Quanzheng Li
CAMCA, Massachusetts General Hospital and Harvard Medical School
MAST-Pro: Dynamic Mixture-of-Experts for Adaptive Segmentation of Pan-Tumors with Knowledge-Driven Prompts
Mar 18, 2025Abstract:Accurate tumor segmentation is crucial for cancer diagnosis and treatment. While foundation models have advanced general-purpose segmentation, existing methods still struggle with: (1) limited incorporation of medical priors, (2) imbalance between generic and tumor-specific features, and (3) high computational costs for clinical adaptation. To address these challenges, we propose MAST-Pro (Mixture-of-experts for Adaptive Segmentation of pan-Tumors with knowledge-driven Prompts), a novel framework that integrates dynamic Mixture-of-Experts (D-MoE) and knowledge-driven prompts for pan-tumor segmentation. Specifically, text and anatomical prompts provide domain-specific priors, guiding tumor representation learning, while D-MoE dynamically selects experts to balance generic and tumor-specific feature learning, improving segmentation accuracy across diverse tumor types. To enhance efficiency, we employ Parameter-Efficient Fine-Tuning (PEFT), optimizing MAST-Pro with significantly reduced computational overhead. Experiments on multi-anatomical tumor datasets demonstrate that MAST-Pro outperforms state-of-the-art approaches, achieving up to a 5.20% improvement in average DSC while reducing trainable parameters by 91.04%, without compromising accuracy.
Anatomically and Metabolically Informed Diffusion for Unified Denoising and Segmentation in Low-Count PET Imaging
Mar 17, 2025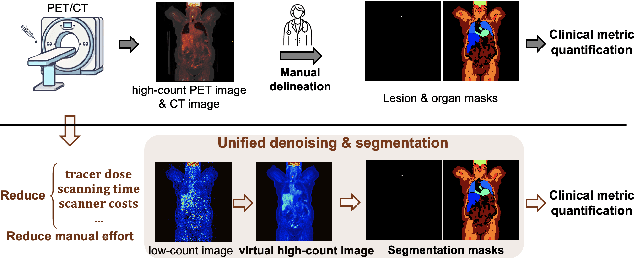
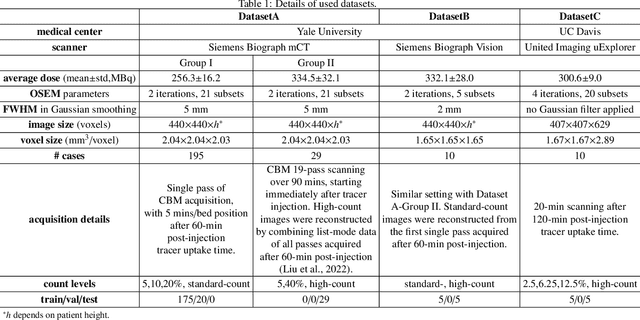
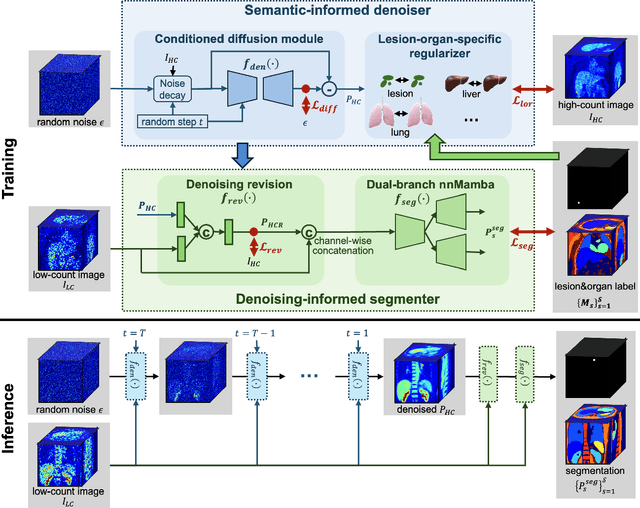
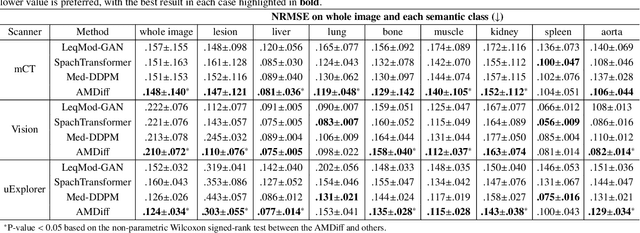
Abstract:Positron emission tomography (PET) image denoising, along with lesion and organ segmentation, are critical steps in PET-aided diagnosis. However, existing methods typically treat these tasks independently, overlooking inherent synergies between them as correlated steps in the analysis pipeline. In this work, we present the anatomically and metabolically informed diffusion (AMDiff) model, a unified framework for denoising and lesion/organ segmentation in low-count PET imaging. By integrating multi-task functionality and exploiting the mutual benefits of these tasks, AMDiff enables direct quantification of clinical metrics, such as total lesion glycolysis (TLG), from low-count inputs. The AMDiff model incorporates a semantic-informed denoiser based on diffusion strategy and a denoising-informed segmenter utilizing nnMamba architecture. The segmenter constrains denoised outputs via a lesion-organ-specific regularizer, while the denoiser enhances the segmenter by providing enriched image information through a denoising revision module. These components are connected via a warming-up mechanism to optimize multitask interactions. Experiments on multi-vendor, multi-center, and multi-noise-level datasets demonstrate the superior performance of AMDiff. For test cases below 20% of the clinical count levels from participating sites, AMDiff achieves TLG quantification biases of -26.98%, outperforming its ablated versions which yield biases of -35.85% (without the lesion-organ-specific regularizer) and -40.79% (without the denoising revision module).
Prediction of Frozen Region Growth in Kidney Cryoablation Intervention Using a 3D Flow-Matching Model
Mar 06, 2025Abstract:This study presents a 3D flow-matching model designed to predict the progression of the frozen region (iceball) during kidney cryoablation. Precise intraoperative guidance is critical in cryoablation to ensure complete tumor eradication while preserving adjacent healthy tissue. However, conventional methods, typically based on physics driven or diffusion based simulations, are computationally demanding and often struggle to represent complex anatomical structures accurately. To address these limitations, our approach leverages intraoperative CT imaging to inform the model. The proposed 3D flow matching model is trained to learn a continuous deformation field that maps early-stage CT scans to future predictions. This transformation not only estimates the volumetric expansion of the iceball but also generates corresponding segmentation masks, effectively capturing spatial and morphological changes over time. Quantitative analysis highlights the model robustness, demonstrating strong agreement between predictions and ground-truth segmentations. The model achieves an Intersection over Union (IoU) score of 0.61 and a Dice coefficient of 0.75. By integrating real time CT imaging with advanced deep learning techniques, this approach has the potential to enhance intraoperative guidance in kidney cryoablation, improving procedural outcomes and advancing the field of minimally invasive surgery.
Developing a PET/CT Foundation Model for Cross-Modal Anatomical and Functional Imaging
Mar 04, 2025



Abstract:In oncology, Positron Emission Tomography-Computed Tomography (PET/CT) is widely used in cancer diagnosis, staging, and treatment monitoring, as it combines anatomical details from CT with functional metabolic activity and molecular marker expression information from PET. However, existing artificial intelligence-driven PET/CT analyses rely predominantly on task-specific models trained from scratch or on limited datasets, limiting their generalizability and robustness. To address this, we propose a foundation model approach specifically designed for multimodal PET/CT imaging. We introduce the Cross-Fraternal Twin Masked Autoencoder (FratMAE), a novel framework that effectively integrates whole-body anatomical and functional or molecular information. FratMAE employs separate Vision Transformer (ViT) encoders for PET and CT scans, along with cross-attention decoders that enable synergistic interactions between modalities during masked autoencoder training. Additionally, it incorporates textual metadata to enhance PET representation learning. By pre-training on PET/CT datasets, FratMAE captures intricate cross-modal relationships and global uptake patterns, achieving superior performance on downstream tasks and demonstrating its potential as a generalizable foundation model.
Pseudoinverse Diffusion Models for Generative CT Image Reconstruction from Low Dose Data
Feb 20, 2025



Abstract:Score-based diffusion models have significantly advanced generative deep learning for image processing. Measurement conditioned models have also been applied to inverse problems such as CT reconstruction. However, the conventional approach, culminating in white noise, often requires a high number of reverse process update steps and score function evaluations. To address this limitation, we propose an alternative forward process in score-based diffusion models that aligns with the noise characteristics of low-dose CT reconstructions, rather than converging to white noise. This method significantly reduces the number of required score function evaluations, enhancing efficiency and maintaining familiar noise textures for radiologists, Our approach not only accelerates the generative process but also retains CT noise correlations, a key aspect often criticized by clinicians for deep learning reconstructions. In this work, we rigorously define a matrix-controlled stochastic process for this purpose and validate it through computational experiments. Using a dataset from The Cancer Genome Atlas Liver Hepatocellular Carcinoma (TCGA-LIHC), we simulate low-dose CT measurements and train our model, comparing it with a baseline scalar diffusion process and conditional diffusion model. Our results demonstrate the superiority of our pseudoinverse diffusion model in terms of efficiency and the ability to produce high-quality reconstructions that are familiar in texture to medical professionals in a low number of score function evaluations. This advancement paves the way for more efficient and clinically practical diffusion models in medical imaging, particularly beneficial in scenarios demanding rapid reconstructions or lower radiation exposure.
Enhancing Cognition and Explainability of Multimodal Foundation Models with Self-Synthesized Data
Feb 19, 2025Abstract:Large multimodal models (LMMs) have shown impressive capabilities in a wide range of visual tasks. However, they often struggle with fine-grained visual reasoning, failing to identify domain-specific objectives and provide justifiable explanations for their predictions. To address this, we propose a novel visual rejection sampling framework to improve the cognition and explainability of LMMs using self-synthesized data. Specifically, visual fine-tuning requires images, queries, and target answers. Our approach begins by synthesizing interpretable answers that include human-verifiable visual features. These features are based on expert-defined concepts, carefully selected based on their alignment with the image content. After each round of fine-tuning, we apply a reward model-free filtering mechanism to select the highest-quality interpretable answers for the next round of tuning. This iterative process of data synthesis and fine-tuning progressively improves the model's ability to generate accurate and reasonable explanations. Experimental results demonstrate the effectiveness of our method in improving both the accuracy and explainability of specialized visual classification tasks.
SearchRAG: Can Search Engines Be Helpful for LLM-based Medical Question Answering?
Feb 18, 2025Abstract:Large Language Models (LLMs) have shown remarkable capabilities in general domains but often struggle with tasks requiring specialized knowledge. Conventional Retrieval-Augmented Generation (RAG) techniques typically retrieve external information from static knowledge bases, which can be outdated or incomplete, missing fine-grained clinical details essential for accurate medical question answering. In this work, we propose SearchRAG, a novel framework that overcomes these limitations by leveraging real-time search engines. Our method employs synthetic query generation to convert complex medical questions into search-engine-friendly queries and utilizes uncertainty-based knowledge selection to filter and incorporate the most relevant and informative medical knowledge into the LLM's input. Experimental results demonstrate that our method significantly improves response accuracy in medical question answering tasks, particularly for complex questions requiring detailed and up-to-date knowledge.
Fine-Tuning Open-Source Large Language Models to Improve Their Performance on Radiation Oncology Tasks: A Feasibility Study to Investigate Their Potential Clinical Applications in Radiation Oncology
Jan 28, 2025



Abstract:Background: The radiation oncology clinical practice involves many steps relying on the dynamic interplay of abundant text data. Large language models have displayed remarkable capabilities in processing complex text information. But their direct applications in specific fields like radiation oncology remain underexplored. Purpose: This study aims to investigate whether fine-tuning LLMs with domain knowledge can improve the performance on Task (1) treatment regimen generation, Task (2) treatment modality selection (photon, proton, electron, or brachytherapy), and Task (3) ICD-10 code prediction in radiation oncology. Methods: Data for 15,724 patient cases were extracted. Cases where patients had a single diagnostic record, and a clearly identifiable primary treatment plan were selected for preprocessing and manual annotation to have 7,903 cases of the patient diagnosis, treatment plan, treatment modality, and ICD-10 code. Each case was used to construct a pair consisting of patient diagnostics details and an answer (treatment regimen, treatment modality, or ICD-10 code respectively) for the supervised fine-tuning of these three tasks. Open source LLaMA2-7B and Mistral-7B models were utilized for the fine-tuning with the Low-Rank Approximations method. Accuracy and ROUGE-1 score were reported for the fine-tuned models and original models. Clinical evaluation was performed on Task (1) by radiation oncologists, while precision, recall, and F-1 score were evaluated for Task (2) and (3). One-sided Wilcoxon signed-rank tests were used to statistically analyze the results. Results: Fine-tuned LLMs outperformed original LLMs across all tasks with p-value <= 0.001. Clinical evaluation demonstrated that over 60% of the fine-tuned LLMs-generated treatment regimens were clinically acceptable. Precision, recall, and F1-score showed improved performance of fine-tuned LLMs.
Large Language Models for Bioinformatics
Jan 10, 2025
Abstract:With the rapid advancements in large language model (LLM) technology and the emergence of bioinformatics-specific language models (BioLMs), there is a growing need for a comprehensive analysis of the current landscape, computational characteristics, and diverse applications. This survey aims to address this need by providing a thorough review of BioLMs, focusing on their evolution, classification, and distinguishing features, alongside a detailed examination of training methodologies, datasets, and evaluation frameworks. We explore the wide-ranging applications of BioLMs in critical areas such as disease diagnosis, drug discovery, and vaccine development, highlighting their impact and transformative potential in bioinformatics. We identify key challenges and limitations inherent in BioLMs, including data privacy and security concerns, interpretability issues, biases in training data and model outputs, and domain adaptation complexities. Finally, we highlight emerging trends and future directions, offering valuable insights to guide researchers and clinicians toward advancing BioLMs for increasingly sophisticated biological and clinical applications.
End-to-End Deep Learning for Interior Tomography with Low-Dose X-ray CT
Jan 09, 2025Abstract:Objective: There exist several X-ray computed tomography (CT) scanning strategies to reduce a radiation dose, such as (1) sparse-view CT, (2) low-dose CT, and (3) region-of-interest (ROI) CT (called interior tomography). To further reduce the dose, the sparse-view and/or low-dose CT settings can be applied together with interior tomography. Interior tomography has various advantages in terms of reducing the number of detectors and decreasing the X-ray radiation dose. However, a large patient or small field-of-view (FOV) detector can cause truncated projections, and then the reconstructed images suffer from severe cupping artifacts. In addition, although the low-dose CT can reduce the radiation exposure dose, analytic reconstruction algorithms produce image noise. Recently, many researchers have utilized image-domain deep learning (DL) approaches to remove each artifact and demonstrated impressive performances, and the theory of deep convolutional framelets supports the reason for the performance improvement. Approach: In this paper, we found that the image-domain convolutional neural network (CNN) is difficult to solve coupled artifacts, based on deep convolutional framelets. Significance: To address the coupled problem, we decouple it into two sub-problems: (i) image domain noise reduction inside truncated projection to solve low-dose CT problem and (ii) extrapolation of projection outside truncated projection to solve the ROI CT problem. The decoupled sub-problems are solved directly with a novel proposed end-to-end learning using dual-domain CNNs. Main results: We demonstrate that the proposed method outperforms the conventional image-domain deep learning methods, and a projection-domain CNN shows better performance than the image-domain CNNs which are commonly used by many researchers.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge