Nicholas Monath
Improving Dual-Encoder Training through Dynamic Indexes for Negative Mining
Mar 27, 2023



Abstract:Dual encoder models are ubiquitous in modern classification and retrieval. Crucial for training such dual encoders is an accurate estimation of gradients from the partition function of the softmax over the large output space; this requires finding negative targets that contribute most significantly ("hard negatives"). Since dual encoder model parameters change during training, the use of traditional static nearest neighbor indexes can be sub-optimal. These static indexes (1) periodically require expensive re-building of the index, which in turn requires (2) expensive re-encoding of all targets using updated model parameters. This paper addresses both of these challenges. First, we introduce an algorithm that uses a tree structure to approximate the softmax with provable bounds and that dynamically maintains the tree. Second, we approximate the effect of a gradient update on target encodings with an efficient Nystrom low-rank approximation. In our empirical study on datasets with over twenty million targets, our approach cuts error by half in relation to oracle brute-force negative mining. Furthermore, our method surpasses prior state-of-the-art while using 150x less accelerator memory.
Autoregressive Structured Prediction with Language Models
Nov 17, 2022



Abstract:Recent years have seen a paradigm shift in NLP towards using pretrained language models ({PLM}) for a wide range of tasks. However, there are many difficult design decisions to represent structures (e.g. tagged text, coreference chains) in a way such that they can be captured by PLMs. Prior work on structured prediction with PLMs typically flattens the structured output into a sequence, which limits the quality of structural information being learned and leads to inferior performance compared to classic discriminative models. In this work, we describe an approach to model structures as sequences of actions in an autoregressive manner with PLMs, allowing in-structure dependencies to be learned without any loss. Our approach achieves the new state-of-the-art on all the structured prediction tasks we looked at, namely, named entity recognition, end-to-end relation extraction, and coreference resolution.
Efficient Nearest Neighbor Search for Cross-Encoder Models using Matrix Factorization
Oct 23, 2022Abstract:Efficient k-nearest neighbor search is a fundamental task, foundational for many problems in NLP. When the similarity is measured by dot-product between dual-encoder vectors or $\ell_2$-distance, there already exist many scalable and efficient search methods. But not so when similarity is measured by more accurate and expensive black-box neural similarity models, such as cross-encoders, which jointly encode the query and candidate neighbor. The cross-encoders' high computational cost typically limits their use to reranking candidates retrieved by a cheaper model, such as dual encoder or TF-IDF. However, the accuracy of such a two-stage approach is upper-bounded by the recall of the initial candidate set, and potentially requires additional training to align the auxiliary retrieval model with the cross-encoder model. In this paper, we present an approach that avoids the use of a dual-encoder for retrieval, relying solely on the cross-encoder. Retrieval is made efficient with CUR decomposition, a matrix decomposition approach that approximates all pairwise cross-encoder distances from a small subset of rows and columns of the distance matrix. Indexing items using our approach is computationally cheaper than training an auxiliary dual-encoder model through distillation. Empirically, for k > 10, our approach provides test-time recall-vs-computational cost trade-offs superior to the current widely-used methods that re-rank items retrieved using a dual-encoder or TF-IDF.
Longtonotes: OntoNotes with Longer Coreference Chains
Oct 07, 2022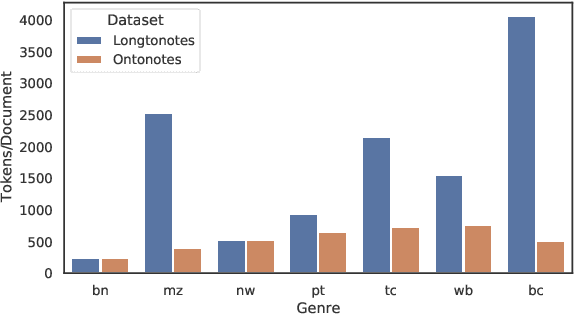
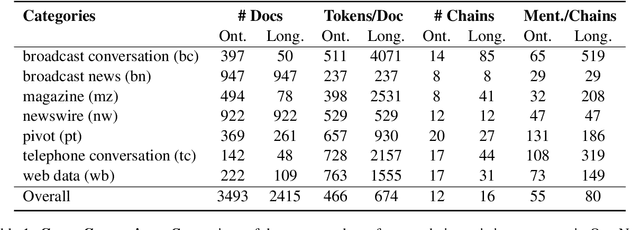
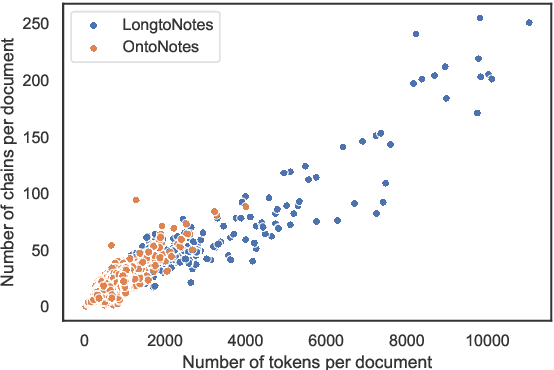
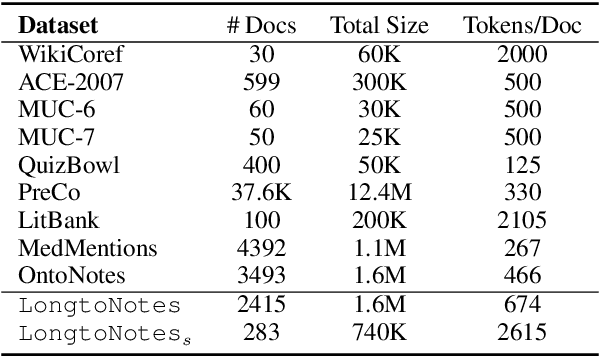
Abstract:Ontonotes has served as the most important benchmark for coreference resolution. However, for ease of annotation, several long documents in Ontonotes were split into smaller parts. In this work, we build a corpus of coreference-annotated documents of significantly longer length than what is currently available. We do so by providing an accurate, manually-curated, merging of annotations from documents that were split into multiple parts in the original Ontonotes annotation process. The resulting corpus, which we call LongtoNotes contains documents in multiple genres of the English language with varying lengths, the longest of which are up to 8x the length of documents in Ontonotes, and 2x those in Litbank. We evaluate state-of-the-art neural coreference systems on this new corpus, analyze the relationships between model architectures/hyperparameters and document length on performance and efficiency of the models, and demonstrate areas of improvement in long-document coreference modeling revealed by our new corpus. Our data and code is available at: https://github.com/kumar-shridhar/LongtoNotes.
Unsupervised Opinion Summarization Using Approximate Geodesics
Sep 15, 2022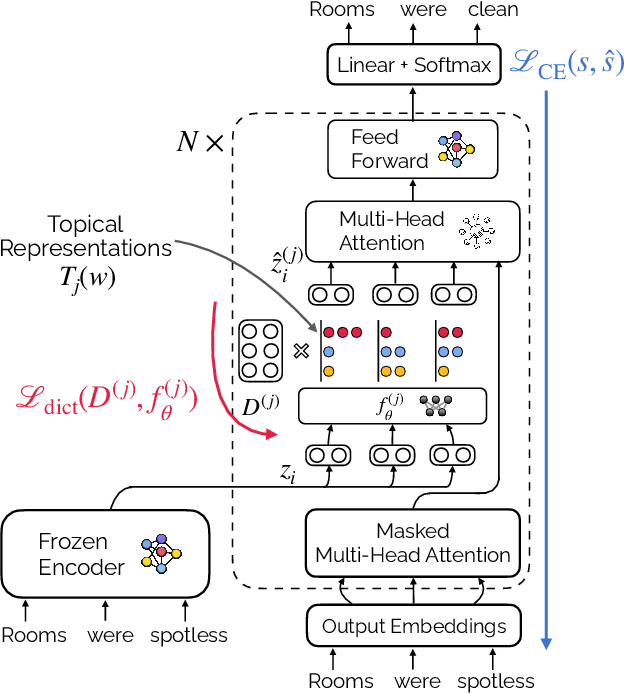
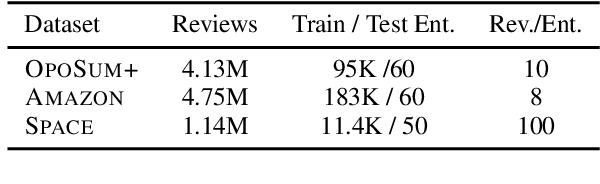
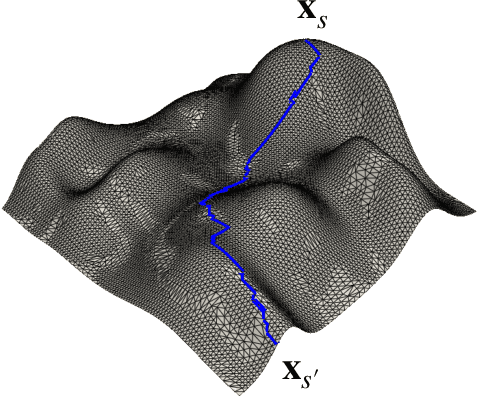
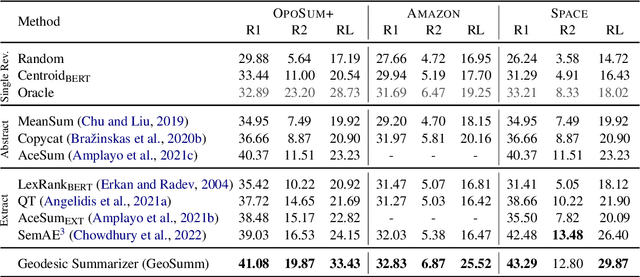
Abstract:Opinion summarization is the task of creating summaries capturing popular opinions from user reviews. In this paper, we introduce Geodesic Summarizer (GeoSumm), a novel system to perform unsupervised extractive opinion summarization. GeoSumm involves an encoder-decoder based representation learning model, that generates representations of text as a distribution over latent semantic units. GeoSumm generates these representations by performing dictionary learning over pre-trained text representations at multiple decoder layers. We then use these representations to quantify the relevance of review sentences using a novel approximate geodesic distance based scoring mechanism. We use the relevance scores to identify popular opinions in order to compose general and aspect-specific summaries. Our proposed model, GeoSumm, achieves state-of-the-art performance on three opinion summarization datasets. We perform additional experiments to analyze the functioning of our model and showcase the generalization ability of {\X} across different domains.
Sublinear Time Approximation of Text Similarity Matrices
Dec 17, 2021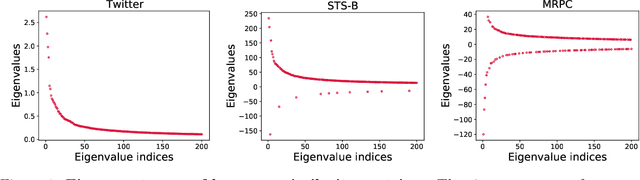
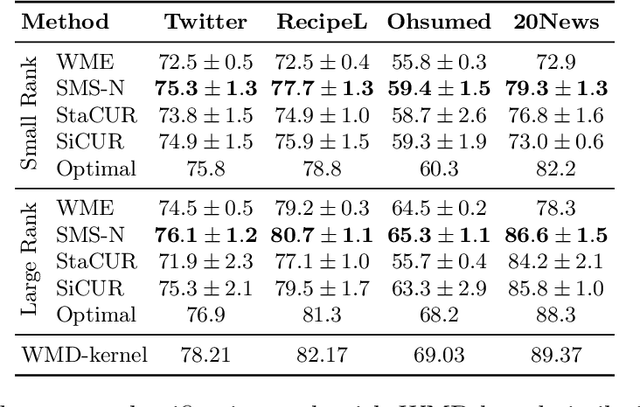
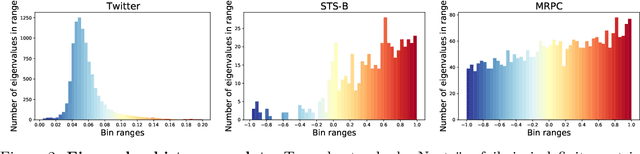
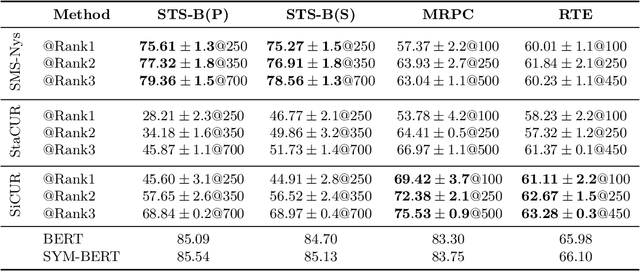
Abstract:We study algorithms for approximating pairwise similarity matrices that arise in natural language processing. Generally, computing a similarity matrix for $n$ data points requires $\Omega(n^2)$ similarity computations. This quadratic scaling is a significant bottleneck, especially when similarities are computed via expensive functions, e.g., via transformer models. Approximation methods reduce this quadratic complexity, often by using a small subset of exactly computed similarities to approximate the remainder of the complete pairwise similarity matrix. Significant work focuses on the efficient approximation of positive semidefinite (PSD) similarity matrices, which arise e.g., in kernel methods. However, much less is understood about indefinite (non-PSD) similarity matrices, which often arise in NLP. Motivated by the observation that many of these matrices are still somewhat close to PSD, we introduce a generalization of the popular Nystr\"{o}m method to the indefinite setting. Our algorithm can be applied to any similarity matrix and runs in sublinear time in the size of the matrix, producing a rank-$s$ approximation with just $O(ns)$ similarity computations. We show that our method, along with a simple variant of CUR decomposition, performs very well in approximating a variety of similarity matrices arising in NLP tasks. We demonstrate high accuracy of the approximated similarity matrices in the downstream tasks of document classification, sentence similarity, and cross-document coreference.
Entity Linking and Discovery via Arborescence-based Supervised Clustering
Sep 02, 2021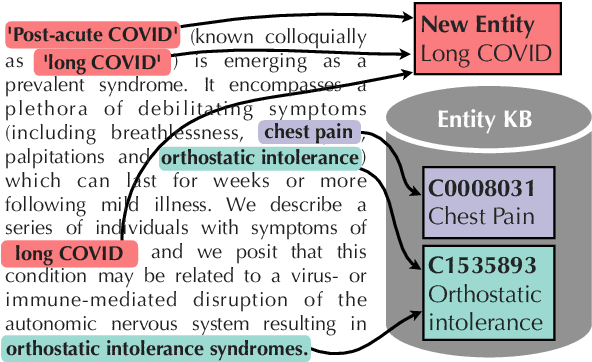
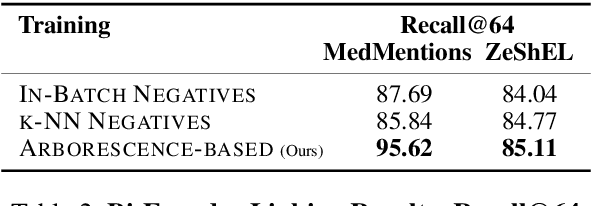
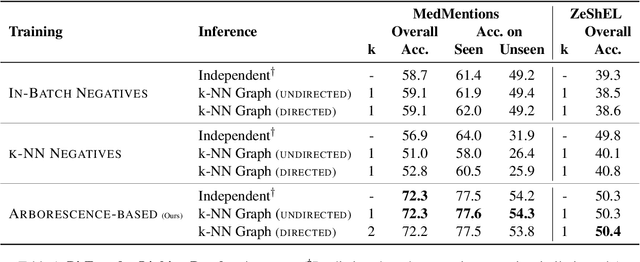
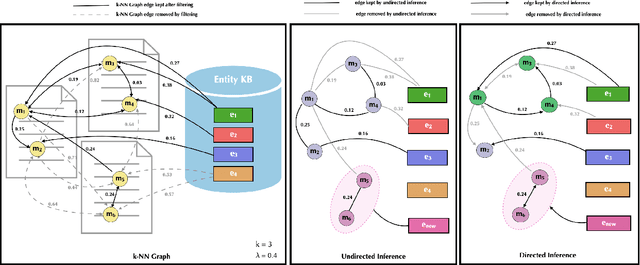

Abstract:Previous work has shown promising results in performing entity linking by measuring not only the affinities between mentions and entities but also those amongst mentions. In this paper, we present novel training and inference procedures that fully utilize mention-to-mention affinities by building minimum arborescences (i.e., directed spanning trees) over mentions and entities across documents in order to make linking decisions. We also show that this method gracefully extends to entity discovery, enabling the clustering of mentions that do not have an associated entity in the knowledge base. We evaluate our approach on the Zero-Shot Entity Linking dataset and MedMentions, the largest publicly available biomedical dataset, and show significant improvements in performance for both entity linking and discovery compared to identically parameterized models. We further show significant efficiency improvements with only a small loss in accuracy over previous work, which use more computationally expensive models.
Exact and Approximate Hierarchical Clustering Using A*
Apr 14, 2021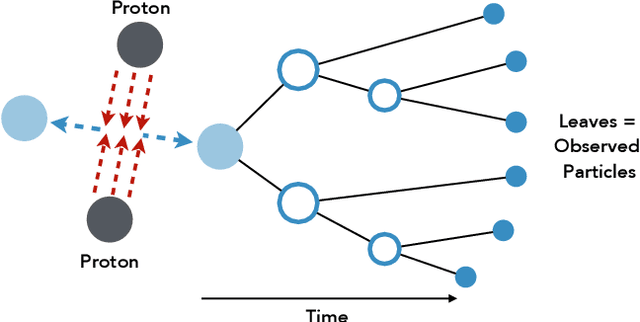
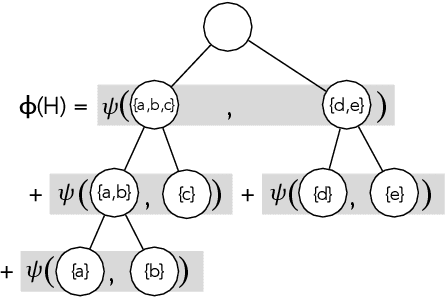
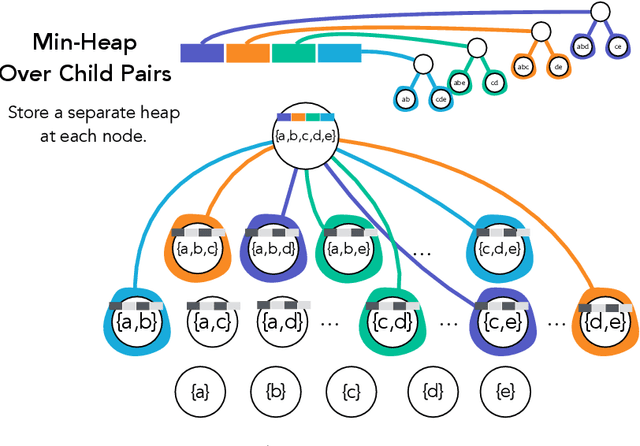
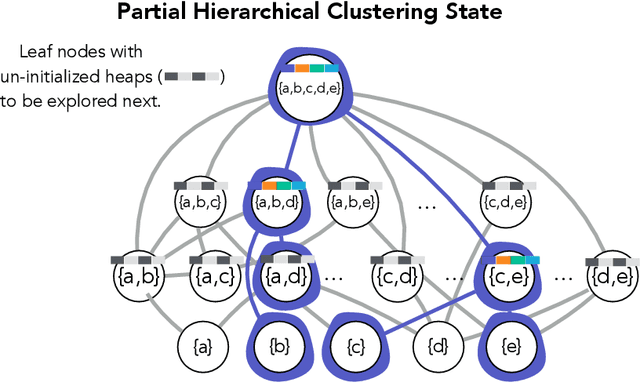
Abstract:Hierarchical clustering is a critical task in numerous domains. Many approaches are based on heuristics and the properties of the resulting clusterings are studied post hoc. However, in several applications, there is a natural cost function that can be used to characterize the quality of the clustering. In those cases, hierarchical clustering can be seen as a combinatorial optimization problem. To that end, we introduce a new approach based on A* search. We overcome the prohibitively large search space by combining A* with a novel \emph{trellis} data structure. This combination results in an exact algorithm that scales beyond previous state of the art, from a search space with $10^{12}$ trees to $10^{15}$ trees, and an approximate algorithm that improves over baselines, even in enormous search spaces that contain more than $10^{1000}$ trees. We empirically demonstrate that our method achieves substantially higher quality results than baselines for a particle physics use case and other clustering benchmarks. We describe how our method provides significantly improved theoretical bounds on the time and space complexity of A* for clustering.
Model-Agnostic Graph Regularization for Few-Shot Learning
Feb 14, 2021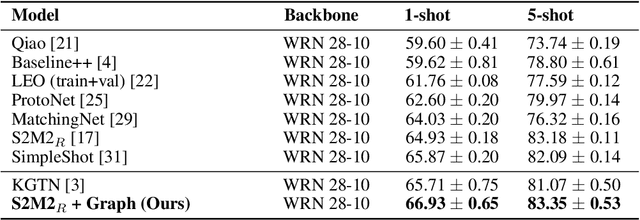
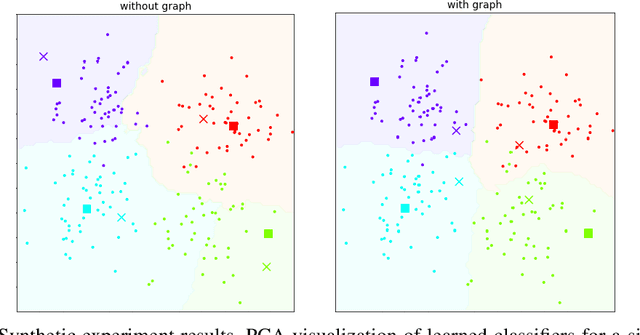
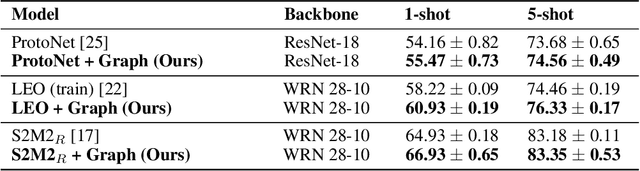
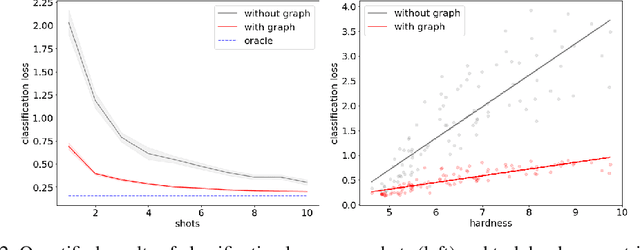
Abstract:In many domains, relationships between categories are encoded in the knowledge graph. Recently, promising results have been achieved by incorporating knowledge graph as side information in hard classification tasks with severely limited data. However, prior models consist of highly complex architectures with many sub-components that all seem to impact performance. In this paper, we present a comprehensive empirical study on graph embedded few-shot learning. We introduce a graph regularization approach that allows a deeper understanding of the impact of incorporating graph information between labels. Our proposed regularization is widely applicable and model-agnostic, and boosts the performance of any few-shot learning model, including fine-tuning, metric-based, and optimization-based meta-learning. Our approach improves the performance of strong base learners by up to 2% on Mini-ImageNet and 6.7% on ImageNet-FS, outperforming state-of-the-art graph embedded methods. Additional analyses reveal that graph regularizing models result in a lower loss for more difficult tasks, such as those with fewer shots and less informative support examples.
Scalable Bottom-Up Hierarchical Clustering
Nov 04, 2020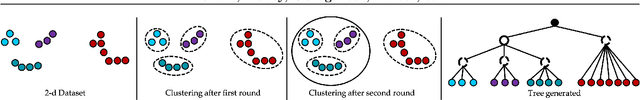
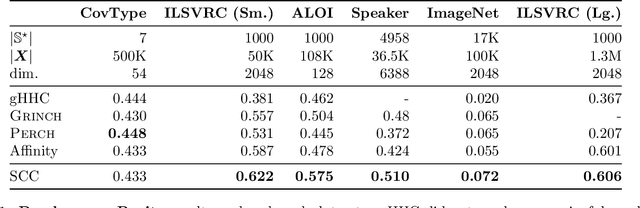
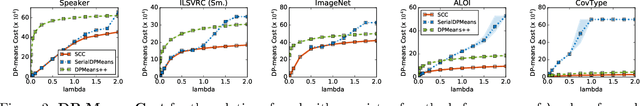
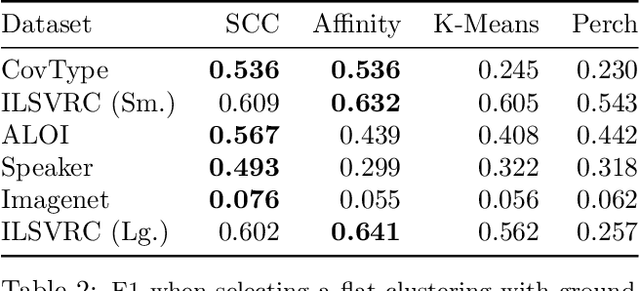
Abstract:Bottom-up algorithms such as the classic hierarchical agglomerative clustering, are highly effective for hierarchical as well as flat clustering. However, the large number of rounds and their sequential nature limit the scalability of agglomerative clustering. In this paper, we present an alternative round-based bottom-up hierarchical clustering, the Sub-Cluster Component Algorithm (SCC), that scales gracefully to massive datasets. Our method builds many sub-clusters in parallel in a given round and requires many fewer rounds -- usually an order of magnitude smaller than classic agglomerative clustering. Our theoretical analysis shows that, under a modest separability assumption, SCC will contain the optimal flat clustering. SCC also provides a 2-approx solution to the DP-means objective, thereby introducing a novel application of hierarchical clustering methods. Empirically, SCC finds better hierarchies and flat clusterings even when the data does not satisfy the separability assumption. We demonstrate the scalability of our method by applying it to a dataset of 30 billion points and showing that SCC produces higher quality clusterings than the state-of-the-art.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge