Leo Anthony Celi
MIT
Towards long-tailed, multi-label disease classification from chest X-ray: Overview of the CXR-LT challenge
Oct 24, 2023



Abstract:Many real-world image recognition problems, such as diagnostic medical imaging exams, are "long-tailed" $\unicode{x2013}$ there are a few common findings followed by many more relatively rare conditions. In chest radiography, diagnosis is both a long-tailed and multi-label problem, as patients often present with multiple findings simultaneously. While researchers have begun to study the problem of long-tailed learning in medical image recognition, few have studied the interaction of label imbalance and label co-occurrence posed by long-tailed, multi-label disease classification. To engage with the research community on this emerging topic, we conducted an open challenge, CXR-LT, on long-tailed, multi-label thorax disease classification from chest X-rays (CXRs). We publicly release a large-scale benchmark dataset of over 350,000 CXRs, each labeled with at least one of 26 clinical findings following a long-tailed distribution. We synthesize common themes of top-performing solutions, providing practical recommendations for long-tailed, multi-label medical image classification. Finally, we use these insights to propose a path forward involving vision-language foundation models for few- and zero-shot disease classification.
Causal thinking for decision making on Electronic Health Records: why and how
Aug 03, 2023



Abstract:Accurate predictions, as with machine learning, may not suffice to provide optimal healthcare for every patient. Indeed, prediction can be driven by shortcuts in the data, such as racial biases. Causal thinking is needed for data-driven decisions. Here, we give an introduction to the key elements, focusing on routinely-collected data, electronic health records (EHRs) and claims data. Using such data to assess the value of an intervention requires care: temporal dependencies and existing practices easily confound the causal effect. We present a step-by-step framework to help build valid decision making from real-life patient records by emulating a randomized trial before individualizing decisions, eg with machine learning. Our framework highlights the most important pitfalls and considerations in analysing EHRs or claims data to draw causal conclusions. We illustrate the various choices in studying the effect of albumin on sepsis mortality in the Medical Information Mart for Intensive Care database (MIMIC-IV). We study the impact of various choices at every step, from feature extraction to causal-estimator selection. In a tutorial spirit, the code and the data are openly available.
Evaluating the Impact of Social Determinants on Health Prediction
May 22, 2023



Abstract:Social determinants of health (SDOH) -- the conditions in which people live, grow, and age -- play a crucial role in a person's health and well-being. There is a large, compelling body of evidence in population health studies showing that a wide range of SDOH is strongly correlated with health outcomes. Yet, a majority of the risk prediction models based on electronic health records (EHR) do not incorporate a comprehensive set of SDOH features as they are often noisy or simply unavailable. Our work links a publicly available EHR database, MIMIC-IV, to well-documented SDOH features. We investigate the impact of such features on common EHR prediction tasks across different patient populations. We find that community-level SDOH features do not improve model performance for a general patient population, but can improve data-limited model fairness for specific subpopulations. We also demonstrate that SDOH features are vital for conducting thorough audits of algorithmic biases beyond protective attributes. We hope the new integrated EHR-SDOH database will enable studies on the relationship between community health and individual outcomes and provide new benchmarks to study algorithmic biases beyond race, gender, and age.
Towards clinical AI fairness: A translational perspective
Apr 26, 2023
Abstract:Artificial intelligence (AI) has demonstrated the ability to extract insights from data, but the issue of fairness remains a concern in high-stakes fields such as healthcare. Despite extensive discussion and efforts in algorithm development, AI fairness and clinical concerns have not been adequately addressed. In this paper, we discuss the misalignment between technical and clinical perspectives of AI fairness, highlight the barriers to AI fairness' translation to healthcare, advocate multidisciplinary collaboration to bridge the knowledge gap, and provide possible solutions to address the clinical concerns pertaining to AI fairness.
Early Diagnosis of Chronic Obstructive Pulmonary Disease from Chest X-Rays using Transfer Learning and Fusion Strategies
Nov 13, 2022Abstract:Chronic obstructive pulmonary disease (COPD) is one of the most common chronic illnesses in the world and the third leading cause of mortality worldwide. It is often underdiagnosed or not diagnosed until later in the disease course. Spirometry tests are the gold standard for diagnosing COPD but can be difficult to obtain, especially in resource-poor countries. Chest X-rays (CXRs), however, are readily available and may serve as a screening tool to identify patients with COPD who should undergo further testing. Currently, no research applies deep learning (DL) algorithms that use large multi-site and multi-modal data to detect COPD patients and evaluate fairness across demographic groups. We use three CXR datasets in our study, CheXpert to pre-train models, MIMIC-CXR to develop, and Emory-CXR to validate our models. The CXRs from patients in the early stage of COPD and not on mechanical ventilation are selected for model training and validation. We visualize the Grad-CAM heatmaps of the true positive cases on the base model for both MIMIC-CXR and Emory-CXR test datasets. We further propose two fusion schemes, (1) model-level fusion, including bagging and stacking methods using MIMIC-CXR, and (2) data-level fusion, including multi-site data using MIMIC-CXR and Emory-CXR, and multi-modal using MIMIC-CXRs and MIMIC-IV EHR, to improve the overall model performance. Fairness analysis is performed to evaluate if the fusion schemes have a discrepancy in the performance among different demographic groups. The results demonstrate that DL models can detect COPD using CXRs, which can facilitate early screening, especially in low-resource regions where CXRs are more accessible than spirometry. The multi-site data fusion scheme could improve the model generalizability on the Emory-CXR test data. Further studies on using CXR or other modalities to predict COPD ought to be in future work.
Learning to Ask Like a Physician
Jun 06, 2022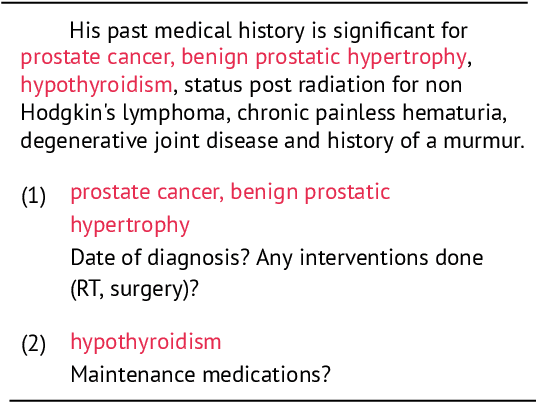
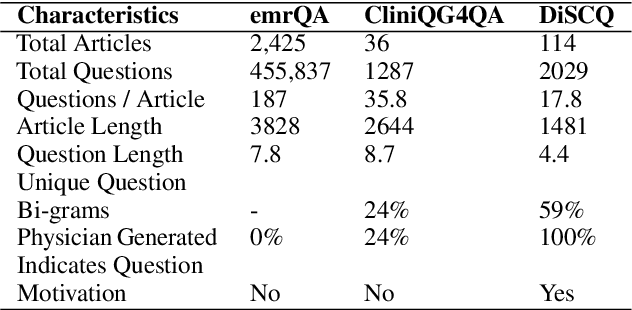
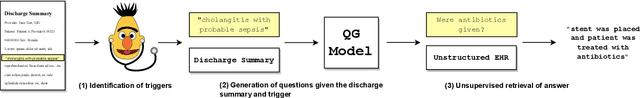
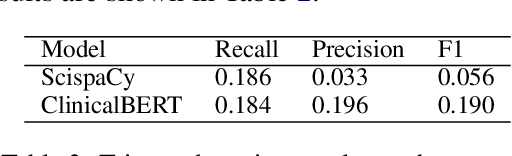
Abstract:Existing question answering (QA) datasets derived from electronic health records (EHR) are artificially generated and consequently fail to capture realistic physician information needs. We present Discharge Summary Clinical Questions (DiSCQ), a newly curated question dataset composed of 2,000+ questions paired with the snippets of text (triggers) that prompted each question. The questions are generated by medical experts from 100+ MIMIC-III discharge summaries. We analyze this dataset to characterize the types of information sought by medical experts. We also train baseline models for trigger detection and question generation (QG), paired with unsupervised answer retrieval over EHRs. Our baseline model is able to generate high quality questions in over 62% of cases when prompted with human selected triggers. We release this dataset (and all code to reproduce baseline model results) to facilitate further research into realistic clinical QA and QG: https://github.com/elehman16/discq.
Write It Like You See It: Detectable Differences in Clinical Notes By Race Lead To Differential Model Recommendations
May 08, 2022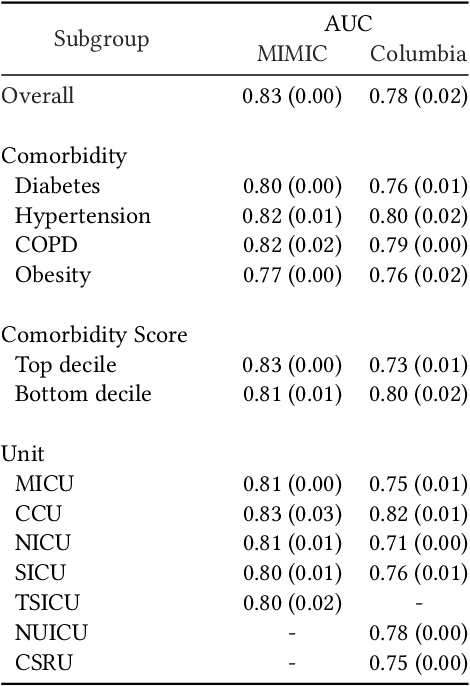
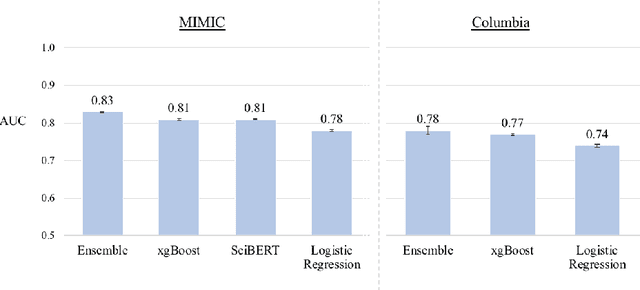


Abstract:Clinical notes are becoming an increasingly important data source for machine learning (ML) applications in healthcare. Prior research has shown that deploying ML models can perpetuate existing biases against racial minorities, as bias can be implicitly embedded in data. In this study, we investigate the level of implicit race information available to ML models and human experts and the implications of model-detectable differences in clinical notes. Our work makes three key contributions. First, we find that models can identify patient self-reported race from clinical notes even when the notes are stripped of explicit indicators of race. Second, we determine that human experts are not able to accurately predict patient race from the same redacted clinical notes. Finally, we demonstrate the potential harm of this implicit information in a simulation study, and show that models trained on these race-redacted clinical notes can still perpetuate existing biases in clinical treatment decisions.
Identification of Subgroups With Similar Benefits in Off-Policy Policy Evaluation
Nov 28, 2021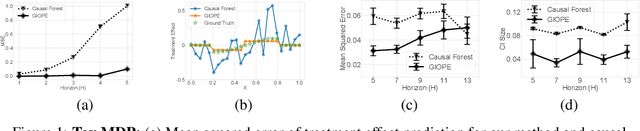
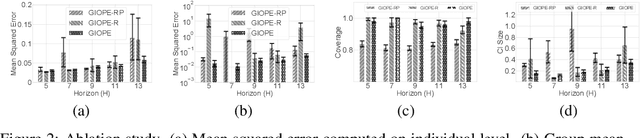
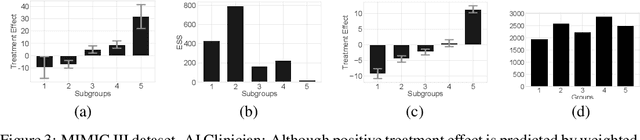
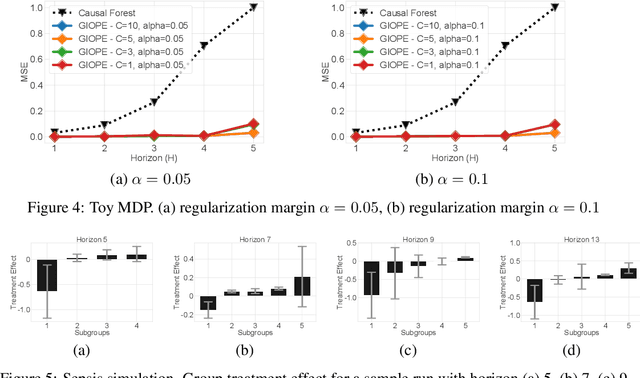
Abstract:Off-policy policy evaluation methods for sequential decision making can be used to help identify if a proposed decision policy is better than a current baseline policy. However, a new decision policy may be better than a baseline policy for some individuals but not others. This has motivated a push towards personalization and accurate per-state estimates of heterogeneous treatment effects (HTEs). Given the limited data present in many important applications, individual predictions can come at a cost to accuracy and confidence in such predictions. We develop a method to balance the need for personalization with confident predictions by identifying subgroups where it is possible to confidently estimate the expected difference in a new decision policy relative to a baseline. We propose a novel loss function that accounts for uncertainty during the subgroup partitioning phase. In experiments, we show that our method can be used to form accurate predictions of HTEs where other methods struggle.
Developing and validating multi-modal models for mortality prediction in COVID-19 patients: a multi-center retrospective study
Sep 01, 2021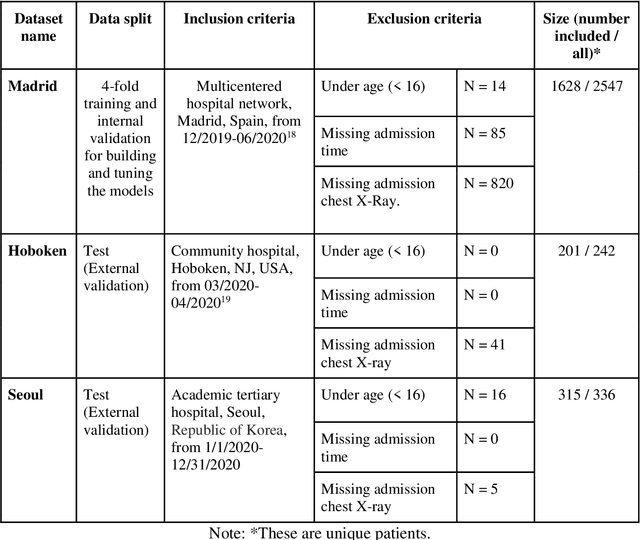
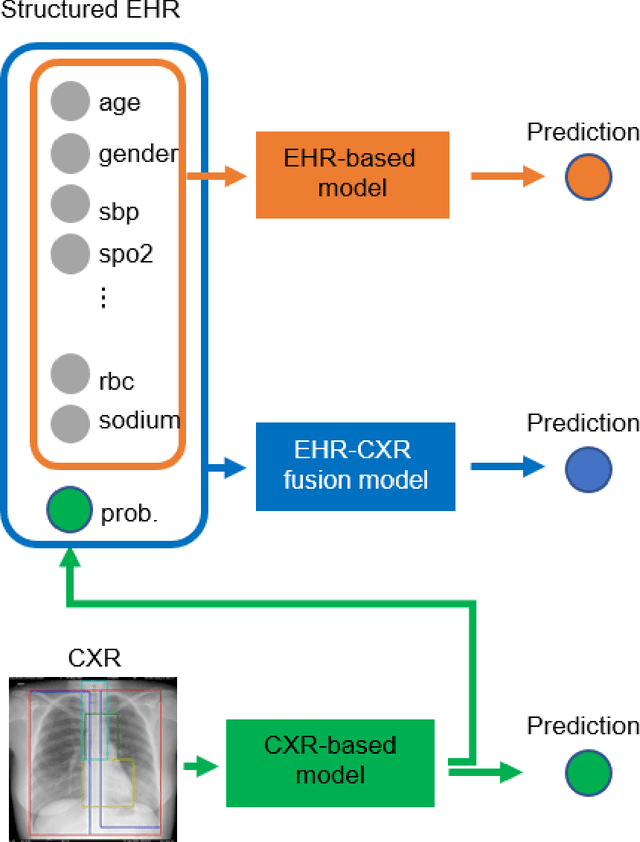
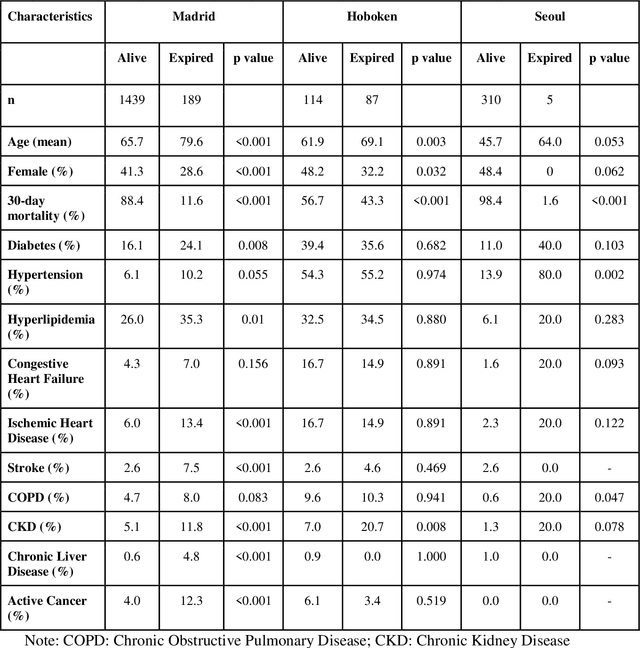
Abstract:The unprecedented global crisis brought about by the COVID-19 pandemic has sparked numerous efforts to create predictive models for the detection and prognostication of SARS-CoV-2 infections with the goal of helping health systems allocate resources. Machine learning models, in particular, hold promise for their ability to leverage patient clinical information and medical images for prediction. However, most of the published COVID-19 prediction models thus far have little clinical utility due to methodological flaws and lack of appropriate validation. In this paper, we describe our methodology to develop and validate multi-modal models for COVID-19 mortality prediction using multi-center patient data. The models for COVID-19 mortality prediction were developed using retrospective data from Madrid, Spain (N=2547) and were externally validated in patient cohorts from a community hospital in New Jersey, USA (N=242) and an academic center in Seoul, Republic of Korea (N=336). The models we developed performed differently across various clinical settings, underscoring the need for a guided strategy when employing machine learning for clinical decision-making. We demonstrated that using features from both the structured electronic health records and chest X-ray imaging data resulted in better 30-day-mortality prediction performance across all three datasets (areas under the receiver operating characteristic curves: 0.85 (95% confidence interval: 0.83-0.87), 0.76 (0.70-0.82), and 0.95 (0.92-0.98)). We discuss the rationale for the decisions made at every step in developing the models and have made our code available to the research community. We employed the best machine learning practices for clinical model development. Our goal is to create a toolkit that would assist investigators and organizations in building multi-modal models for prediction, classification and/or optimization.
Reading Race: AI Recognises Patient's Racial Identity In Medical Images
Jul 21, 2021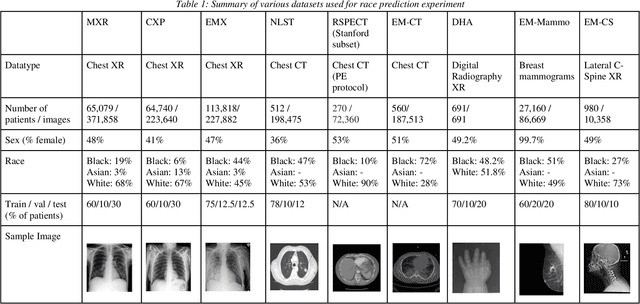
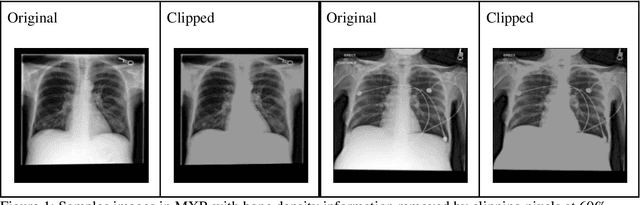
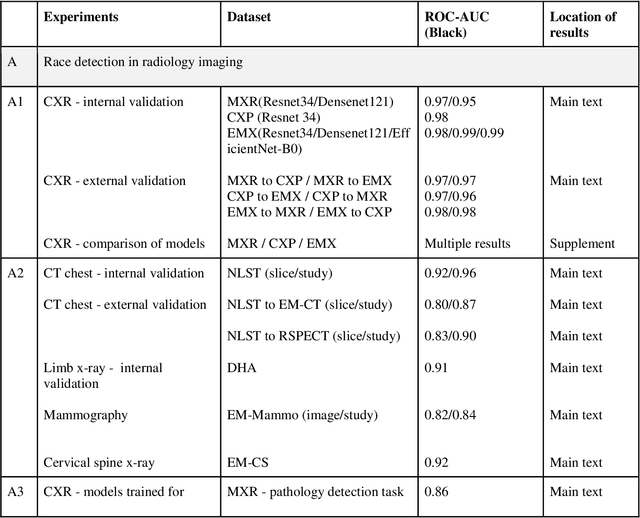
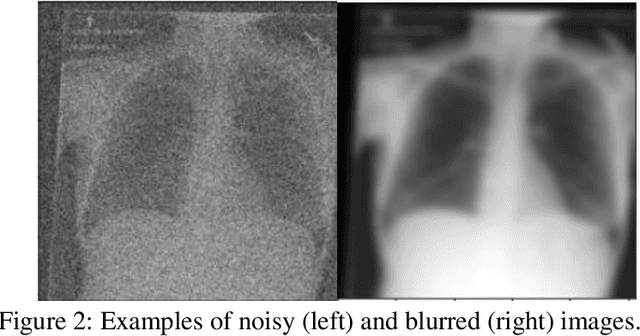
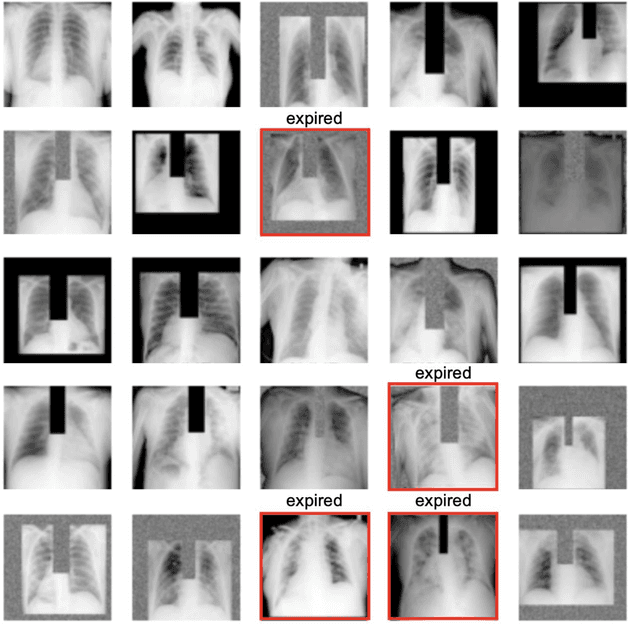
Abstract:Background: In medical imaging, prior studies have demonstrated disparate AI performance by race, yet there is no known correlation for race on medical imaging that would be obvious to the human expert interpreting the images. Methods: Using private and public datasets we evaluate: A) performance quantification of deep learning models to detect race from medical images, including the ability of these models to generalize to external environments and across multiple imaging modalities, B) assessment of possible confounding anatomic and phenotype population features, such as disease distribution and body habitus as predictors of race, and C) investigation into the underlying mechanism by which AI models can recognize race. Findings: Standard deep learning models can be trained to predict race from medical images with high performance across multiple imaging modalities. Our findings hold under external validation conditions, as well as when models are optimized to perform clinically motivated tasks. We demonstrate this detection is not due to trivial proxies or imaging-related surrogate covariates for race, such as underlying disease distribution. Finally, we show that performance persists over all anatomical regions and frequency spectrum of the images suggesting that mitigation efforts will be challenging and demand further study. Interpretation: We emphasize that model ability to predict self-reported race is itself not the issue of importance. However, our findings that AI can trivially predict self-reported race -- even from corrupted, cropped, and noised medical images -- in a setting where clinical experts cannot, creates an enormous risk for all model deployments in medical imaging: if an AI model secretly used its knowledge of self-reported race to misclassify all Black patients, radiologists would not be able to tell using the same data the model has access to.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge