Lingyun Huang
Weakly-Supervised Universal Lesion Segmentation with Regional Level Set Loss
May 03, 2021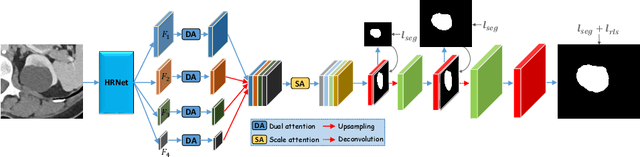
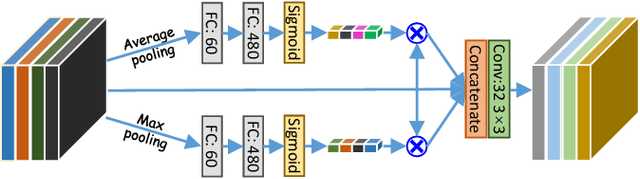
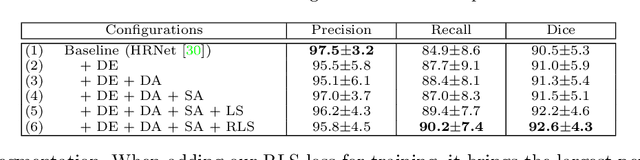
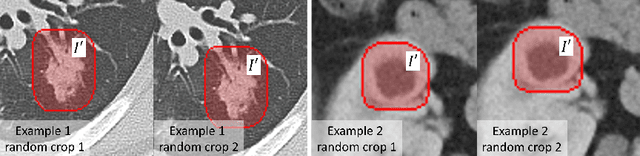
Abstract:Accurately segmenting a variety of clinically significant lesions from whole body computed tomography (CT) scans is a critical task on precision oncology imaging, denoted as universal lesion segmentation (ULS). Manual annotation is the current clinical practice, being highly time-consuming and inconsistent on tumor's longitudinal assessment. Effectively training an automatic segmentation model is desirable but relies heavily on a large number of pixel-wise labelled data. Existing weakly-supervised segmentation approaches often struggle with regions nearby the lesion boundaries. In this paper, we present a novel weakly-supervised universal lesion segmentation method by building an attention enhanced model based on the High-Resolution Network (HRNet), named AHRNet, and propose a regional level set (RLS) loss for optimizing lesion boundary delineation. AHRNet provides advanced high-resolution deep image features by involving a decoder, dual-attention and scale attention mechanisms, which are crucial to performing accurate lesion segmentation. RLS can optimize the model reliably and effectively in a weakly-supervised fashion, forcing the segmentation close to lesion boundary. Extensive experimental results demonstrate that our method achieves the best performance on the publicly large-scale DeepLesion dataset and a hold-out test set.
Scalable Semi-supervised Landmark Localization for X-ray Images using Few-shot Deep Adaptive Graph
Apr 29, 2021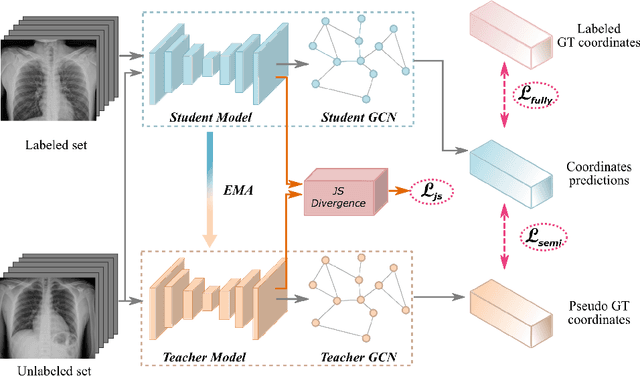
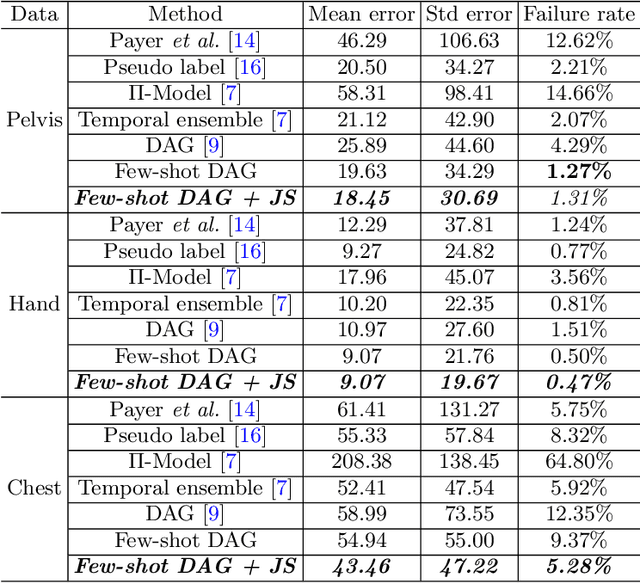
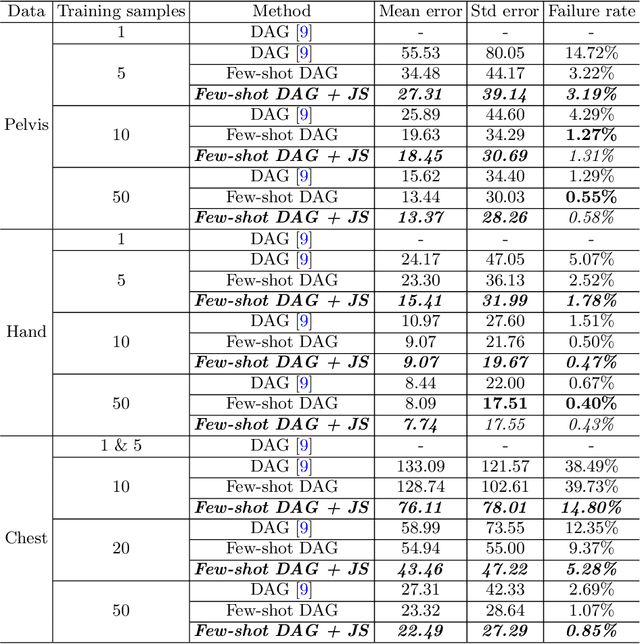
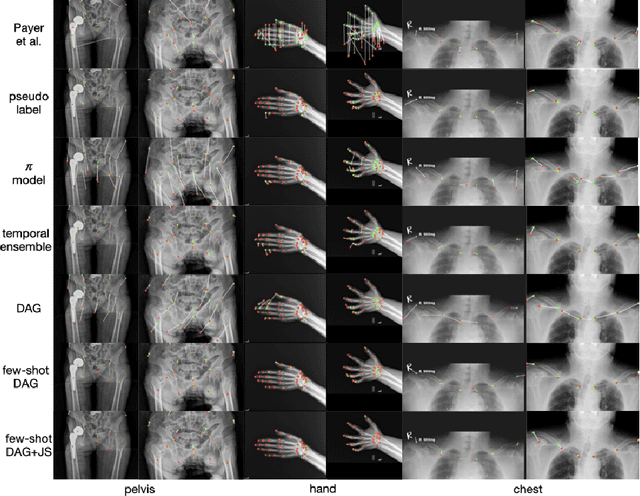
Abstract:Landmark localization plays an important role in medical image analysis. Learning based methods, including CNN and GCN, have demonstrated the state-of-the-art performance. However, most of these methods are fully-supervised and heavily rely on manual labeling of a large training dataset. In this paper, based on a fully-supervised graph-based method, DAG, we proposed a semi-supervised extension of it, termed few-shot DAG, \ie five-shot DAG. It first trains a DAG model on the labeled data and then fine-tunes the pre-trained model on the unlabeled data with a teacher-student SSL mechanism. In addition to the semi-supervised loss, we propose another loss using JS divergence to regulate the consistency of the intermediate feature maps. We extensively evaluated our method on pelvis, hand and chest landmark detection tasks. Our experiment results demonstrate consistent and significant improvements over previous methods.
Semi-Supervised Learning for Bone Mineral Density Estimation in Hip X-ray Images
Mar 24, 2021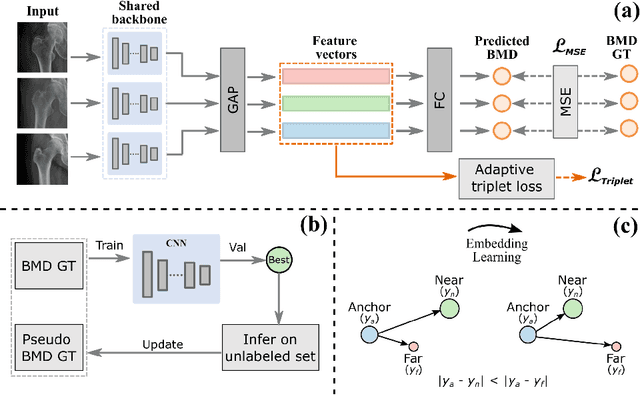

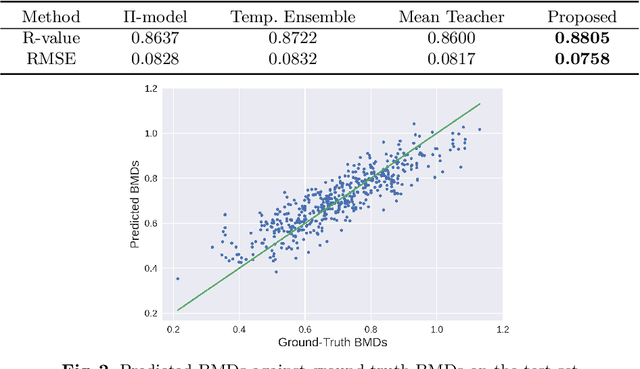
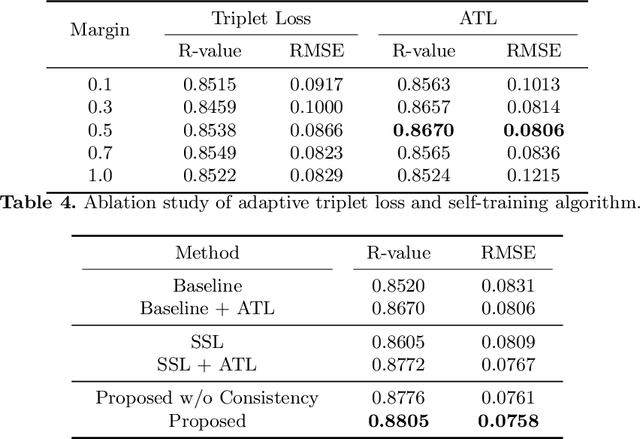
Abstract:Bone mineral density (BMD) is a clinically critical indicator of osteoporosis, usually measured by dual-energy X-ray absorptiometry (DEXA). Due to the limited accessibility of DEXA machines and examinations, osteoporosis is often under-diagnosed and under-treated, leading to increased fragility fracture risks. Thus it is highly desirable to obtain BMDs with alternative cost-effective and more accessible medical imaging examinations such as X-ray plain films. In this work, we formulate the BMD estimation from plain hip X-ray images as a regression problem. Specifically, we propose a new semi-supervised self-training algorithm to train the BMD regression model using images coupled with DEXA measured BMDs and unlabeled images with pseudo BMDs. Pseudo BMDs are generated and refined iteratively for unlabeled images during self-training. We also present a novel adaptive triplet loss to improve the model's regression accuracy. On an in-house dataset of 1,090 images (819 unique patients), our BMD estimation method achieves a high Pearson correlation coefficient of 0.8805 to ground-truth BMDs. It offers good feasibility to use the more accessible and cheaper X-ray imaging for opportunistic osteoporosis screening.
Hetero-Modal Learning and Expansive Consistency Constraints for Semi-Supervised Detection from Multi-Sequence Data
Mar 24, 2021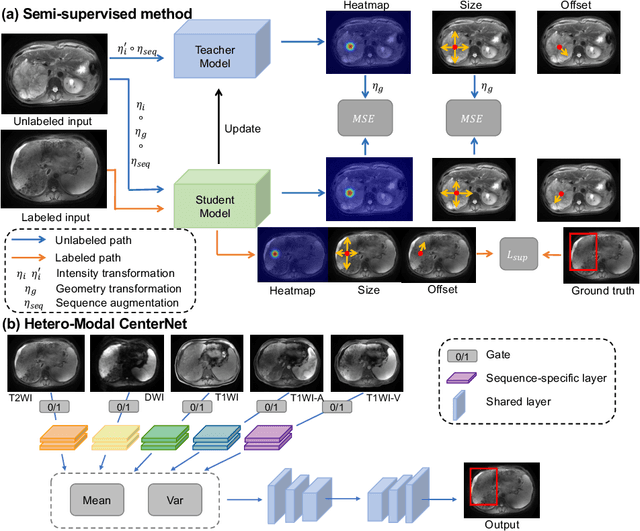
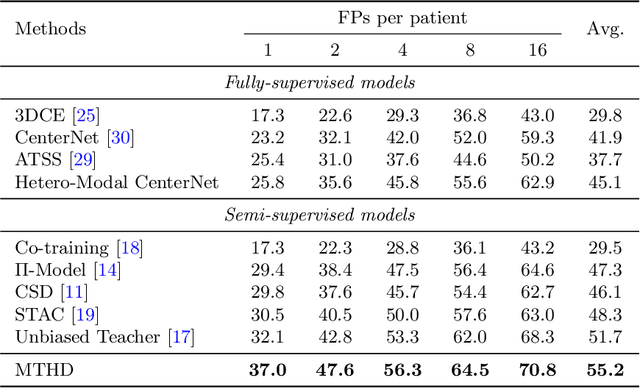
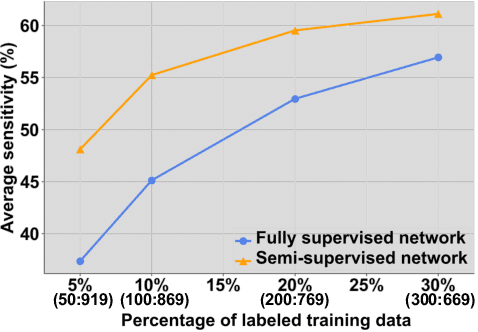
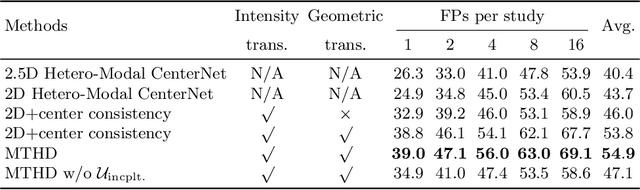
Abstract:Lesion detection serves a critical role in early diagnosis and has been well explored in recent years due to methodological advancesand increased data availability. However, the high costs of annotations hinder the collection of large and completely labeled datasets, motivating semi-supervised detection approaches. In this paper, we introduce mean teacher hetero-modal detection (MTHD), which addresses two important gaps in current semi-supervised detection. First, it is not obvious how to enforce unlabeled consistency constraints across the very different outputs of various detectors, which has resulted in various compromises being used in the state of the art. Using an anchor-free framework, MTHD formulates a mean teacher approach without such compromises, enforcing consistency on the soft-output of object centers and size. Second, multi-sequence data is often critical, e.g., for abdominal lesion detection, but unlabeled data is often missing sequences. To deal with this, MTHD incorporates hetero-modal learning in its framework. Unlike prior art, MTHD is able to incorporate an expansive set of consistency constraints that include geometric transforms and random sequence combinations. We train and evaluate MTHD on liver lesion detection using the largest MR lesion dataset to date (1099 patients with >5000 volumes). MTHD surpasses the best fully-supervised and semi-supervised competitors by 10.1% and 3.5%, respectively, in average sensitivity.
Sequential Learning on Liver Tumor Boundary Semantics and Prognostic Biomarker Mining
Mar 09, 2021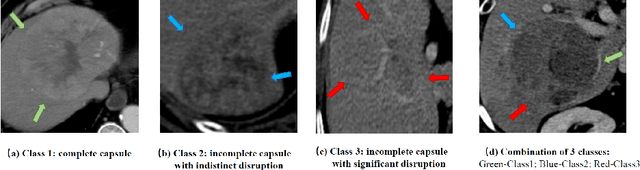
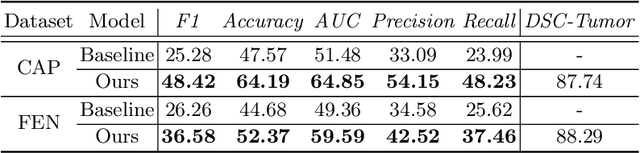
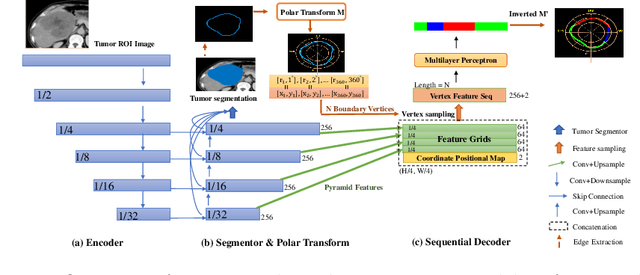
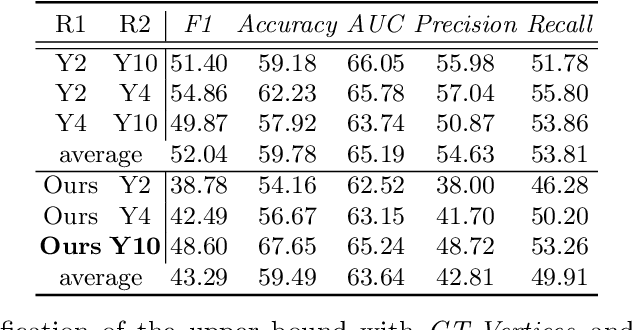
Abstract:The boundary of tumors (hepatocellular carcinoma, or HCC) contains rich semantics: capsular invasion, visibility, smoothness, folding and protuberance, etc. Capsular invasion on tumor boundary has proven to be clinically correlated with the prognostic indicator, microvascular invasion (MVI). Investigating tumor boundary semantics has tremendous clinical values. In this paper, we propose the first and novel computational framework that disentangles the task into two components: spatial vertex localization and sequential semantic classification. (1) A HCC tumor segmentor is built for tumor mask boundary extraction, followed by polar transform representing the boundary with radius and angle. Vertex generator is used to produce fixed-length boundary vertices where vertex features are sampled on the corresponding spatial locations. (2) The sampled deep vertex features with positional embedding are mapped into a sequential space and decoded by a multilayer perceptron (MLP) for semantic classification. Extensive experiments on tumor capsule semantics demonstrate the effectiveness of our framework. Mining the correlation between the boundary semantics and MVI status proves the feasibility to integrate this boundary semantics as a valid HCC prognostic biomarker.
Knowledge Distillation with Adaptive Asymmetric Label Sharpening for Semi-supervised Fracture Detection in Chest X-rays
Dec 30, 2020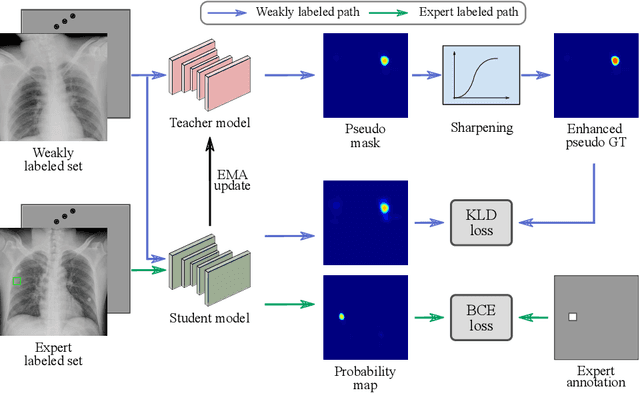
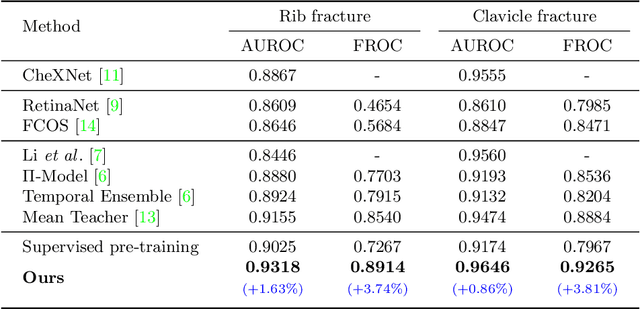
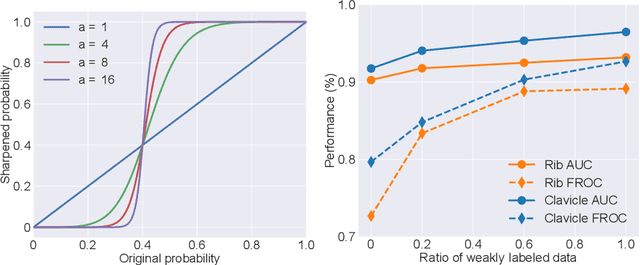
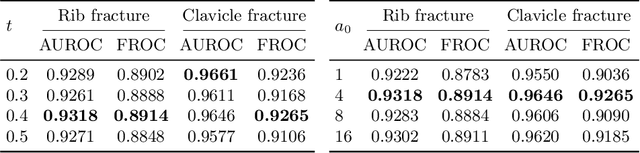
Abstract:Exploiting available medical records to train high performance computer-aided diagnosis (CAD) models via the semi-supervised learning (SSL) setting is emerging to tackle the prohibitively high labor costs involved in large-scale medical image annotations. Despite the extensive attentions received on SSL, previous methods failed to 1) account for the low disease prevalence in medical records and 2) utilize the image-level diagnosis indicated from the medical records. Both issues are unique to SSL for CAD models. In this work, we propose a new knowledge distillation method that effectively exploits large-scale image-level labels extracted from the medical records, augmented with limited expert annotated region-level labels, to train a rib and clavicle fracture CAD model for chest X-ray (CXR). Our method leverages the teacher-student model paradigm and features a novel adaptive asymmetric label sharpening (AALS) algorithm to address the label imbalance problem that specially exists in medical domain. Our approach is extensively evaluated on all CXR (N = 65,845) from the trauma registry of anonymous hospital over a period of 9 years (2008-2016), on the most common rib and clavicle fractures. The experiment results demonstrate that our method achieves the state-of-the-art fracture detection performance, i.e., an area under receiver operating characteristic curve (AUROC) of 0.9318 and a free-response receiver operating characteristic (FROC) score of 0.8914 on the rib fractures, significantly outperforming previous approaches by an AUROC gap of 1.63% and an FROC improvement by 3.74%. Consistent performance gains are also observed for clavicle fracture detection.
Fully-Automated Liver Tumor Localization and Characterization from Multi-Phase MR Volumes Using Key-Slice ROI Parsing: A Physician-Inspired Approach
Dec 15, 2020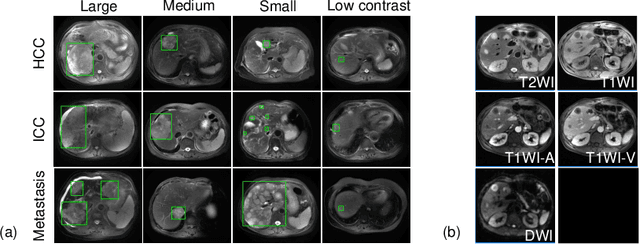
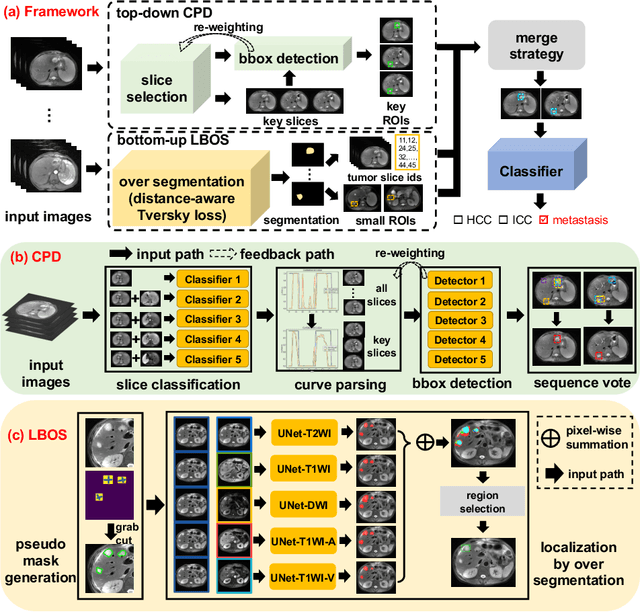
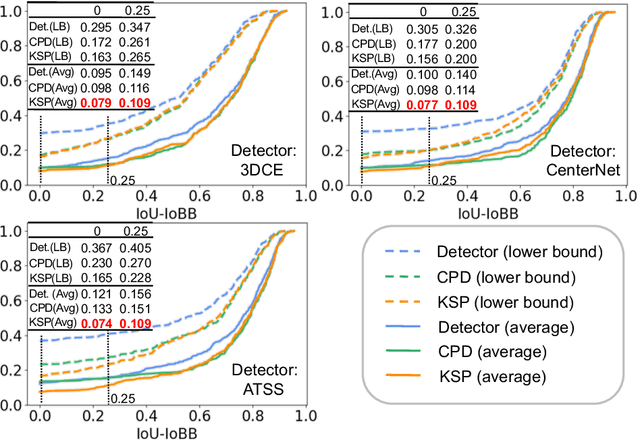

Abstract:Using radiological scans to identify liver tumors is crucial for proper patient treatment. This is highly challenging, as top radiologists only achieve F1 scores of roughly 80% (hepatocellular carcinoma (HCC) vs. others) with only moderate inter-rater agreement, even when using multi-phase magnetic resonance (MR) imagery. Thus, there is great impetus for computer-aided diagnosis (CAD) solutions. A critical challengeis to reliably parse a 3D MR volume to localize diagnosable regions of interest (ROI). In this paper, we break down this problem using a key-slice parser (KSP), which emulates physician workflows by first identifying key slices and then localize their corresponding key ROIs. Because performance demands are so extreme, (not to miss any key ROI),our KSP integrates complementary modules--top-down classification-plus-detection (CPD) and bottom-up localization-by-over-segmentation(LBOS). The CPD uses a curve-parsing and detection confidence to re-weight classifier confidences. The LBOS uses over-segmentation to flag CPD failure cases and provides its own ROIs. For scalability, LBOS is only weakly trained on pseudo-masks using a new distance-aware Tversky loss. We evaluate our approach on the largest multi-phase MR liver lesion test dataset to date (430 biopsy-confirmed patients). Experiments demonstrate that our KSP can localize diagnosable ROIs with high reliability (85% patients have an average overlap of >= 40% with the ground truth). Moreover, we achieve an HCC vs. others F1 score of 0.804, providing a fully-automated CAD solution comparable with top human physicians.
3D Graph Anatomy Geometry-Integrated Network for Pancreatic Mass Segmentation, Diagnosis, and Quantitative Patient Management
Dec 08, 2020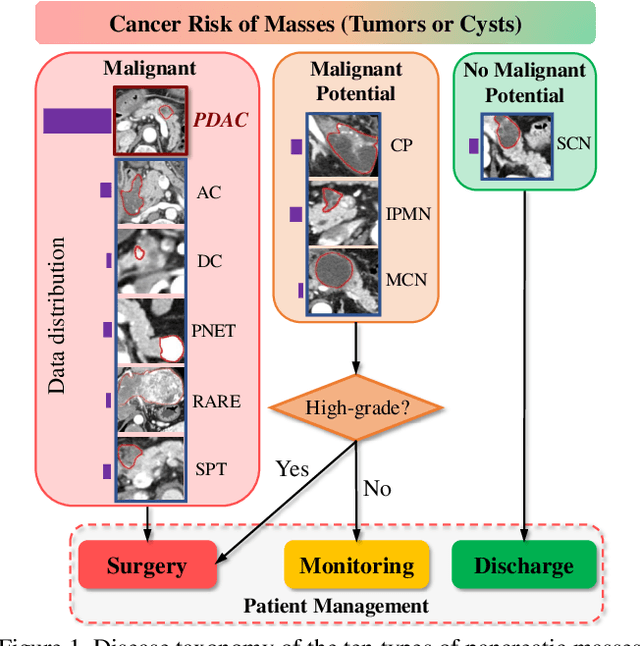
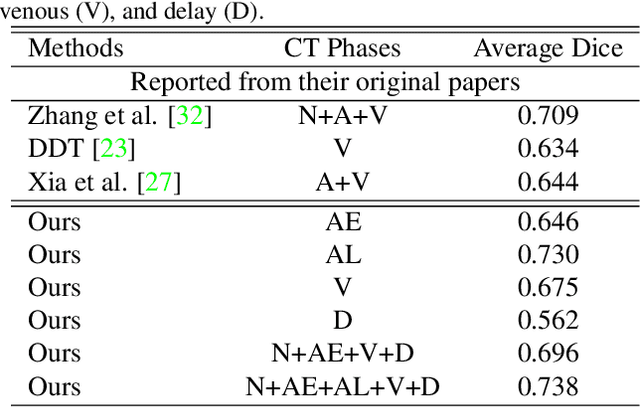
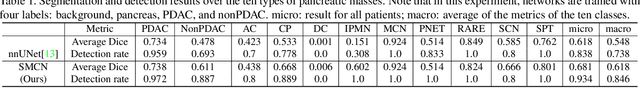
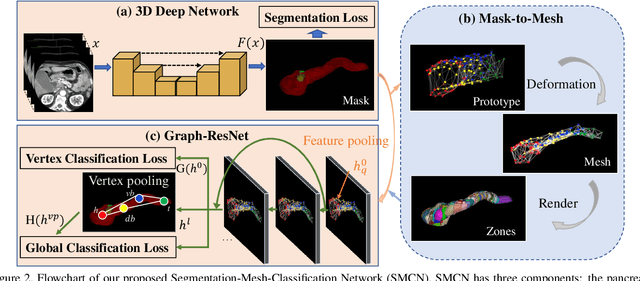

Abstract:The pancreatic disease taxonomy includes ten types of masses (tumors or cysts)[20,8]. Previous work focuses on developing segmentation or classification methods only for certain mass types. Differential diagnosis of all mass types is clinically highly desirable [20] but has not been investigated using an automated image understanding approach. We exploit the feasibility to distinguish pancreatic ductal adenocarcinoma (PDAC) from the nine other nonPDAC masses using multi-phase CT imaging. Both image appearance and the 3D organ-mass geometry relationship are critical. We propose a holistic segmentation-mesh-classification network (SMCN) to provide patient-level diagnosis, by fully utilizing the geometry and location information, which is accomplished by combining the anatomical structure and the semantic detection-by-segmentation network. SMCN learns the pancreas and mass segmentation task and builds an anatomical correspondence-aware organ mesh model by progressively deforming a pancreas prototype on the raw segmentation mask (i.e., mask-to-mesh). A new graph-based residual convolutional network (Graph-ResNet), whose nodes fuse the information of the mesh model and feature vectors extracted from the segmentation network, is developed to produce the patient-level differential classification results. Extensive experiments on 661 patients' CT scans (five phases per patient) show that SMCN can improve the mass segmentation and detection accuracy compared to the strong baseline method nnUNet (e.g., for nonPDAC, Dice: 0.611 vs. 0.478; detection rate: 89% vs. 70%), achieve similar sensitivity and specificity in differentiating PDAC and nonPDAC as expert radiologists (i.e., 94% and 90%), and obtain results comparable to a multimodality test [20] that combines clinical, imaging, and molecular testing for clinical management of patients.
Learning from Multiple Datasets with Heterogeneous and Partial Labels for Universal Lesion Detection in CT
Sep 05, 2020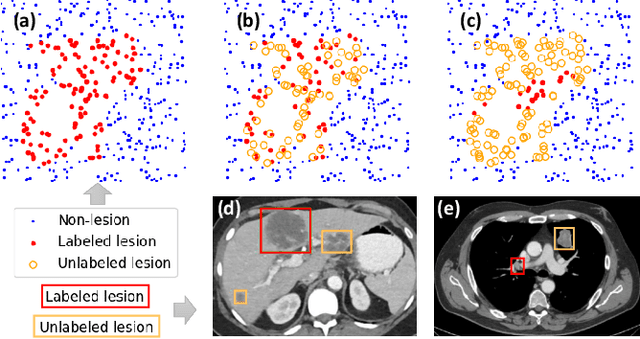
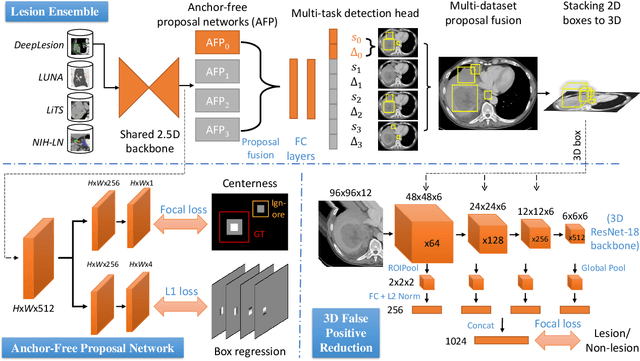
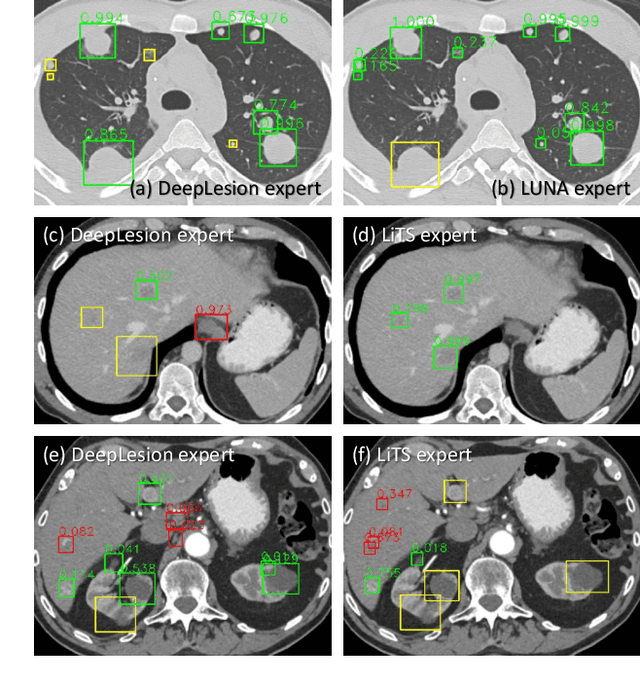
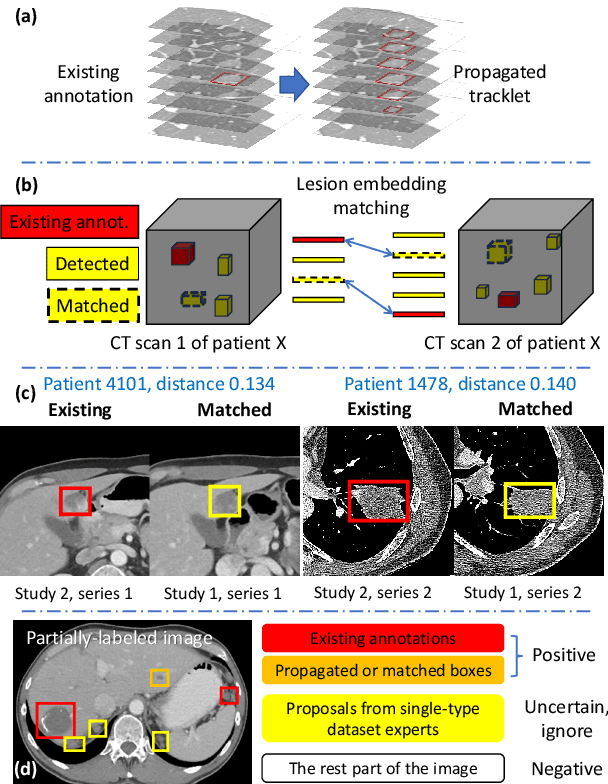
Abstract:Large-scale datasets with high-quality labels are desired for training accurate deep learning models. However, due to annotation costs, medical imaging datasets are often either partially-labeled or small. For example, DeepLesion is a large-scale CT image dataset with lesions of various types, but it also has many unlabeled lesions (missing annotations). When training a lesion detector on a partially-labeled dataset, the missing annotations will generate incorrect negative signals and degrade performance. Besides DeepLesion, there are several small single-type datasets, such as LUNA for lung nodules and LiTS for liver tumors. Such datasets have heterogeneous label scopes, i.e., different lesion types are labeled in different datasets with other types ignored. In this work, we aim to tackle the problem of heterogeneous and partial labels, and develop a universal lesion detection algorithm to detect a comprehensive variety of lesions. First, we build a simple yet effective lesion detection framework named Lesion ENSemble (LENS). LENS can efficiently learn from multiple heterogeneous lesion datasets in a multi-task fashion and leverage their synergy by feature sharing and proposal fusion. Next, we propose strategies to mine missing annotations from partially-labeled datasets by exploiting clinical prior knowledge and cross-dataset knowledge transfer. Finally, we train our framework on four public lesion datasets and evaluate it on 800 manually-labeled sub-volumes in DeepLesion. On this challenging task, our method brings a relative improvement of 49% compared to the current state-of-the-art approach.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge