Sonification as a Reliable Alternative to Conventional Visual Surgical Navigation
Jun 30, 2022Sasan Matinfar, Mehrdad Salehi, Daniel Suter, Matthias Seibold, Navid Navab, Shervin Dehghani, Florian Wanivenhaus, Philipp Fürnstahl, Mazda Farshad, Nassir Navab
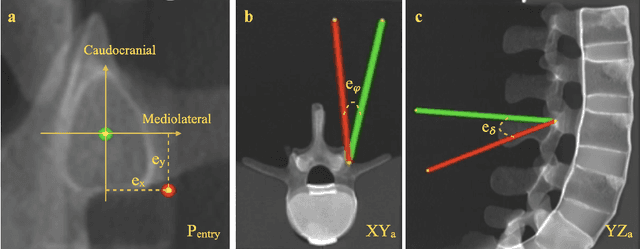
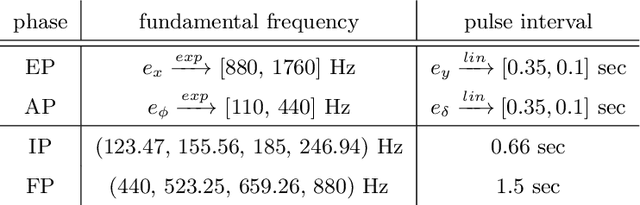
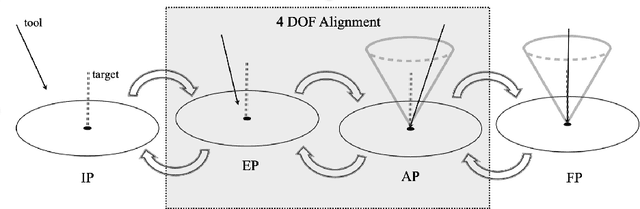
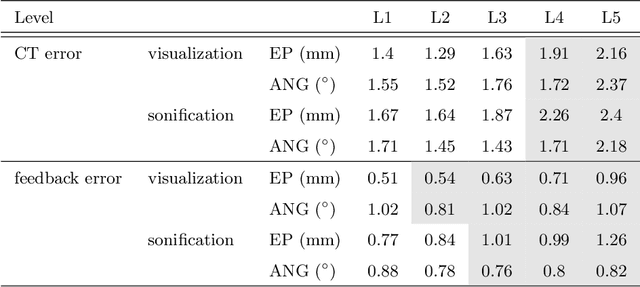
Despite the undeniable advantages of image-guided surgical assistance systems in terms of accuracy, such systems have not yet fully met surgeons' needs or expectations regarding usability, time efficiency, and their integration into the surgical workflow. On the other hand, perceptual studies have shown that presenting independent but causally correlated information via multimodal feedback involving different sensory modalities can improve task performance. This article investigates an alternative method for computer-assisted surgical navigation, introduces a novel sonification methodology for navigated pedicle screw placement, and discusses advanced solutions based on multisensory feedback. The proposed method comprises a novel sonification solution for alignment tasks in four degrees of freedom based on frequency modulation (FM) synthesis. We compared the resulting accuracy and execution time of the proposed sonification method with visual navigation, which is currently considered the state of the art. We conducted a phantom study in which 17 surgeons executed the pedicle screw placement task in the lumbar spine, guided by either the proposed sonification-based or the traditional visual navigation method. The results demonstrated that the proposed method is as accurate as the state of the art while decreasing the surgeon's need to focus on visual navigation displays instead of the natural focus on surgical tools and targeted anatomy during task execution.
DeStripe: A Self2Self Spatio-Spectral Graph Neural Network with Unfolded Hessian for Stripe Artifact Removal in Light-sheet Microscopy
Jun 27, 2022Yu Liu, Kurt Weiss, Nassir Navab, Carsten Marr, Jan Huisken, Tingying Peng
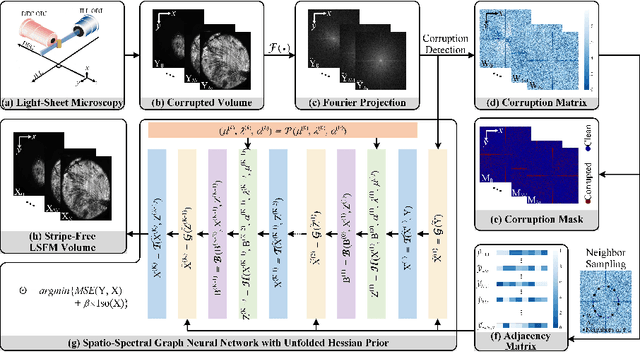
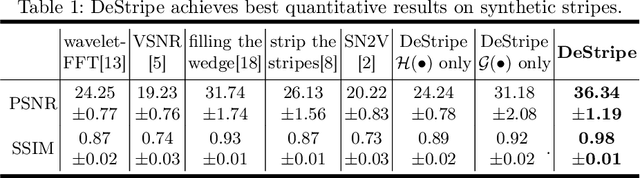

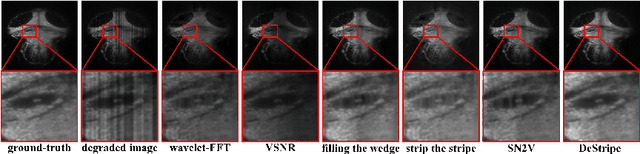
Light-sheet fluorescence microscopy (LSFM) is a cutting-edge volumetric imaging technique that allows for three-dimensional imaging of mesoscopic samples with decoupled illumination and detection paths. Although the selective excitation scheme of such a microscope provides intrinsic optical sectioning that minimizes out-of-focus fluorescence background and sample photodamage, it is prone to light absorption and scattering effects, which results in uneven illumination and striping artifacts in the images adversely. To tackle this issue, in this paper, we propose a blind stripe artifact removal algorithm in LSFM, called DeStripe, which combines a self-supervised spatio-spectral graph neural network with unfolded Hessian prior. Specifically, inspired by the desirable properties of Fourier transform in condensing striping information into isolated values in the frequency domain, DeStripe firstly localizes the potentially corrupted Fourier coefficients by exploiting the structural difference between unidirectional stripe artifacts and more isotropic foreground images. Affected Fourier coefficients can then be fed into a graph neural network for recovery, with a Hessian regularization unrolled to further ensure structures in the standard image space are well preserved. Since in realistic, stripe-free LSFM barely exists with a standard image acquisition protocol, DeStripe is equipped with a Self2Self denoising loss term, enabling artifact elimination without access to stripe-free ground truth images. Competitive experimental results demonstrate the efficacy of DeStripe in recovering corrupted biomarkers in LSFM with both synthetic and real stripe artifacts.
U-PET: MRI-based Dementia Detection with Joint Generation of Synthetic FDG-PET Images
Jun 16, 2022Marcel Kollovieh, Matthias Keicher, Stephan Wunderlich, Hendrik Burwinkel, Thomas Wendler, Nassir Navab
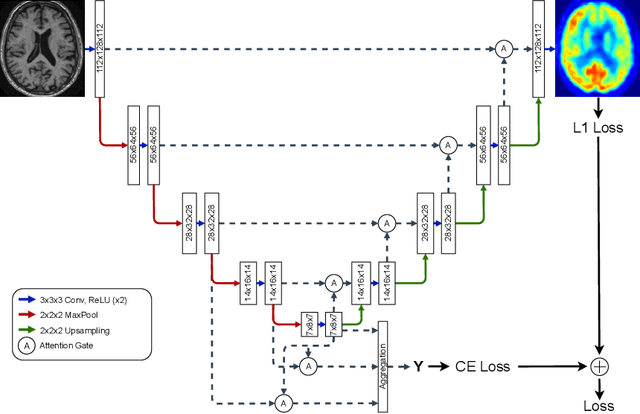
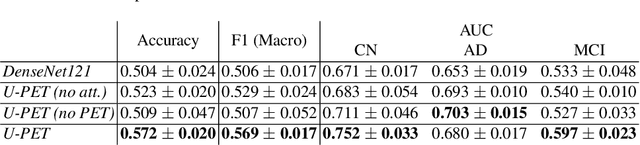
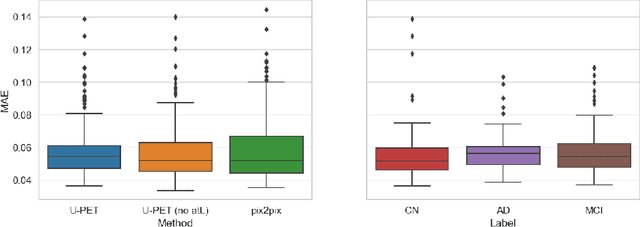
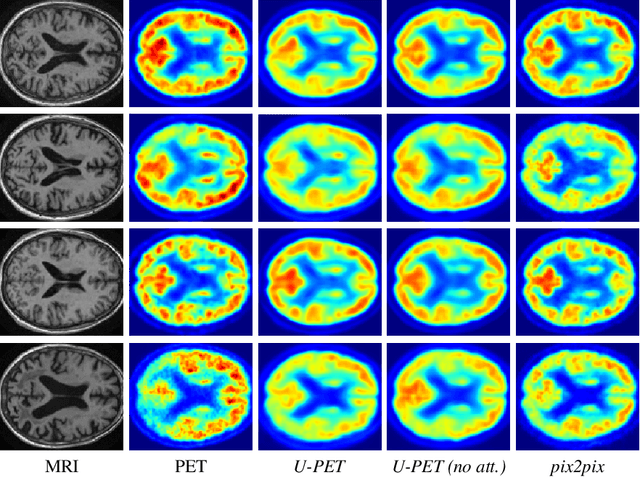
Alzheimer's disease (AD) is the most common cause of dementia. An early detection is crucial for slowing down the disease and mitigating risks related to the progression. While the combination of MRI and FDG-PET is the best image-based tool for diagnosis, FDG-PET is not always available. The reliable detection of Alzheimer's disease with only MRI could be beneficial, especially in regions where FDG-PET might not be affordable for all patients. To this end, we propose a multi-task method based on U-Net that takes T1-weighted MR images as an input to generate synthetic FDG-PET images and classifies the dementia progression of the patient into cognitive normal (CN), cognitive impairment (MCI), and AD. The attention gates used in both task heads can visualize the most relevant parts of the brain, guiding the examiner and adding interpretability. Results show the successful generation of synthetic FDG-PET images and a performance increase in disease classification over the naive single-task baseline.
Virtual embeddings and self-consistency for self-supervised learning
Jun 15, 2022Tariq Bdair, Hossam Abdelhamid, Nassir Navab, Shadi Albarqouni
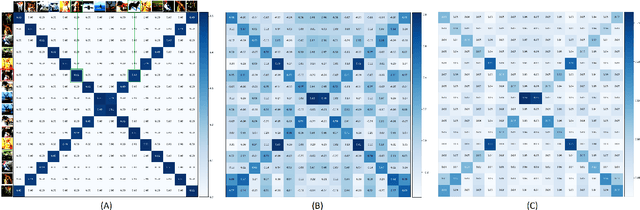
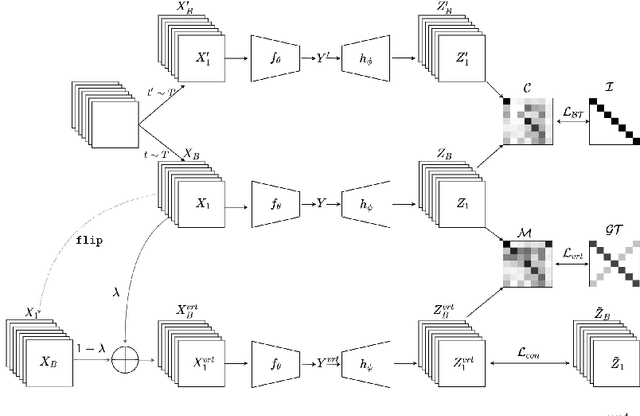
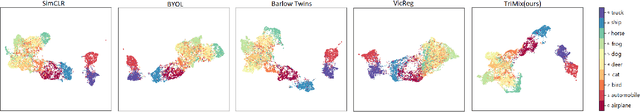
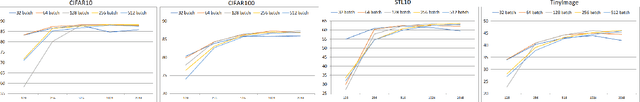

Self-supervised Learning (SSL) has recently gained much attention due to the high cost and data limitation in the training of supervised learning models. The current paradigm in the SSL is to utilize data augmentation at the input space to create different views of the same images and train a model to maximize the representations between similar images and minimize them for different ones. While this approach achieves state-of-the-art (SOTA) results in various downstream tasks, it still lakes the opportunity to investigate the latent space augmentation. This paper proposes TriMix, a novel concept for SSL that generates virtual embeddings through linear interpolation of the data, thus providing the model with novel representations. Our strategy focuses on training the model to extract the original embeddings from virtual ones, hence, better representation learning. Additionally, we propose a self-consistency term that improves the consistency between the virtual and actual embeddings. We validate TriMix on eight benchmark datasets consisting of natural and medical images with an improvement of 2.71% and 0.41% better than the second-best models for both data types. Further, our approach outperformed the current methods in semi-supervised learning, particularly in low data regimes. Besides, our pre-trained models showed better transfer to other datasets.
PRO-TIP: Phantom for RObust automatic ultrasound calibration by TIP detection
Jun 13, 2022Matteo Ronchetti, Julia Rackerseder, Maria Tirindelli, Mehrdad Salehi, Nassir Navab, Wolfgang Wein, Oliver Zettinig
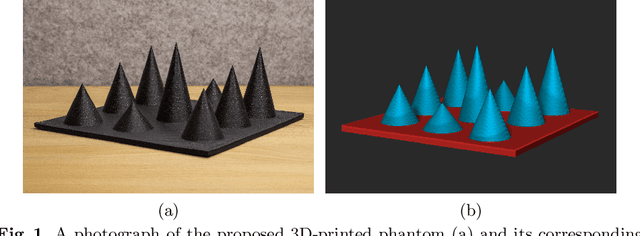
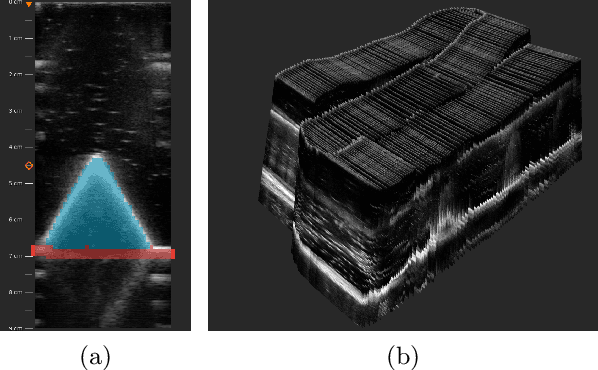

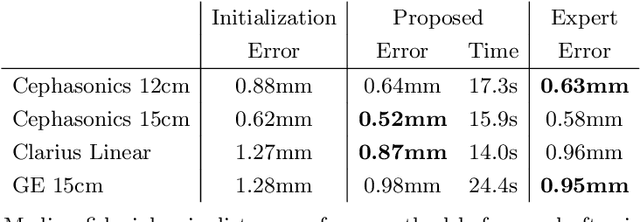
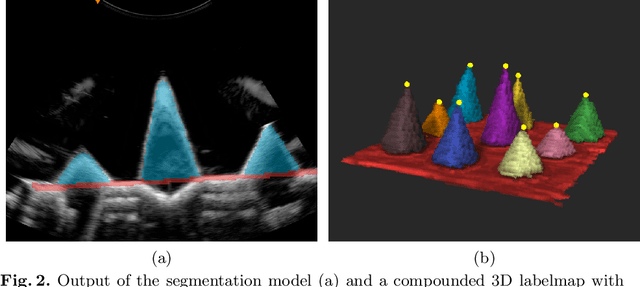

We propose a novel method to automatically calibrate tracked ultrasound probes. To this end we design a custom phantom consisting of nine cones with different heights. The tips are used as key points to be matched between multiple sweeps. We extract them using a convolutional neural network to segment the cones in every ultrasound frame and then track them across the sweep. The calibration is robustly estimated using RANSAC and later refined employing image based techniques. Our phantom can be 3D-printed and offers many advantages over state-of-the-art methods. The phantom design and algorithm code are freely available online. Since our phantom does not require a tracking target on itself, ease of use is improved over currently used techniques. The fully automatic method generalizes to new probes and different vendors, as shown in our experiments. Our approach produces results comparable to calibrations obtained by a domain expert.
BFS-Net: Weakly Supervised Cell Instance Segmentation from Bright-Field Microscopy Z-Stacks
Jun 09, 2022Shervin Dehghani, Benjamin Busam, Nassir Navab, Ali Nasseri
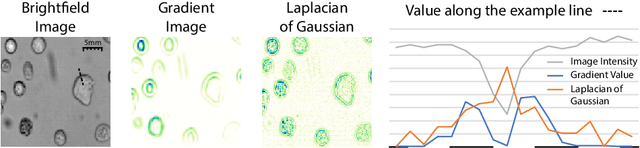
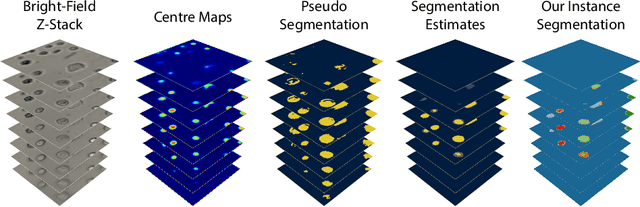
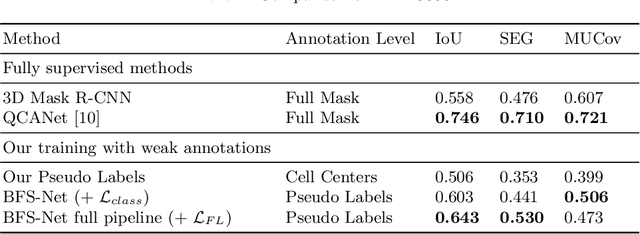
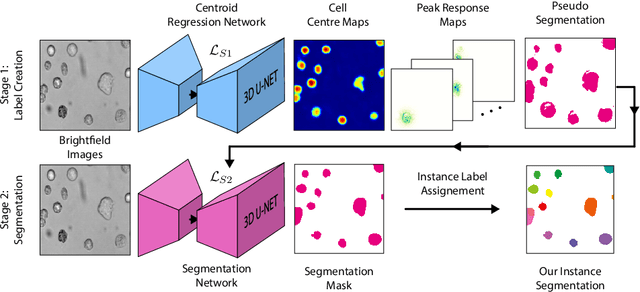

Despite its broad availability, volumetric information acquisition from Bright-Field Microscopy (BFM) is inherently difficult due to the projective nature of the acquisition process. We investigate the prediction of 3D cell instances from a set of BFM Z-Stack images. We propose a novel two-stage weakly supervised method for volumetric instance segmentation of cells which only requires approximate cell centroids annotation. Created pseudo-labels are thereby refined with a novel refinement loss with Z-stack guidance. The evaluations show that our approach can generalize not only to BFM Z-Stack data, but to other 3D cell imaging modalities. A comparison of our pipeline against fully supervised methods indicates that the significant gain in reduced data collection and labelling results in minor performance difference.
PhoCaL: A Multi-Modal Dataset for Category-Level Object Pose Estimation with Photometrically Challenging Objects
May 18, 2022Pengyuan Wang, HyunJun Jung, Yitong Li, Siyuan Shen, Rahul Parthasarathy Srikanth, Lorenzo Garattoni, Sven Meier, Nassir Navab, Benjamin Busam
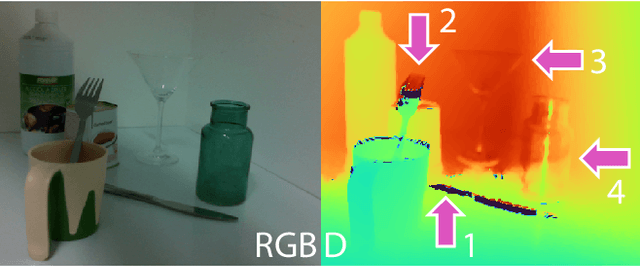
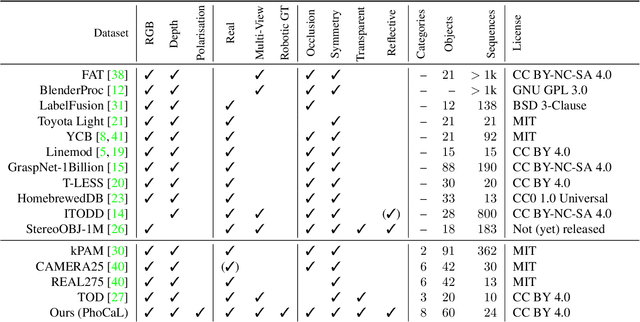



Object pose estimation is crucial for robotic applications and augmented reality. Beyond instance level 6D object pose estimation methods, estimating category-level pose and shape has become a promising trend. As such, a new research field needs to be supported by well-designed datasets. To provide a benchmark with high-quality ground truth annotations to the community, we introduce a multimodal dataset for category-level object pose estimation with photometrically challenging objects termed PhoCaL. PhoCaL comprises 60 high quality 3D models of household objects over 8 categories including highly reflective, transparent and symmetric objects. We developed a novel robot-supported multi-modal (RGB, depth, polarisation) data acquisition and annotation process. It ensures sub-millimeter accuracy of the pose for opaque textured, shiny and transparent objects, no motion blur and perfect camera synchronisation. To set a benchmark for our dataset, state-of-the-art RGB-D and monocular RGB methods are evaluated on the challenging scenes of PhoCaL.
Weakly-supervised Biomechanically-constrained CT/MRI Registration of the Spine
May 16, 2022Bailiang Jian, Mohammad Farid Azampour, Francesca De Benetti, Johannes Oberreuter, Christina Bukas, Alexandra S. Gersing, Sarah C. Foreman, Anna-Sophia Dietrich, Jon Rischewski, Jan S. Kirschke, Nassir Navab, Thomas Wendler
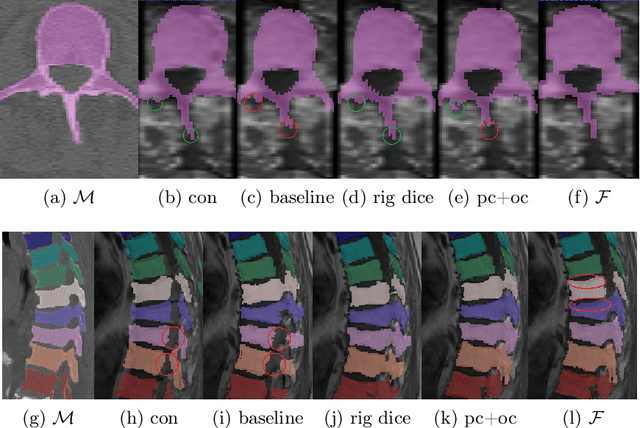
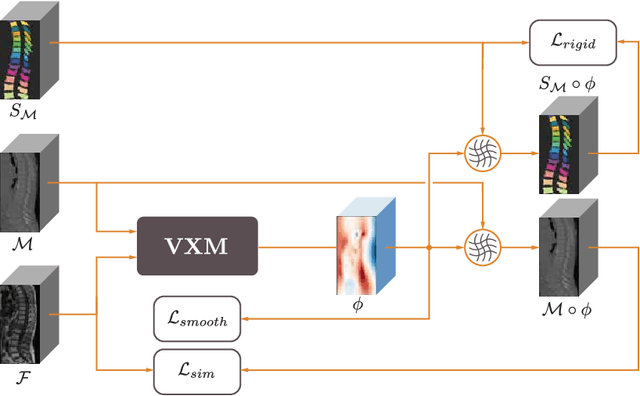
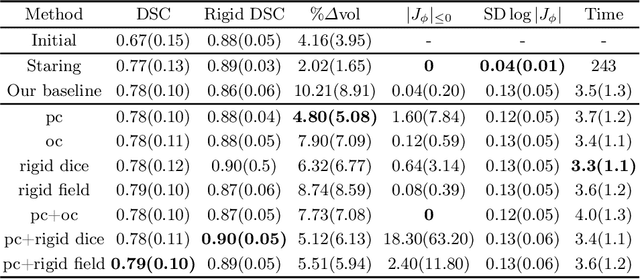
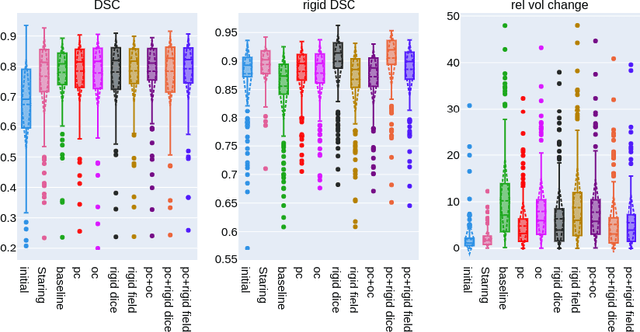

CT and MRI are two of the most informative modalities in spinal diagnostics and treatment planning. CT is useful when analysing bony structures, while MRI gives information about the soft tissue. Thus, fusing the information of both modalities can be very beneficial. Registration is the first step for this fusion. While the soft tissues around the vertebra are deformable, each vertebral body is constrained to move rigidly. We propose a weakly-supervised deep learning framework that preserves the rigidity and the volume of each vertebra while maximizing the accuracy of the registration. To achieve this goal, we introduce anatomy-aware losses for training the network. We specifically design these losses to depend only on the CT label maps since automatic vertebra segmentation in CT gives more accurate results contrary to MRI. We evaluate our method on an in-house dataset of 167 patients. Our results show that adding the anatomy-aware losses increases the plausibility of the inferred transformation while keeping the accuracy untouched.
VesNet-RL: Simulation-based Reinforcement Learning for Real-World US Probe Navigation
May 10, 2022Yuan Bi, Zhongliang Jiang, Yuan Gao, Thomas Wendler, Angelos Karlas, Nassir Navab
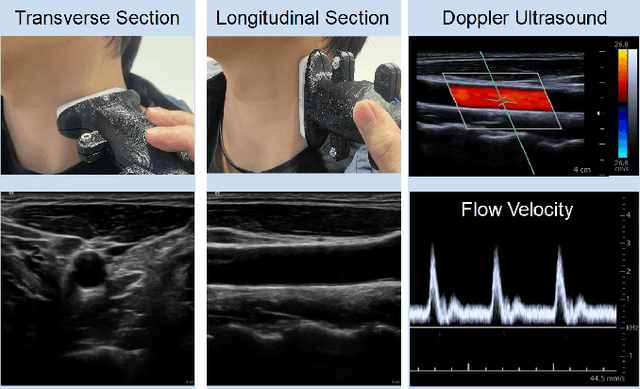
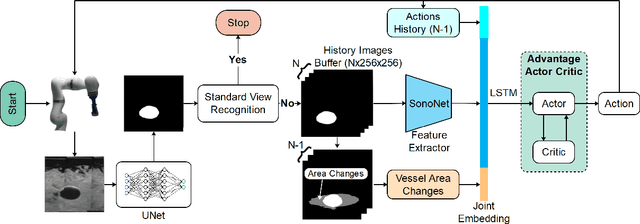
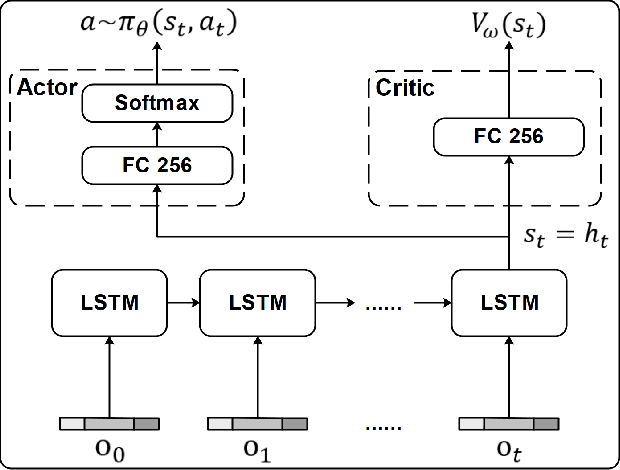
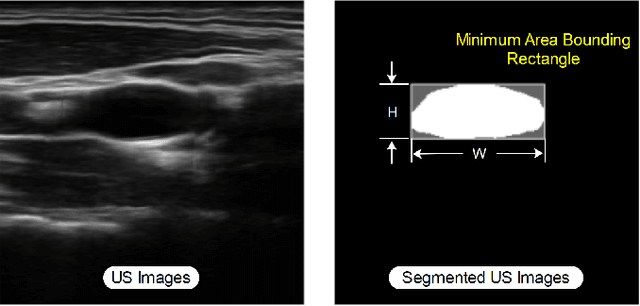
Ultrasound (US) is one of the most common medical imaging modalities since it is radiation-free, low-cost, and real-time. In freehand US examinations, sonographers often navigate a US probe to visualize standard examination planes with rich diagnostic information. However, reproducibility and stability of the resulting images often suffer from intra- and inter-operator variation. Reinforcement learning (RL), as an interaction-based learning method, has demonstrated its effectiveness in visual navigating tasks; however, RL is limited in terms of generalization. To address this challenge, we propose a simulation-based RL framework for real-world navigation of US probes towards the standard longitudinal views of vessels. A UNet is used to provide binary masks from US images; thereby, the RL agent trained on simulated binary vessel images can be applied in real scenarios without further training. To accurately characterize actual states, a multi-modality state representation structure is introduced to facilitate the understanding of environments. Moreover, considering the characteristics of vessels, a novel standard view recognition approach based on the minimum bounding rectangle is proposed to terminate the searching process. To evaluate the effectiveness of the proposed method, the trained policy is validated virtually on 3D volumes of a volunteer's in-vivo carotid artery, and physically on custom-designed gel phantoms using robotic US. The results demonstrate that proposed approach can effectively and accurately navigate the probe towards the longitudinal view of vessels.
Affective Medical Estimation and Decision Making via Visualized Learning and Deep Learning
May 09, 2022Mohammad Eslami, Solale Tabarestani, Ehsan Adeli, Glyn Elwyn, Tobias Elze, Mengyu Wang, Nazlee Zebardast, Nassir Navab, Malek Adjouadi
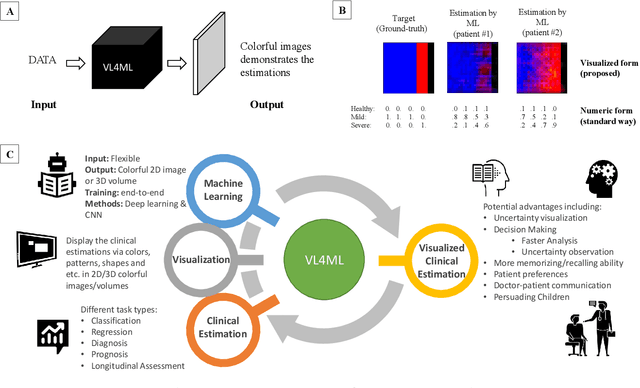
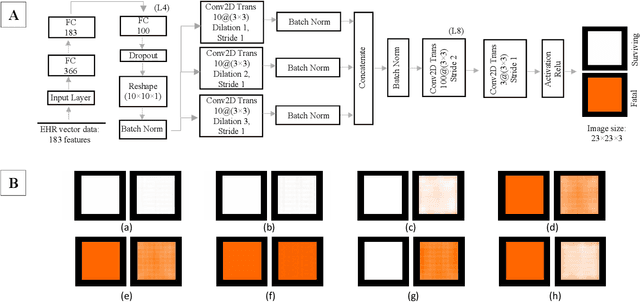
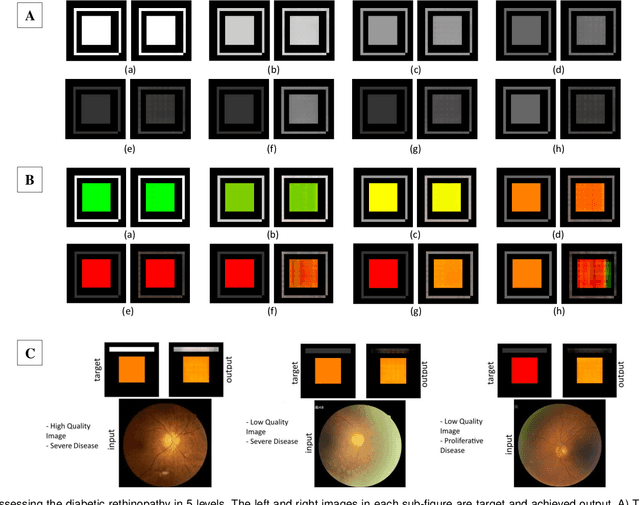
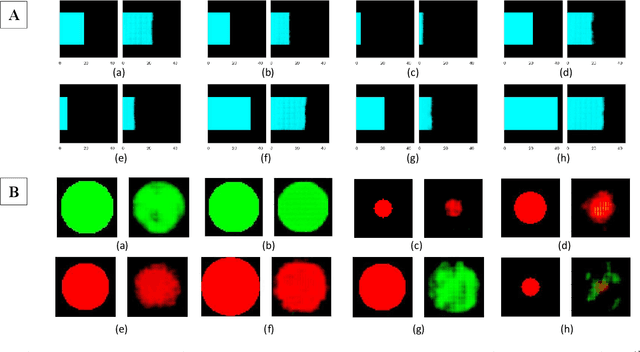
With the advent of sophisticated machine learning (ML) techniques and the promising results they yield, especially in medical applications, where they have been investigated for different tasks to enhance the decision-making process. Since visualization is such an effective tool for human comprehension, memorization, and judgment, we have presented a first-of-its-kind estimation approach we refer to as Visualized Learning for Machine Learning (VL4ML) that not only can serve to assist physicians and clinicians in making reasoned medical decisions, but it also allows to appreciate the uncertainty visualization, which could raise incertitude in making the appropriate classification or prediction. For the proof of concept, and to demonstrate the generalized nature of this visualized estimation approach, five different case studies are examined for different types of tasks including classification, regression, and longitudinal prediction. A survey analysis with more than 100 individuals is also conducted to assess users' feedback on this visualized estimation method. The experiments and the survey demonstrate the practical merits of the VL4ML that include: (1) appreciating visually clinical/medical estimations; (2) getting closer to the patients' preferences; (3) improving doctor-patient communication, and (4) visualizing the uncertainty introduced through the black box effect of the deployed ML algorithm. All the source codes are shared via a GitHub repository.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge