase organization of the hackathon; we describe the proposed methods and the on-site results. Additionally, we present post-competition results where we show how the presented methods perform on an independent set of lung cancer slides, which was not part of the initial competition, as well as a comparison on lymphocyte assessment between presented methods and a panel of pathologists. We show that some of the participants were capable to achieve pathologist-level performance at lymphocyte assessment. After the hackathon, LYSTO was left as a lightweight plug-and-play benchmark dataset on grand-challenge website, together with an automatic evaluation platform. LYSTO has supported a number of research in lymphocyte assessment in oncology. LYSTO will be a long-lasting educational challenge for deep learning and digital pathology, it is available at https://lysto.grand-challenge.org/.
Nuclear Segmentation and Classification: On Color & Compression Generalization
Jan 09, 2023Abstract:Since the introduction of digital and computational pathology as a field, one of the major problems in the clinical application of algorithms has been the struggle to generalize well to examples outside the distribution of the training data. Existing work to address this in both pathology and natural images has focused almost exclusively on classification tasks. We explore and evaluate the robustness of the 7 best performing nuclear segmentation and classification models from the largest computational pathology challenge for this problem to date, the CoNIC challenge. We demonstrate that existing state-of-the-art (SoTA) models are robust towards compression artifacts but suffer substantial performance reduction when subjected to shifts in the color domain. We find that using stain normalization to address the domain shift problem can be detrimental to the model performance. On the other hand, neural style transfer is more consistent in improving test performance when presented with large color variations in the wild.
Stain-Robust Mitotic Figure Detection for MIDOG 2022 Challenge
Aug 26, 2022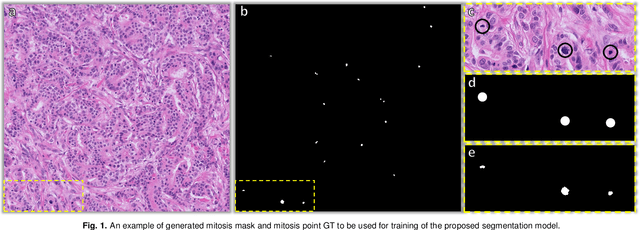
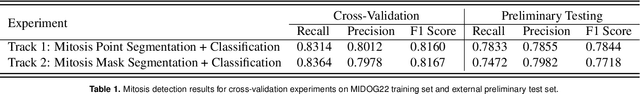
Abstract:The detection of mitotic figures from different scanners/sites remains an important topic of research, owing to its potential in assisting clinicians with tumour grading. The MItosis DOmain Generalization (MIDOG) 2022 challenge aims to test the robustness of detection models on unseen data from multiple scanners and tissue types for this task. We present a short summary of the approach employed by the TIA Centre team to address this challenge. Our approach is based on a hybrid detection model, where mitotic candidates are segmented, before being refined by a deep learning classifier. Cross-validation on the training images achieved the F1-score of 0.816 and 0.784 on the preliminary test set, demonstrating the generalizability of our model to unseen data from new scanners.
TIAger: Tumor-Infiltrating Lymphocyte Scoring in Breast Cancer for the TiGER Challenge
Jun 23, 2022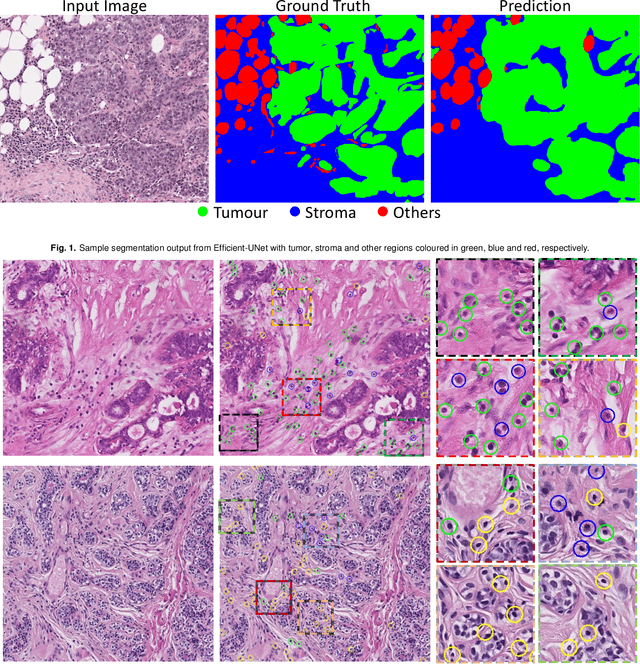
Abstract:The quantification of tumor-infiltrating lymphocytes (TILs) has been shown to be an independent predictor for prognosis of breast cancer patients. Typically, pathologists give an estimate of the proportion of the stromal region that contains TILs to obtain a TILs score. The Tumor InfiltratinG lymphocytes in breast cancER (TiGER) challenge, aims to assess the prognostic significance of computer-generated TILs scores for predicting survival as part of a Cox proportional hazards model. For this challenge, as the TIAger team, we have developed an algorithm to first segment tumor vs. stroma, before localising the tumor bulk region for TILs detection. Finally, we use these outputs to generate a TILs score for each case. On preliminary testing, our approach achieved a tumor-stroma weighted Dice score of 0.791 and a FROC score of 0.572 for lymphocytic detection. For predicting survival, our model achieved a C-index of 0.719. These results achieved first place across the preliminary testing leaderboards of the TiGER challenge.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge