Walter Simson
Ultrasound Autofocusing: Common Midpoint Phase Error Optimization via Differentiable Beamforming
Oct 03, 2024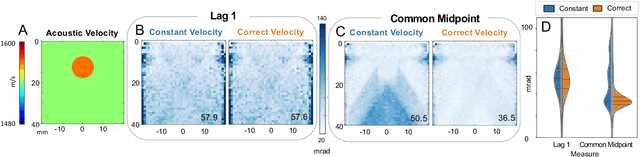
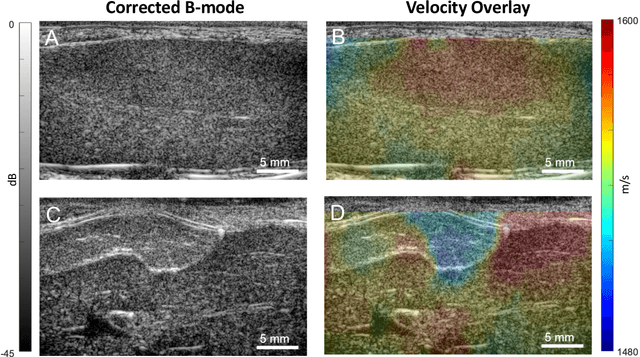
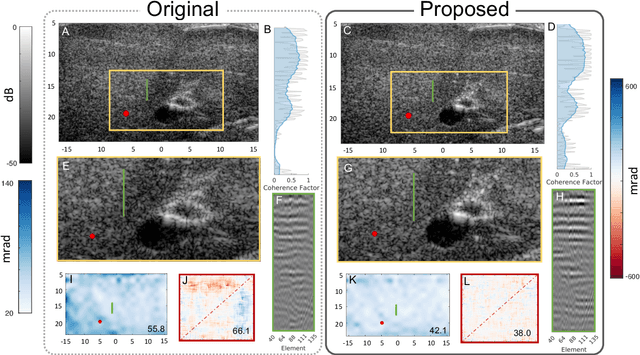

Abstract:Wavefield imaging reconstructs physical properties from wavefield measurements across an aperture, using modalities like radar, optics, sonar, seismic, and ultrasound imaging. Propagation of a wavefront from unknown sources through heterogeneous media causes phase aberrations that degrade the coherence of the wavefront leading to reduced image resolution and contrast. Adaptive imaging techniques attempt to correct phase aberration and restore coherence leading to improved focus. We propose an autofocusing paradigm for aberration correction in ultrasound imaging by fitting an acoustic velocity field to pressure measurements, via optimization of the common midpoint phase error (CMPE), using a straight-ray wave propagation model for beamforming in diffusely scattering media. We show that CMPE induced by heterogeneous acoustic velocity is a robust measure of phase aberration that can be used for acoustic autofocusing. CMPE is optimized iteratively using a differentiable beamforming approach to simultaneously improve the image focus while estimating the acoustic velocity field of the interrogated medium. The approach relies solely on wavefield measurements using a straight-ray integral solution of the two-way time-of-flight without explicit numerical time-stepping models of wave propagation. We demonstrate method performance through in silico simulations, in vitro phantom measurements, and in vivo mammalian models, showing practical applications in distributed aberration quantification, correction, and velocity estimation for medical ultrasound autofocusing.
PHOCUS: Physics-Based Deconvolution for Ultrasound Resolution Enhancement
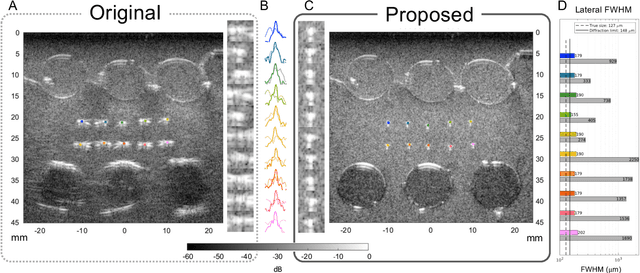
Aug 07, 2024Abstract:Ultrasound is widely used in medical diagnostics allowing for accessible and powerful imaging but suffers from resolution limitations due to diffraction and the finite aperture of the imaging system, which restricts diagnostic use. The impulse function of an ultrasound imaging system is called the point spread function (PSF), which is convolved with the spatial distribution of reflectors in the image formation process. Recovering high-resolution reflector distributions by removing image distortions induced by the convolution process improves image clarity and detail. Conventionally, deconvolution techniques attempt to rectify the imaging system's dependent PSF, working directly on the radio-frequency (RF) data. However, RF data is often not readily accessible. Therefore, we introduce a physics-based deconvolution process using a modeled PSF, working directly on the more commonly available B-mode images. By leveraging Implicit Neural Representations (INRs), we learn a continuous mapping from spatial locations to their respective echogenicity values, effectively compensating for the discretized image space. Our contribution consists of a novel methodology for retrieving a continuous echogenicity map directly from a B-mode image through a differentiable physics-based rendering pipeline for ultrasound resolution enhancement. We qualitatively and quantitatively evaluate our approach on synthetic data, demonstrating improvements over traditional methods in metrics such as PSNR and SSIM. Furthermore, we show qualitative enhancements on an ultrasound phantom and an in-vivo acquisition of a carotid artery.
Implicit Neural Representations for Breathing-compensated Volume Reconstruction in Robotic Ultrasound Aorta Screening
Nov 08, 2023Abstract:Ultrasound (US) imaging is widely used in diagnosing and staging abdominal diseases due to its lack of non-ionizing radiation and prevalent availability. However, significant inter-operator variability and inconsistent image acquisition hinder the widespread adoption of extensive screening programs. Robotic ultrasound systems have emerged as a promising solution, offering standardized acquisition protocols and the possibility of automated acquisition. Additionally, these systems enable access to 3D data via robotic tracking, enhancing volumetric reconstruction for improved ultrasound interpretation and precise disease diagnosis. However, the interpretability of 3D US reconstruction of abdominal images can be affected by the patient's breathing motion. This study introduces a method to compensate for breathing motion in 3D US compounding by leveraging implicit neural representations. Our approach employs a robotic ultrasound system for automated screenings. To demonstrate the method's effectiveness, we evaluate our proposed method for the diagnosis and monitoring of abdominal aorta aneurysms as a representative use case. Our experiments demonstrate that our proposed pipeline facilitates robust automated robotic acquisition, mitigating artifacts from breathing motion, and yields smoother 3D reconstructions for enhanced screening and medical diagnosis.
LOTUS: Learning to Optimize Task-based US representations
Jul 29, 2023



Abstract:Anatomical segmentation of organs in ultrasound images is essential to many clinical applications, particularly for diagnosis and monitoring. Existing deep neural networks require a large amount of labeled data for training in order to achieve clinically acceptable performance. Yet, in ultrasound, due to characteristic properties such as speckle and clutter, it is challenging to obtain accurate segmentation boundaries, and precise pixel-wise labeling of images is highly dependent on the expertise of physicians. In contrast, CT scans have higher resolution and improved contrast, easing organ identification. In this paper, we propose a novel approach for learning to optimize task-based ultra-sound image representations. Given annotated CT segmentation maps as a simulation medium, we model acoustic propagation through tissue via ray-casting to generate ultrasound training data. Our ultrasound simulator is fully differentiable and learns to optimize the parameters for generating physics-based ultrasound images guided by the downstream segmentation task. In addition, we train an image adaptation network between real and simulated images to achieve simultaneous image synthesis and automatic segmentation on US images in an end-to-end training setting. The proposed method is evaluated on aorta and vessel segmentation tasks and shows promising quantitative results. Furthermore, we also conduct qualitative results of optimized image representations on other organs.
Self-Supervised Learning for Physiologically-Based Pharmacokinetic Modeling in Dynamic PET
May 17, 2023



Abstract:Dynamic positron emission tomography imaging (dPET) provides temporally resolved images of a tracer enabling a quantitative measure of physiological processes. Voxel-wise physiologically-based pharmacokinetic (PBPK) modeling of the time activity curves (TAC) can provide relevant diagnostic information for clinical workflow. Conventional fitting strategies for TACs are slow and ignore the spatial relation between neighboring voxels. We train a spatio-temporal UNet to estimate the kinetic parameters given TAC from F-18-fluorodeoxyglucose (FDG) dPET. This work introduces a self-supervised loss formulation to enforce the similarity between the measured TAC and those generated with the learned kinetic parameters. Our method provides quantitatively comparable results at organ-level to the significantly slower conventional approaches, while generating pixel-wise parametric images which are consistent with expected physiology. To the best of our knowledge, this is the first self-supervised network that allows voxel-wise computation of kinetic parameters consistent with a non-linear kinetic model. The code will become publicly available upon acceptance.
Spotlight on nerves: Portable multispectral optoacoustic imaging of peripheral nerve vascularization and morphology
Jul 28, 2022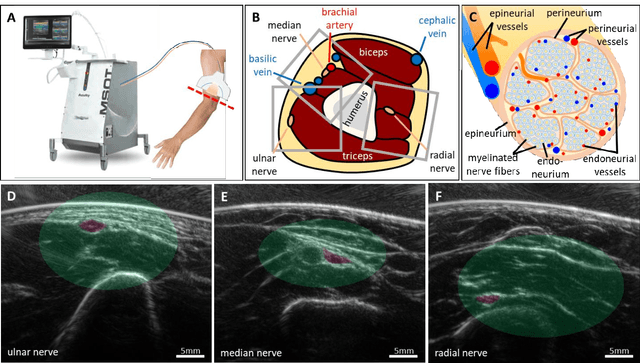

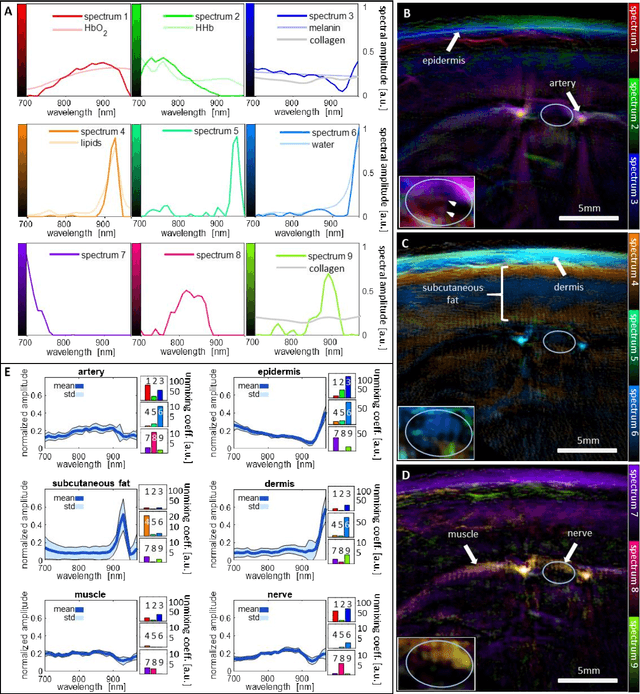
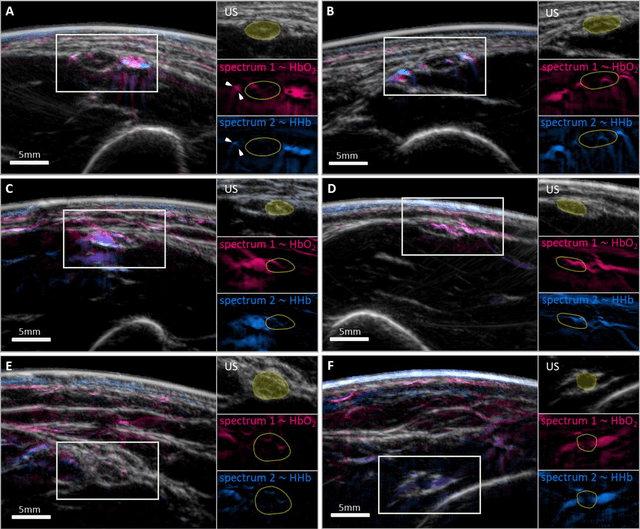
Abstract:Various morphological and functional parameters of peripheral nerves and their vascular supply are indicative of pathological changes due to injury or disease. Based on recent improvements in optoacoustic image quality, we explore the ability of multispectral optoacoustic tomography, in tandem with ultrasound imaging (OPUS), to investigate the vascular environment and morphology of peripheral nerves in vivo in a pilot study on healthy volunteers. We showcase the unique ability of optoacoustic imaging to visualize the vasa nervorum by observing intraneurial vessels in healthy nerves in vivo for the first time. In addition, we demonstrate that the label-free spectral optoacoustic contrast of the perfused connective tissue of peripheral nerves can be linked to the endogenous contrast of haemoglobin and collagen. We introduce metrics to analyze the composition of tissue based on its optoacoustic contrast and show that the high-resolution spectral contrast reveals specific differences between nervous tissue and reference tissue in the nerve's surrounding. We discuss how this showcased extraction of peripheral nerve characteristics using multispectral optoacoustic and ultrasound imaging can offer new insights into the pathophysiology of nerve damage and neuropathies, for example, in the context of diabetes.
CACTUSS: Common Anatomical CT-US Space for US examinations
Jul 18, 2022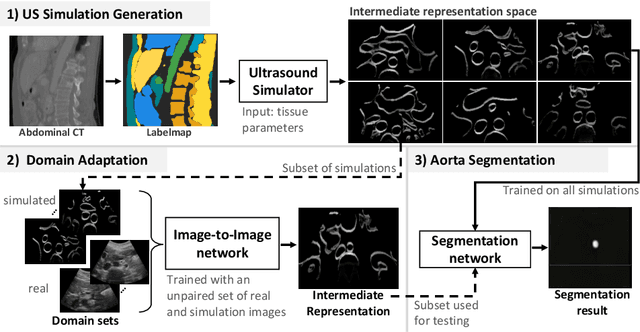
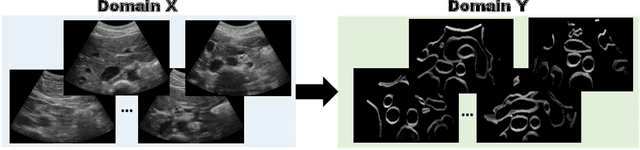
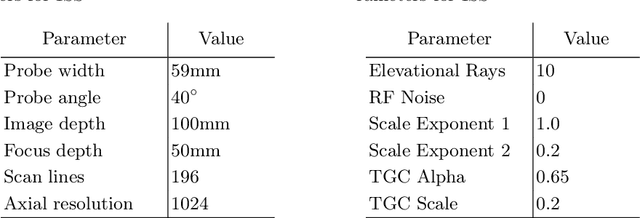
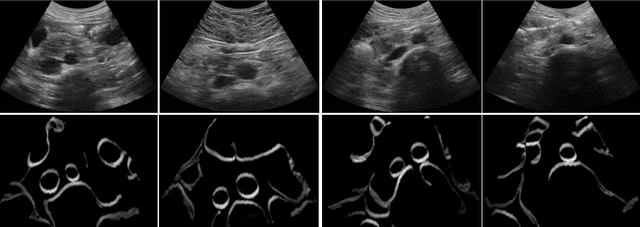
Abstract:Abdominal aortic aneurysm (AAA) is a vascular disease in which a section of the aorta enlarges, weakening its walls and potentially rupturing the vessel. Abdominal ultrasound has been utilized for diagnostics, but due to its limited image quality and operator dependency, CT scans are usually required for monitoring and treatment planning. Recently, abdominal CT datasets have been successfully utilized to train deep neural networks for automatic aorta segmentation. Knowledge gathered from this solved task could therefore be leveraged to improve US segmentation for AAA diagnosis and monitoring. To this end, we propose CACTUSS: a common anatomical CT-US space, which acts as a virtual bridge between CT and US modalities to enable automatic AAA screening sonography. CACTUSS makes use of publicly available labelled data to learn to segment based on an intermediary representation that inherits properties from both US and CT. We train a segmentation network in this new representation and employ an additional image-to-image translation network which enables our model to perform on real B-mode images. Quantitative comparisons against fully supervised methods demonstrate the capabilities of CACTUSS in terms of Dice Score and diagnostic metrics, showing that our method also meets the clinical requirements for AAA scanning and diagnosis.
Rethinking Ultrasound Augmentation: A Physics-Inspired Approach
May 05, 2021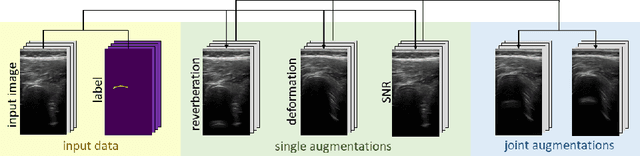
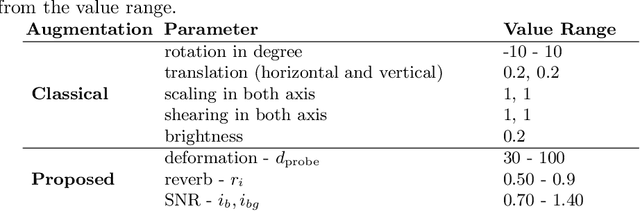

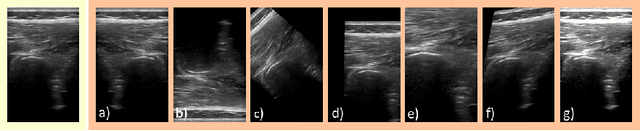
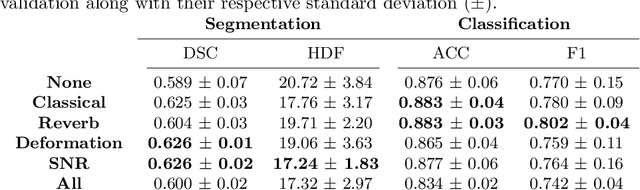
Abstract:Medical Ultrasound (US), despite its wide use, is characterized by artifacts and operator dependency. Those attributes hinder the gathering and utilization of US datasets for the training of Deep Neural Networks used for Computer-Assisted Intervention Systems. Data augmentation is commonly used to enhance model generalization and performance. However, common data augmentation techniques, such as affine transformations do not align with the physics of US and, when used carelessly can lead to unrealistic US images. To this end, we propose a set of physics-inspired transformations, including deformation, reverb and Signal-to-Noise Ratio, that we apply on US B-mode images for data augmentation. We evaluate our method on a new spine US dataset for the tasks of bone segmentation and classification.
Ultrasound-Guided Robotic Navigation with Deep Reinforcement Learning
Apr 07, 2020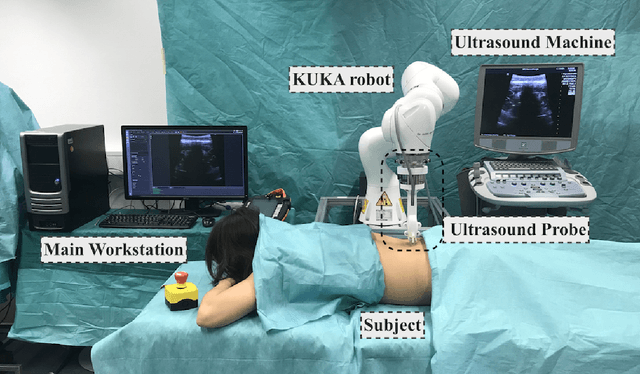
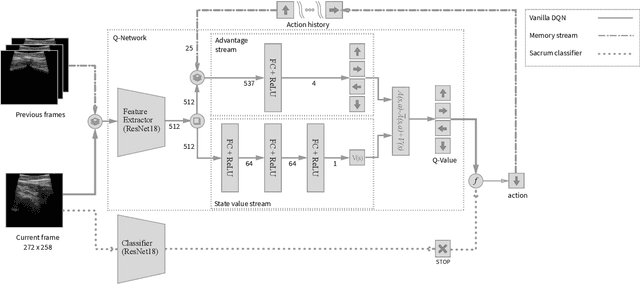
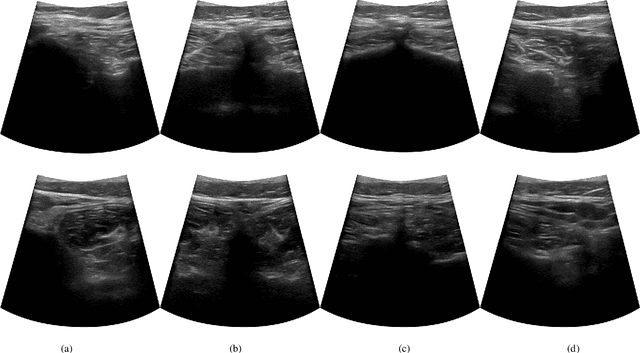
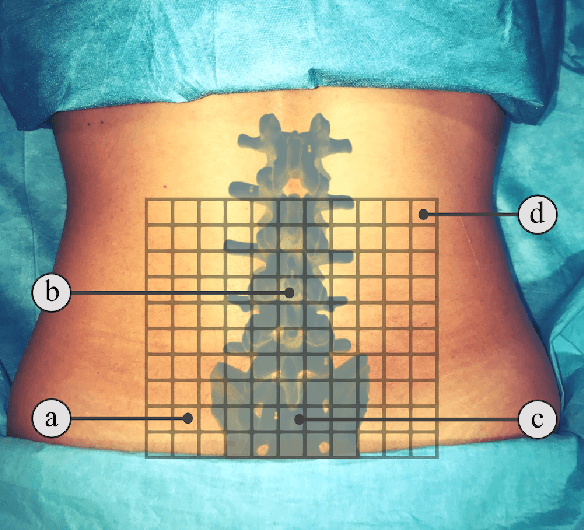
Abstract:In this paper we introduce the first reinforcement learning (RL) based robotic navigation method which utilizes ultrasound (US) images as an input. Our approach combines state-of-the-art RL techniques, specifically deep Q-networks (DQN) with memory buffers and a binary classifier for deciding when to terminate the task. Our method is trained and evaluated on an in-house collected data-set of 34 volunteers and when compared to pure RL and supervised learning (SL) techniques, it performs substantially better, which highlights the suitability of RL navigation for US-guided procedures. When testing our proposed model, we obtained a 82.91% chance of navigating correctly to the sacrum from 165 different starting positions on 5 different unseen simulated environments.
TeCNO: Surgical Phase Recognition with Multi-Stage Temporal Convolutional Networks
Mar 24, 2020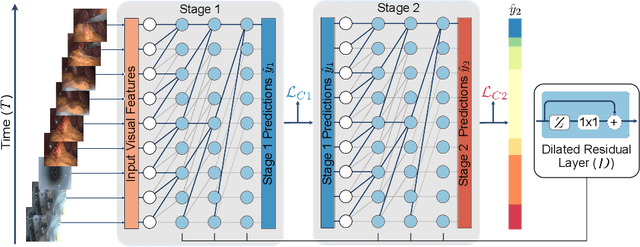
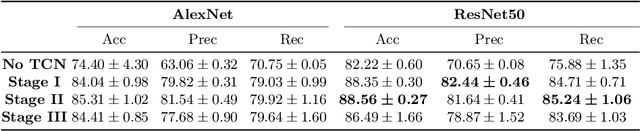
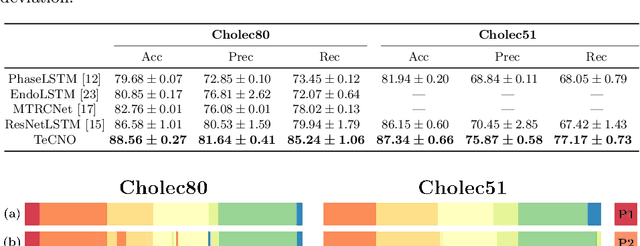
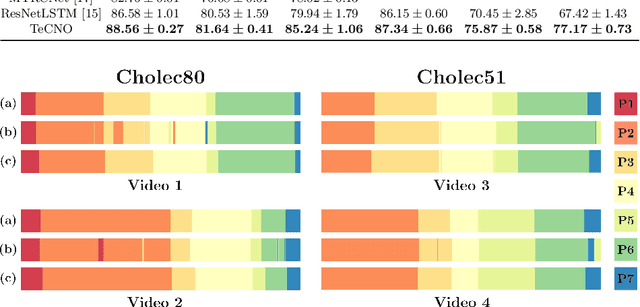
Abstract:Automatic surgical phase recognition is a challenging and crucial task with the potential to improve patient safety and become an integral part of intra-operative decision-support systems. In this paper, we propose, for the first time in workflow analysis, a Multi-Stage Temporal Convolutional Network (MS-TCN) that performs hierarchical prediction refinement for surgical phase recognition. Causal, dilated convolutions allow for a large receptive field and online inference with smooth predictions even during ambiguous transitions. Our method is thoroughly evaluated on two datasets of laparoscopic cholecystectomy videos with and without the use of additional surgical tool information. Outperforming various state-of-the-art LSTM approaches, we verify the suitability of the proposed causal MS-TCN for surgical phase recognition.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge