Lena Maier-Hein
German Cancer Research Center, DKFZ Heidelberg, Helmholtz Imaging, Germany, National Center for Tumor Diseases, Faculty of Mathematics and Computer Science, Heidelberg University, Germany, Medical Faculty, Heidelberg University, Germany
Machine learning-based analysis of hyperspectral images for automated sepsis diagnosis
Jun 15, 2021
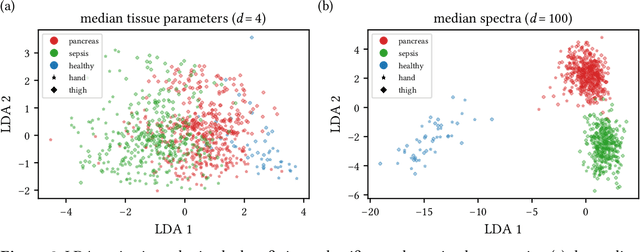
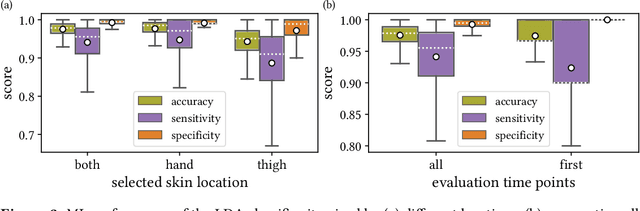

Abstract:Sepsis is a leading cause of mortality and critical illness worldwide. While robust biomarkers for early diagnosis are still missing, recent work indicates that hyperspectral imaging (HSI) has the potential to overcome this bottleneck by monitoring microcirculatory alterations. Automated machine learning-based diagnosis of sepsis based on HSI data, however, has not been explored to date. Given this gap in the literature, we leveraged an existing data set to (1) investigate whether HSI-based automated diagnosis of sepsis is possible and (2) put forth a list of possible confounders relevant for HSI-based tissue classification. While we were able to classify sepsis with an accuracy of over $98\,\%$ using the existing data, our research also revealed several subject-, therapy- and imaging-related confounders that may lead to an overestimation of algorithm performance when not balanced across the patient groups. We conclude that further prospective studies, carefully designed with respect to these confounders, are necessary to confirm the preliminary results obtained in this study.
The Medical Segmentation Decathlon
Jun 10, 2021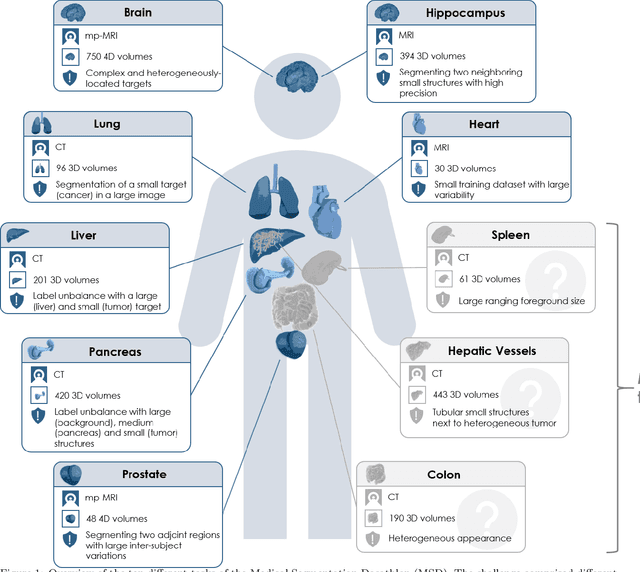
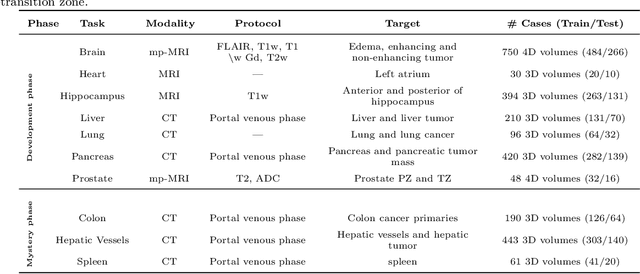
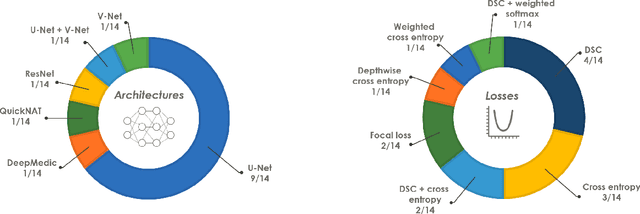
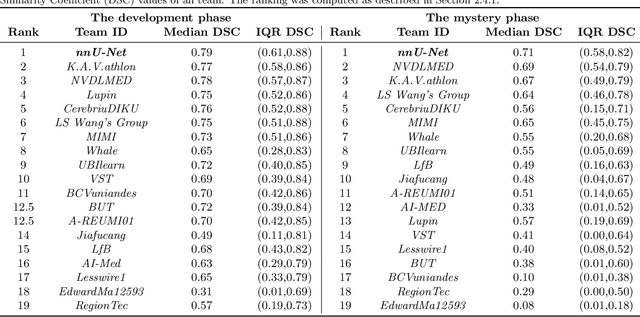
Abstract:International challenges have become the de facto standard for comparative assessment of image analysis algorithms given a specific task. Segmentation is so far the most widely investigated medical image processing task, but the various segmentation challenges have typically been organized in isolation, such that algorithm development was driven by the need to tackle a single specific clinical problem. We hypothesized that a method capable of performing well on multiple tasks will generalize well to a previously unseen task and potentially outperform a custom-designed solution. To investigate the hypothesis, we organized the Medical Segmentation Decathlon (MSD) - a biomedical image analysis challenge, in which algorithms compete in a multitude of both tasks and modalities. The underlying data set was designed to explore the axis of difficulties typically encountered when dealing with medical images, such as small data sets, unbalanced labels, multi-site data and small objects. The MSD challenge confirmed that algorithms with a consistent good performance on a set of tasks preserved their good average performance on a different set of previously unseen tasks. Moreover, by monitoring the MSD winner for two years, we found that this algorithm continued generalizing well to a wide range of other clinical problems, further confirming our hypothesis. Three main conclusions can be drawn from this study: (1) state-of-the-art image segmentation algorithms are mature, accurate, and generalize well when retrained on unseen tasks; (2) consistent algorithmic performance across multiple tasks is a strong surrogate of algorithmic generalizability; (3) the training of accurate AI segmentation models is now commoditized to non AI experts.
Video-rate multispectral imaging in laparoscopic surgery: First-in-human application
May 28, 2021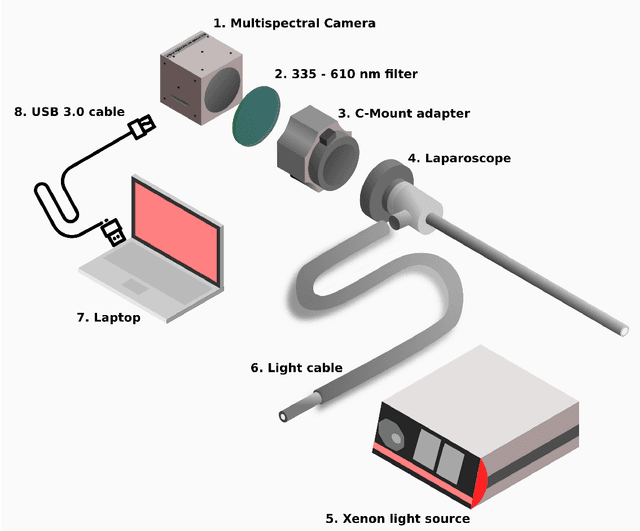
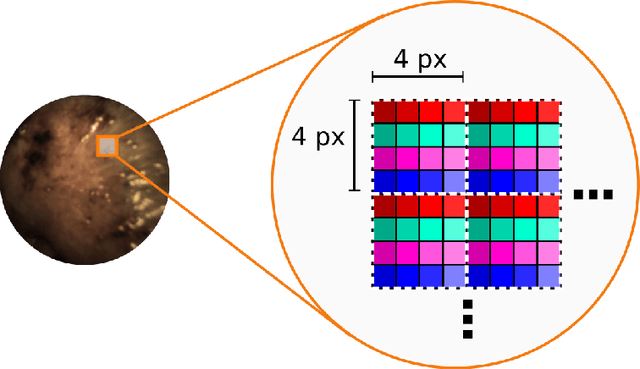
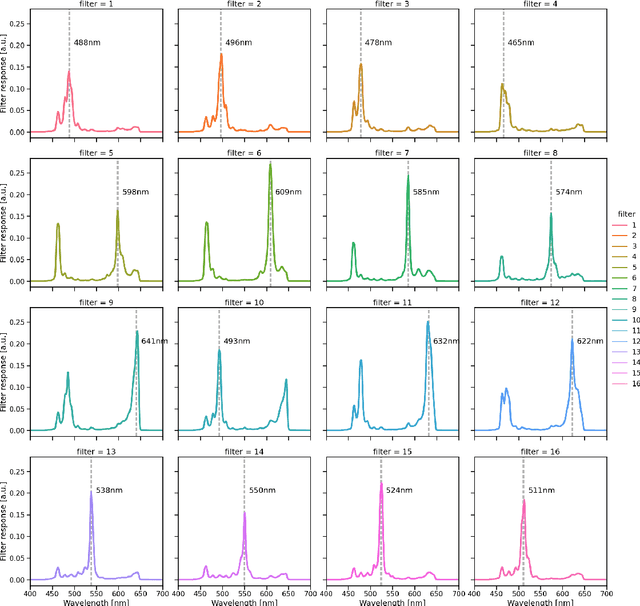
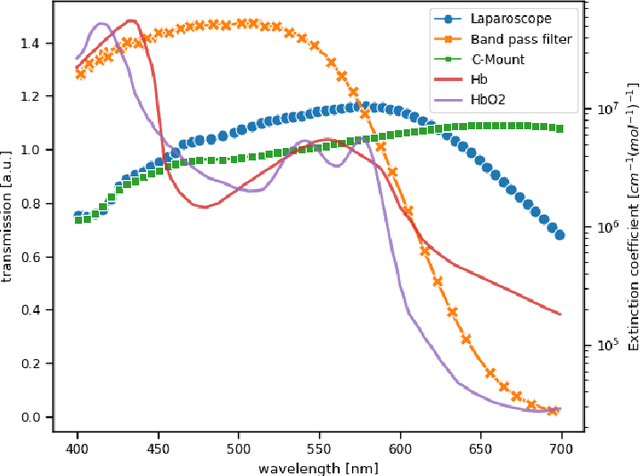
Abstract:Multispectral and hyperspectral imaging (MSI/HSI) can provide clinically relevant information on morphological and functional tissue properties. Application in the operating room (OR), however, has so far been limited by complex hardware setups and slow acquisition times. To overcome these limitations, we propose a novel imaging system for video-rate spectral imaging in the clinical workflow. The system integrates a small snapshot multispectral camera with a standard laparoscope and a clinically commonly used light source, enabling the recording of multispectral images with a spectral dimension of 16 at a frame rate of 25 Hz. An ongoing in patient study shows that multispectral recordings from this system can help detect perfusion changes in partial nephrectomy surgery, thus opening the doors to a wide range of clinical applications.
Semantic segmentation of multispectral photoacoustic images using deep learning
May 20, 2021
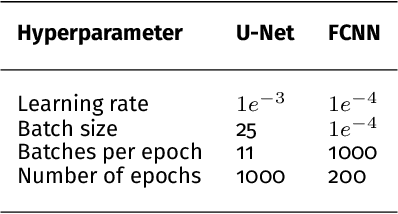


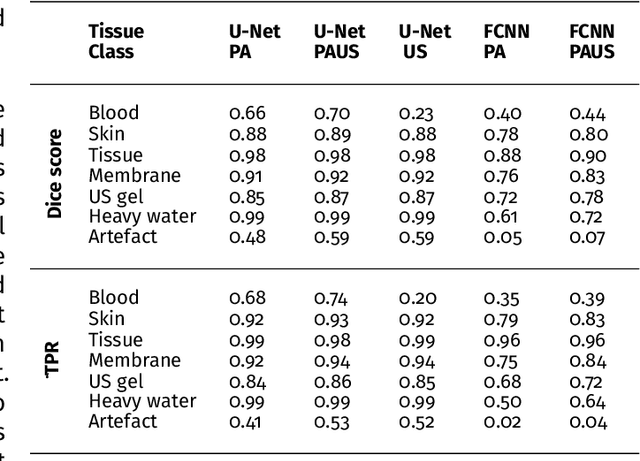
Abstract:Photoacoustic imaging has the potential to revolutionise healthcare due to the valuable information on tissue physiology that is contained in multispectral photoacoustic measurements. Clinical translation of the technology requires conversion of the high-dimensional acquired data into clinically relevant and interpretable information. In this work, we present a deep learning-based approach to semantic segmentation of multispectral photoacoustic images to facilitate the interpretability of recorded images. Manually annotated multispectral photoacoustic imaging data are used as gold standard reference annotations and enable the training of a deep learning-based segmentation algorithm in a supervised manner. Based on a validation study with experimentally acquired data of healthy human volunteers, we show that automatic tissue segmentation can be used to create powerful analyses and visualisations of multispectral photoacoustic images. Due to the intuitive representation of high-dimensional information, such a processing algorithm could be a valuable means to facilitate the clinical translation of photoacoustic imaging.
The Federated Tumor Segmentation (FeTS) Challenge
May 14, 2021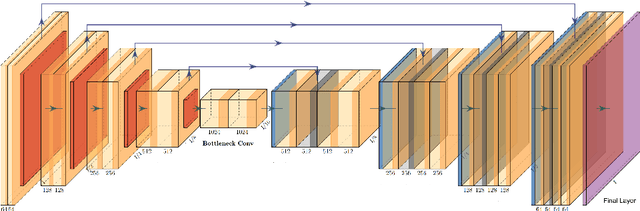
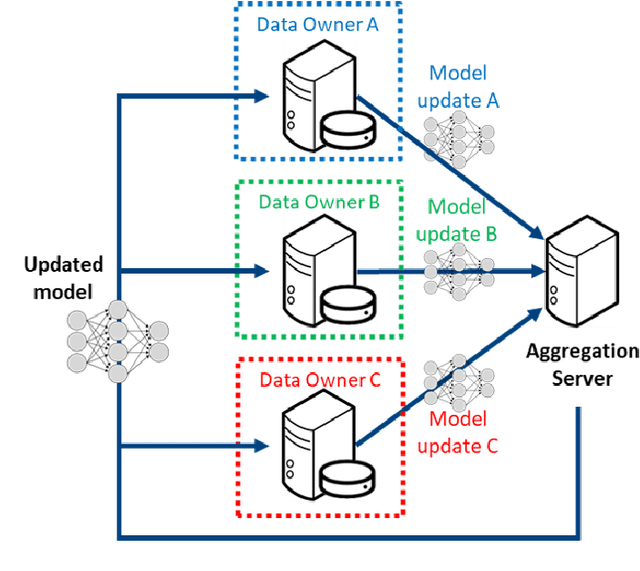
Abstract:This manuscript describes the first challenge on Federated Learning, namely the Federated Tumor Segmentation (FeTS) challenge 2021. International challenges have become the standard for validation of biomedical image analysis methods. However, the actual performance of participating (even the winning) algorithms on "real-world" clinical data often remains unclear, as the data included in challenges are usually acquired in very controlled settings at few institutions. The seemingly obvious solution of just collecting increasingly more data from more institutions in such challenges does not scale well due to privacy and ownership hurdles. Towards alleviating these concerns, we are proposing the FeTS challenge 2021 to cater towards both the development and the evaluation of models for the segmentation of intrinsically heterogeneous (in appearance, shape, and histology) brain tumors, namely gliomas. Specifically, the FeTS 2021 challenge uses clinically acquired, multi-institutional magnetic resonance imaging (MRI) scans from the BraTS 2020 challenge, as well as from various remote independent institutions included in the collaborative network of a real-world federation (https://www.fets.ai/). The goals of the FeTS challenge are directly represented by the two included tasks: 1) the identification of the optimal weight aggregation approach towards the training of a consensus model that has gained knowledge via federated learning from multiple geographically distinct institutions, while their data are always retained within each institution, and 2) the federated evaluation of the generalizability of brain tumor segmentation models "in the wild", i.e. on data from institutional distributions that were not part of the training datasets.
Common Limitations of Image Processing Metrics: A Picture Story
Apr 13, 2021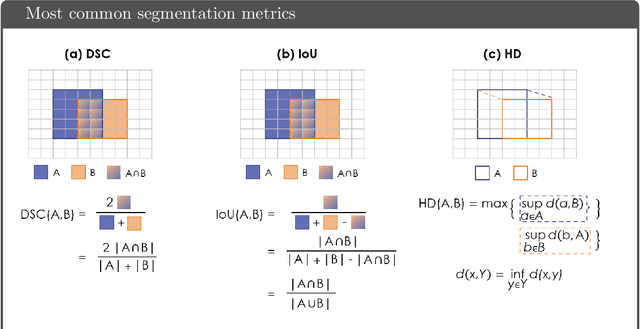
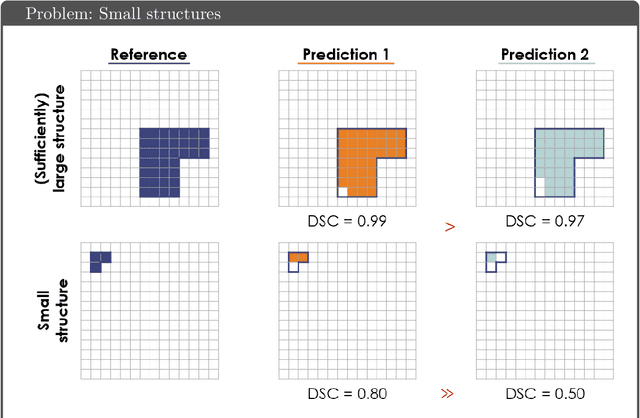
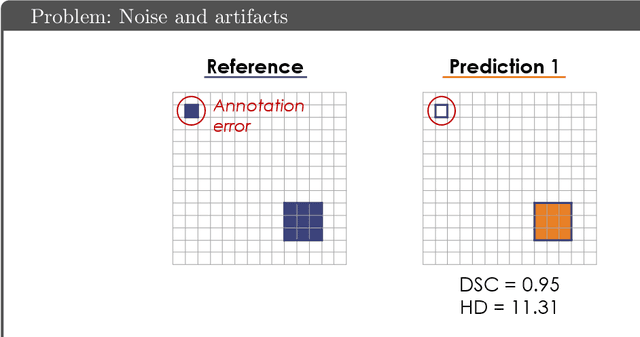
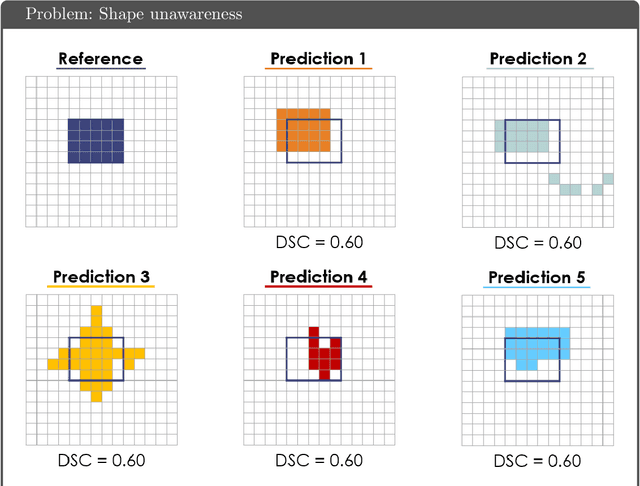
Abstract:While the importance of automatic image analysis is increasing at an enormous pace, recent meta-research revealed major flaws with respect to algorithm validation. Specifically, performance metrics are key for objective, transparent and comparative performance assessment, but relatively little attention has been given to the practical pitfalls when using specific metrics for a given image analysis task. A common mission of several international initiatives is therefore to provide researchers with guidelines and tools to choose the performance metrics in a problem-aware manner. This dynamically updated document has the purpose to illustrate important limitations of performance metrics commonly applied in the field of image analysis. The current version is based on a Delphi process on metrics conducted by an international consortium of image analysis experts.
Data-driven generation of plausible tissue geometries for realistic photoacoustic image synthesis
Mar 29, 2021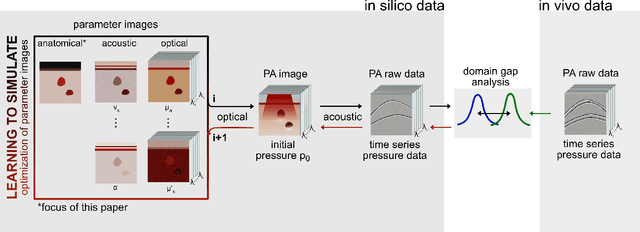
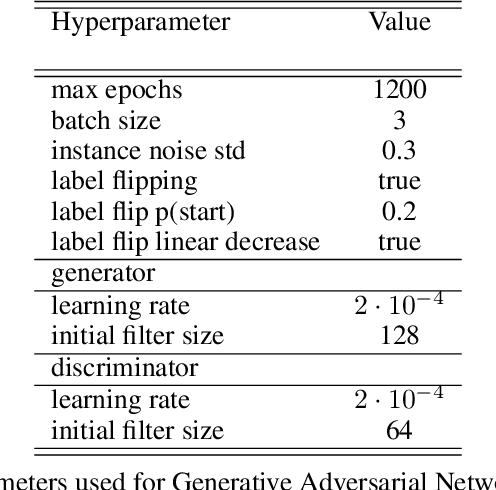
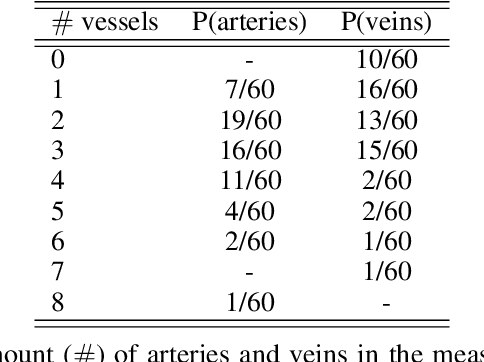
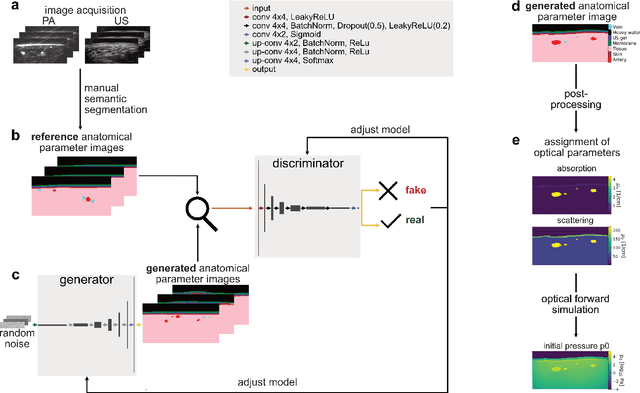

Abstract:Photoacoustic tomography (PAT) has the potential to recover morphological and functional tissue properties such as blood oxygenation with high spatial resolution and in an interventional setting. However, decades of research invested in solving the inverse problem of recovering clinically relevant tissue properties from spectral measurements have failed to produce solutions that can quantify tissue parameters robustly in a clinical setting. Previous attempts to address the limitations of model-based approaches with machine learning were hampered by the absence of labeled reference data needed for supervised algorithm training. While this bottleneck has been tackled by simulating training data, the domain gap between real and simulated images remains a huge unsolved challenge. As a first step to address this bottleneck, we propose a novel approach to PAT data simulation, which we refer to as "learning to simulate". Our approach involves subdividing the challenge of generating plausible simulations into two disjoint problems: (1) Probabilistic generation of realistic tissue morphology, represented by semantic segmentation maps and (2) pixel-wise assignment of corresponding optical and acoustic properties. In the present work, we focus on the first challenge. Specifically, we leverage the concept of Generative Adversarial Networks (GANs) trained on semantically annotated medical imaging data to generate plausible tissue geometries. According to an initial in silico feasibility study our approach is well-suited for contributing to realistic PAT image synthesis and could thus become a fundamental step for deep learning-based quantitative PAT.
Surgical Visual Domain Adaptation: Results from the MICCAI 2020 SurgVisDom Challenge
Feb 26, 2021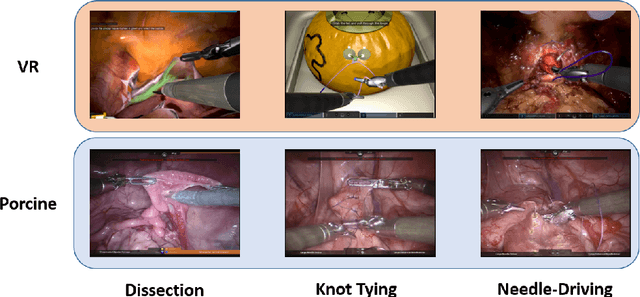
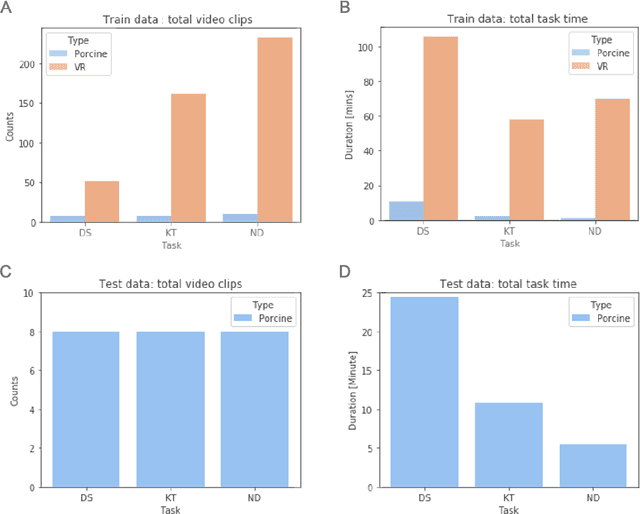
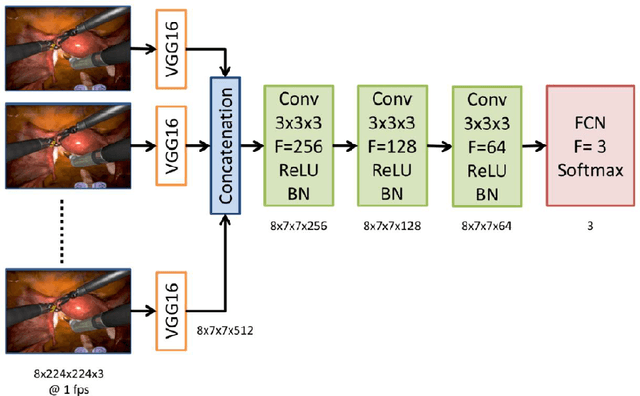
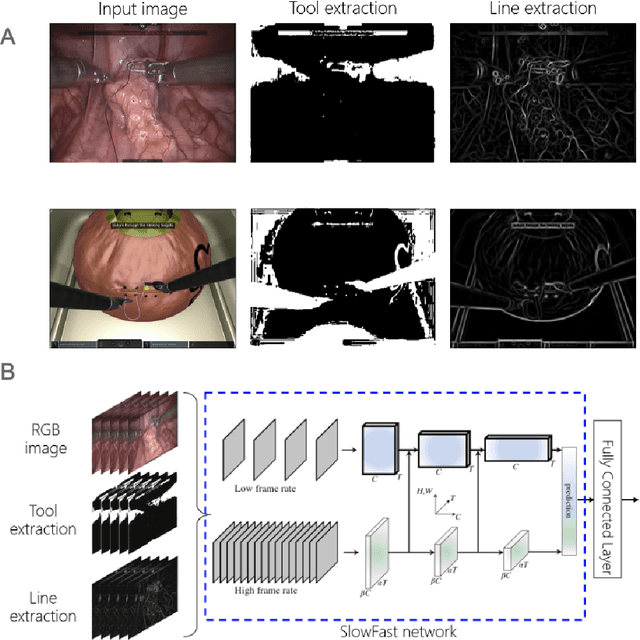
Abstract:Surgical data science is revolutionizing minimally invasive surgery by enabling context-aware applications. However, many challenges exist around surgical data (and health data, more generally) needed to develop context-aware models. This work - presented as part of the Endoscopic Vision (EndoVis) challenge at the Medical Image Computing and Computer Assisted Intervention (MICCAI) 2020 conference - seeks to explore the potential for visual domain adaptation in surgery to overcome data privacy concerns. In particular, we propose to use video from virtual reality (VR) simulations of surgical exercises in robotic-assisted surgery to develop algorithms to recognize tasks in a clinical-like setting. We present the performance of the different approaches to solve visual domain adaptation developed by challenge participants. Our analysis shows that the presented models were unable to learn meaningful motion based features form VR data alone, but did significantly better when small amount of clinical-like data was also made available. Based on these results, we discuss promising methods and further work to address the problem of visual domain adaptation in surgical data science. We also release the challenge dataset publicly at https://www.synapse.org/surgvisdom2020.
Stereo Correspondence and Reconstruction of Endoscopic Data Challenge
Jan 28, 2021
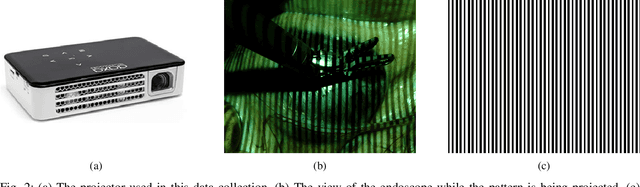

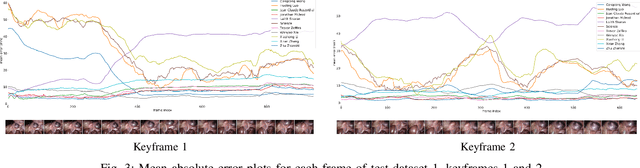
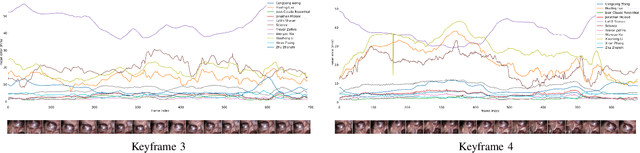
Abstract:The stereo correspondence and reconstruction of endoscopic data sub-challenge was organized during the Endovis challenge at MICCAI 2019 in Shenzhen, China. The task was to perform dense depth estimation using 7 training datasets and 2 test sets of structured light data captured using porcine cadavers. These were provided by a team at Intuitive Surgical. 10 teams participated in the challenge day. This paper contains 3 additional methods which were submitted after the challenge finished as well as a supplemental section from these teams on issues they found with the dataset.
SERV-CT: A disparity dataset from CT for validation of endoscopic 3D reconstruction
Dec 22, 2020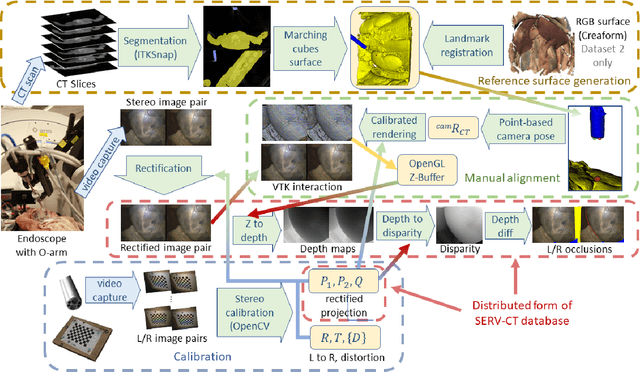
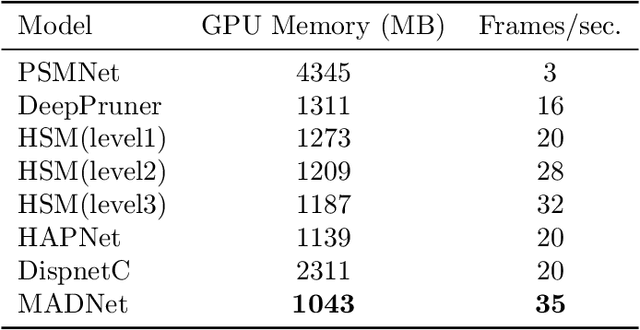
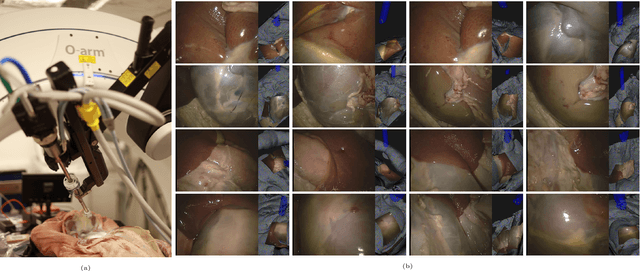
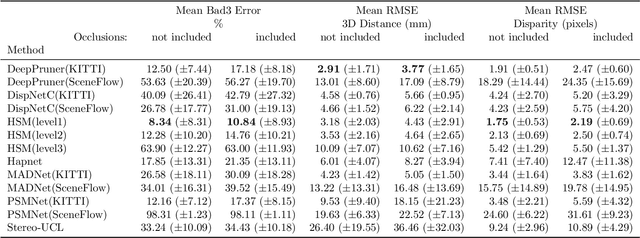
Abstract:In computer vision, reference datasets have been highly successful in promoting algorithmic development in stereo reconstruction. Surgical scenes gives rise to specific problems, including the lack of clear corner features, highly specular surfaces and the presence of blood and smoke. Publicly available datasets have been produced using CT and either phantom images or biological tissue samples covering a relatively small region of the endoscope field-of-view. We present a stereo-endoscopic reconstruction validation dataset based on CT (SERV-CT). Two {\it ex vivo} small porcine full torso cadavers were placed within the view of the endoscope with both the endoscope and target anatomy visible in the CT scan. Orientation of the endoscope was manually aligned to the stereoscopic view. Reference disparities and occlusions were calculated for 8 stereo pairs from each sample. For the second sample an RGB surface was acquired to aid alignment of smooth, featureless surfaces. Repeated manual alignments showed an RMS disparity accuracy of ~2 pixels and a depth accuracy of ~2mm. The reference dataset includes endoscope image pairs with corresponding calibration, disparities, depths and occlusions covering the majority of the endoscopic image and a range of tissue types. Smooth specular surfaces and images with significant variation of depth are included. We assessed the performance of various stereo algorithms from online available repositories. There is a significant variation between algorithms, highlighting some of the challenges of surgical endoscopic images. The SERV-CT dataset provides an easy to use stereoscopic validation for surgical applications with smooth reference disparities and depths with coverage over the majority of the endoscopic images. This complements existing resources well and we hope will aid the development of surgical endoscopic anatomical reconstruction algorithms.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge