Junzhou Huang
Don't Overlook the Support Set: Towards Improving Generalization in Meta-learning
Jul 26, 2020
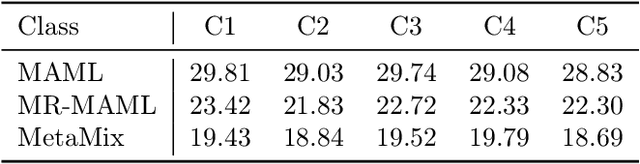
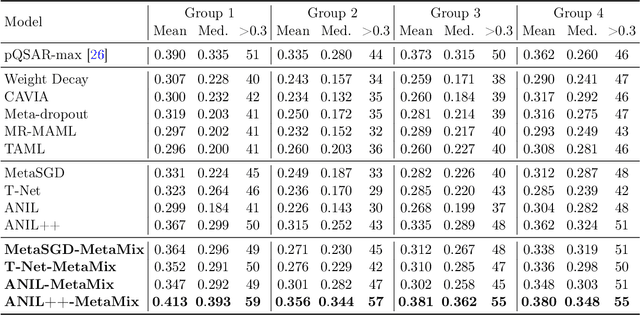
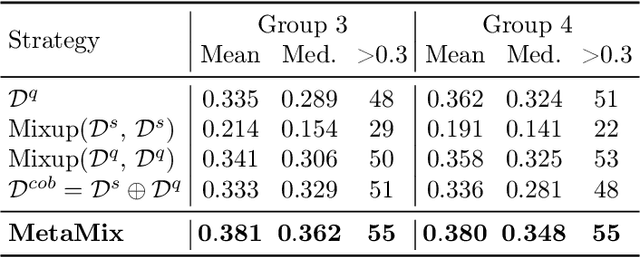
Abstract:Meta-learning has proven to be a powerful paradigm for transferring the knowledge from previously tasks to facilitate the learning of a novel task. Current dominant algorithms train a well-generalized model initialization which is adapted to each task via the support set. The crux, obviously, lies in optimizing the generalization capability of the initialization, which is measured by the performance of the adapted model on the query set of each task. Unfortunately, this generalization measure, evidenced by empirical results, pushes the initialization to overfit the query but fail the support set, which significantly impairs the generalization and adaptation to novel tasks. To address this issue, we include the support set when evaluating the generalization to produce a new meta-training strategy, MetaMix, that linearly combines the input and hidden representations of samples from both the support and query sets. Theoretical studies on classification and regression tasks show how MetaMix can improve the generalization of meta-learning. More remarkably, MetaMix obtains state-of-the-art results by a large margin across many datasets and remains compatible with existing meta-learning algorithms.
Inverse Graph Identification: Can We Identify Node Labels Given Graph Labels?
Jul 12, 2020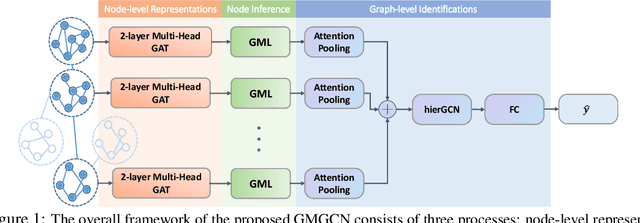
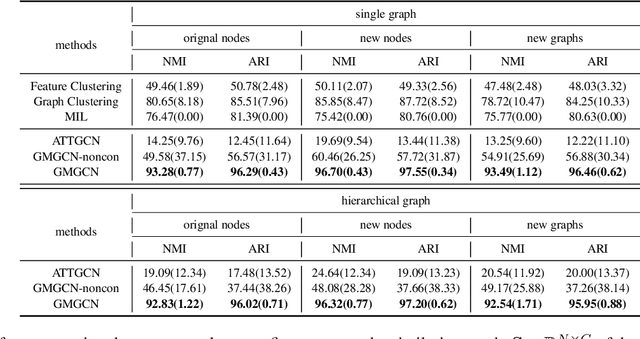
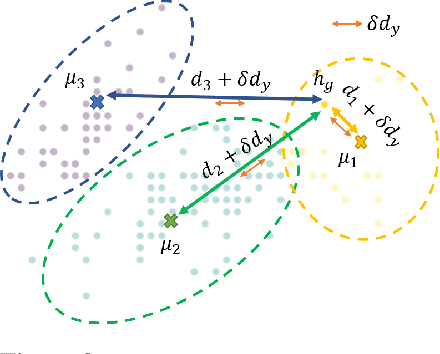
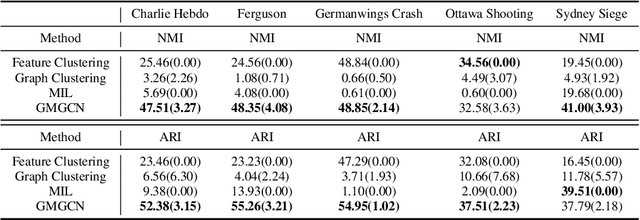
Abstract:Graph Identification (GI) has long been researched in graph learning and is essential in certain applications (e.g. social community detection). Specifically, GI requires to predict the label/score of a target graph given its collection of node features and edge connections. While this task is common, more complex cases arise in practice---we are supposed to do the inverse thing by, for example, grouping similar users in a social network given the labels of different communities. This triggers an interesting thought: can we identify nodes given the labels of the graphs they belong to? Therefore, this paper defines a novel problem dubbed Inverse Graph Identification (IGI), as opposed to GI. Upon a formal discussion of the variants of IGI, we choose a particular case study of node clustering by making use of the graph labels and node features, with an assistance of a hierarchical graph that further characterizes the connections between different graphs. To address this task, we propose Gaussian Mixture Graph Convolutional Network (GMGCN), a simple yet effective method that makes the node-level message passing process using Graph Attention Network (GAT) under the protocol of GI and then infers the category of each node via a Gaussian Mixture Layer (GML). The training of GMGCN is further boosted by a proposed consensus loss to take advantage of the structure of the hierarchical graph. Extensive experiments are conducted to test the rationality of the formulation of IGI. We verify the superiority of the proposed method compared to other baselines on several benchmarks we have built up. We will release our codes along with the benchmark data to facilitate more research attention to the IGI problem.
Breaking the Curse of Space Explosion: Towards Efficient NAS with Curriculum Search
Jul 07, 2020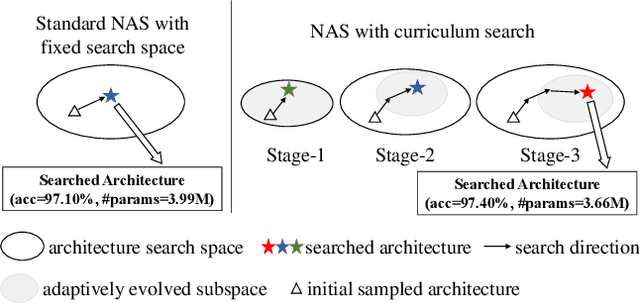
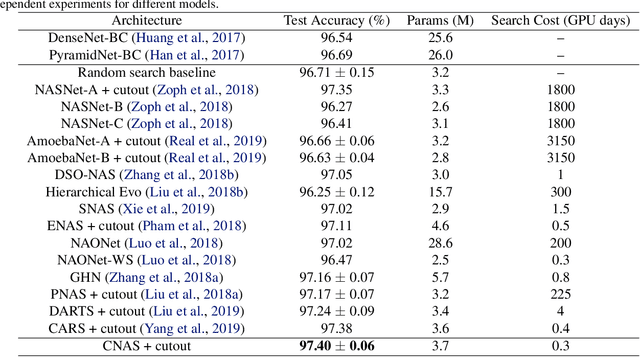
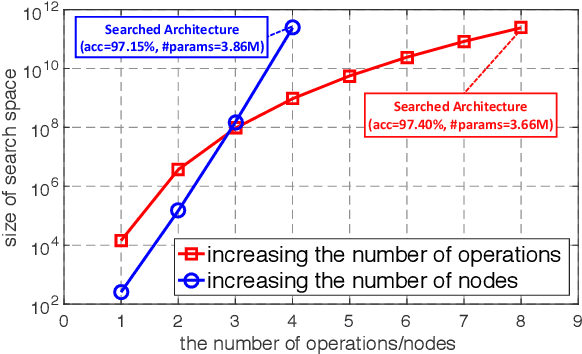
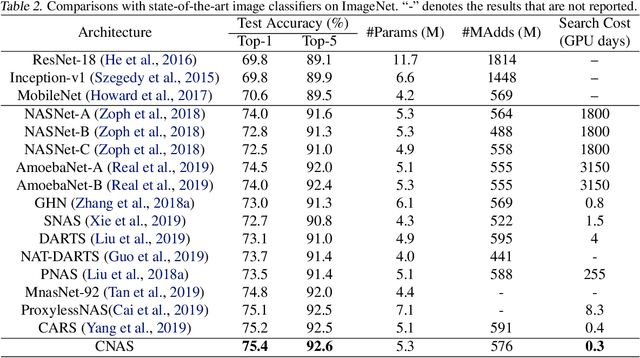
Abstract:Neural architecture search (NAS) has become an important approach to automatically find effective architectures. To cover all possible good architectures, we need to search in an extremely large search space with billions of candidate architectures. More critically, given a large search space, we may face a very challenging issue of space explosion. However, due to the limitation of computational resources, we can only sample a very small proportion of the architectures, which provides insufficient information for the training. As a result, existing methods may often produce suboptimal architectures. To alleviate this issue, we propose a curriculum search method that starts from a small search space and gradually incorporates the learned knowledge to guide the search in a large space. With the proposed search strategy, our Curriculum Neural Architecture Search (CNAS) method significantly improves the search efficiency and finds better architectures than existing NAS methods. Extensive experiments on CIFAR-10 and ImageNet demonstrate the effectiveness of the proposed method.
Collaborative Unsupervised Domain Adaptation for Medical Image Diagnosis
Jul 05, 2020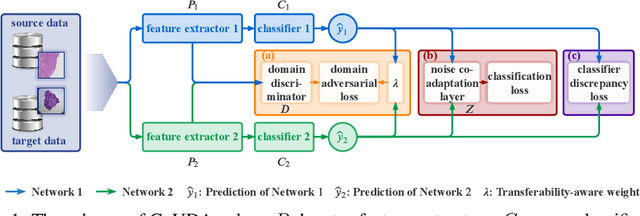
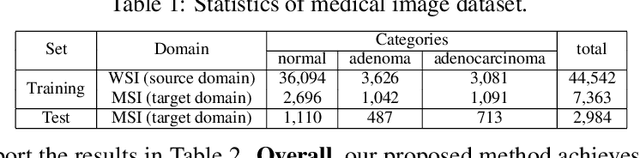
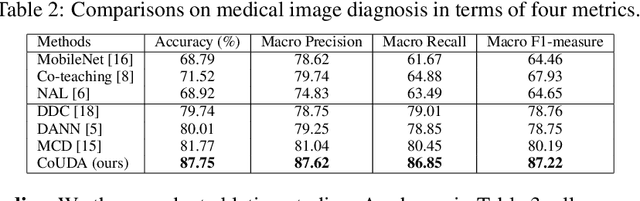
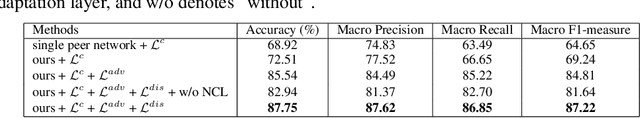
Abstract:Deep learning based medical image diagnosis has shown great potential in clinical medicine. However, it often suffers two major difficulties in real-world applications: 1) only limited labels are available for model training, due to expensive annotation costs over medical images; 2) labeled images may contain considerable label noise (e.g., mislabeling labels) due to diagnostic difficulties of diseases. To address these, we seek to exploit rich labeled data from relevant domains to help the learning in the target task via {Unsupervised Domain Adaptation} (UDA). Unlike most UDA methods that rely on clean labeled data or assume samples are equally transferable, we innovatively propose a Collaborative Unsupervised Domain Adaptation algorithm, which conducts transferability-aware adaptation and conquers label noise in a collaborative way. We theoretically analyze the generalization performance of the proposed method, and also empirically evaluate it on both medical and general images. Promising experimental results demonstrate the superiority and generalization of the proposed method.
GROVER: Self-supervised Message Passing Transformer on Large-scale Molecular Data
Jun 18, 2020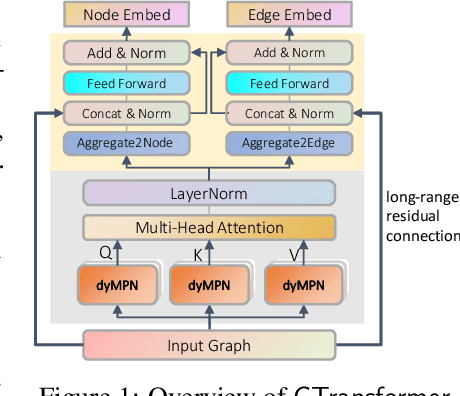
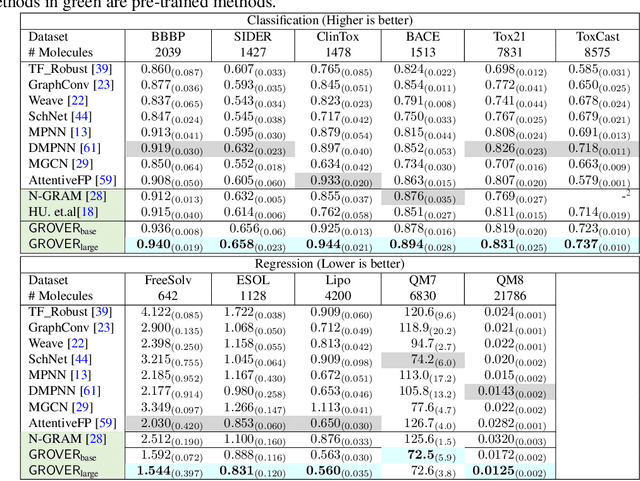
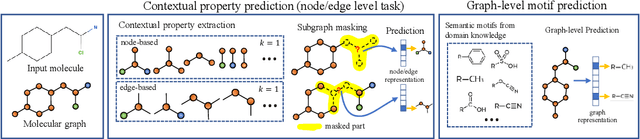
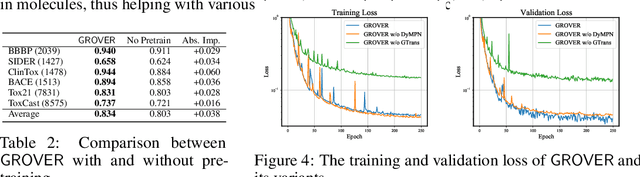
Abstract:How to obtain informative representations of molecules is a crucial prerequisite in AI-driven drug design and discovery. Recent researches abstract molecules as graphs and employ Graph Neural Networks (GNNs) for task-specific and data-driven molecular representation learning. Nevertheless, two "dark clouds" impede the usage of GNNs in real scenarios: (1) insufficient labeled molecules for supervised training; (2) poor generalization capabilities to new-synthesized molecules. To address them both, we propose a novel molecular representation framework, GROVER, which stands for Graph Representation frOm self-superVised mEssage passing tRansformer. With carefully designed self-supervised tasks in node, edge and graph-level, GROVER can learn rich structural and semantic information of molecules from enormous unlabelled molecular data. Rather, to encode such complex information, GROVER integrates Message Passing Networks with the Transformer-style architecture to deliver a class of more expressive encoders of molecules. The flexibility of GROVER allows it to be trained efficiently on large-scale molecular dataset without requiring any supervision, thus being immunized to the two issues mentioned above. We pre-train GROVER with 100 million parameters on 10 million unlabelled molecules---the biggest GNN and the largest training dataset that we have ever met. We then leverage the pre-trained GROVER to downstream molecular property prediction tasks followed by task-specific fine-tuning, where we observe a huge improvement (more than 6% on average) over current state-of-the-art methods on 11 challenging benchmarks. The insights we gained are that well-designed self-supervision losses and largely-expressive pre-trained models enjoy the significant potential on performance boosting.
Multi-View Graph Neural Networks for Molecular Property Prediction
Jun 12, 2020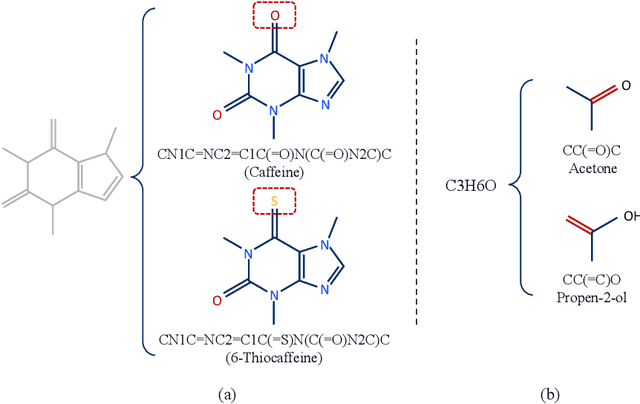
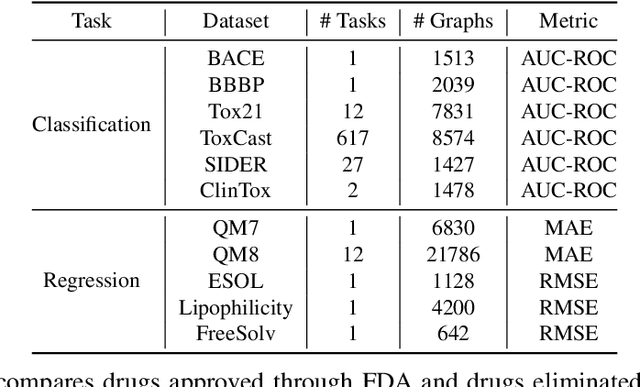
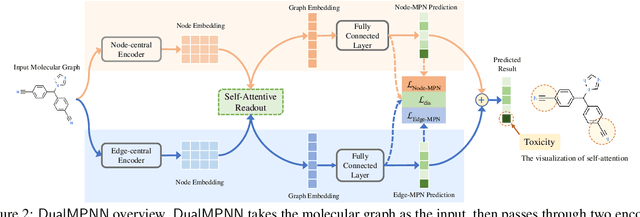
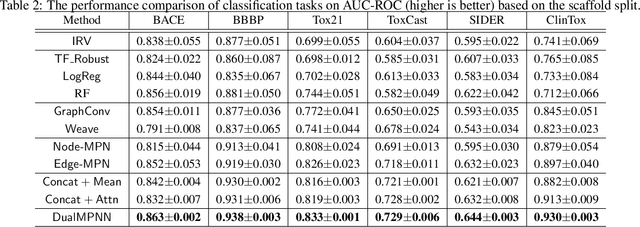
Abstract:The crux of molecular property prediction is to generate meaningful representations of the molecules. One promising route is to exploit the molecular graph structure through Graph Neural Networks (GNNs). It is well known that both atoms and bonds significantly affect the chemical properties of a molecule, so an expressive model shall be able to exploit both node (atom) and edge (bond) information simultaneously. Guided by this observation, we present Multi-View Graph Neural Network (MV-GNN), a multi-view message passing architecture to enable more accurate predictions of molecular properties. In MV-GNN, we introduce a shared self-attentive readout component and disagreement loss to stabilize the training process. This readout component also renders the whole architecture interpretable. We further boost the expressive power of MV-GNN by proposing a cross-dependent message passing scheme that enhances information communication of the two views, which results in the MV-GNN^cross variant. Lastly, we theoretically justify the expressiveness of the two proposed models in terms of distinguishing non-isomorphism graphs. Extensive experiments demonstrate that MV-GNN models achieve remarkably superior performance over the state-of-the-art models on a variety of challenging benchmarks. Meanwhile, visualization results of the node importance are consistent with prior knowledge, which confirms the interpretability power of MV-GNN models.
Co-Heterogeneous and Adaptive Segmentation from Multi-Source and Multi-Phase CT Imaging Data: A Study on Pathological Liver and Lesion Segmentation
Jun 10, 2020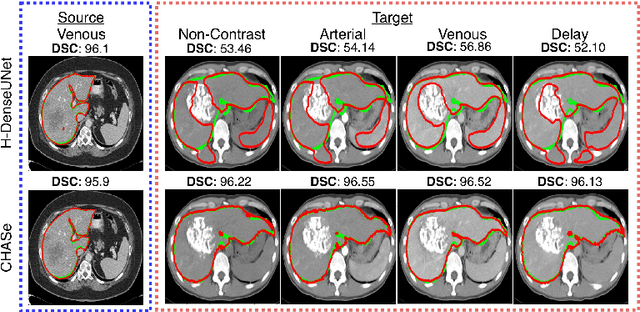
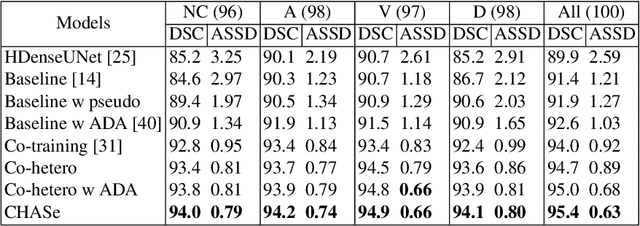
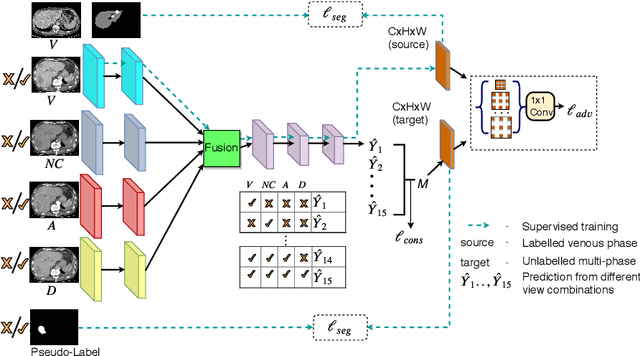
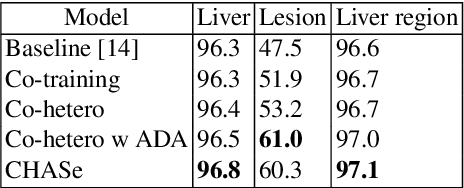
Abstract:In medical imaging, organ/pathology segmentation models trained on current publicly available and fully-annotated datasets usually do not well-represent the heterogeneous modalities, phases, pathologies, and clinical scenarios encountered in real environments. On the other hand, there are tremendous amounts of unlabelled patient imaging scans stored by many modern clinical centers. In this work, we present a novel segmentation strategy, co-heterogenous and adaptive segmentation (CHASe), which only requires a small labeled cohort of single phase imaging data to adapt to any unlabeled cohort of heterogenous multi-phase data with possibly new clinical scenarios and pathologies. To do this, we propose a versatile framework that fuses appearance based semi-supervision, mask based adversarial domain adaptation, and pseudo-labeling. We also introduce co-heterogeneous training, which is a novel integration of co-training and hetero modality learning. We have evaluated CHASe using a clinically comprehensive and challenging dataset of multi-phase computed tomography (CT) imaging studies (1147 patients and 4577 3D volumes). Compared to previous state-of-the-art baselines, CHASe can further improve pathological liver mask Dice-Sorensen coefficients by ranges of $4.2\% \sim 9.4\%$, depending on the phase combinations: e.g., from $84.6\%$ to $94.0\%$ on non-contrast CTs.
COVID-DA: Deep Domain Adaptation from Typical Pneumonia to COVID-19
Apr 30, 2020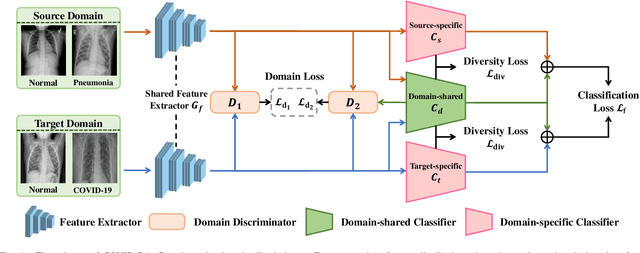
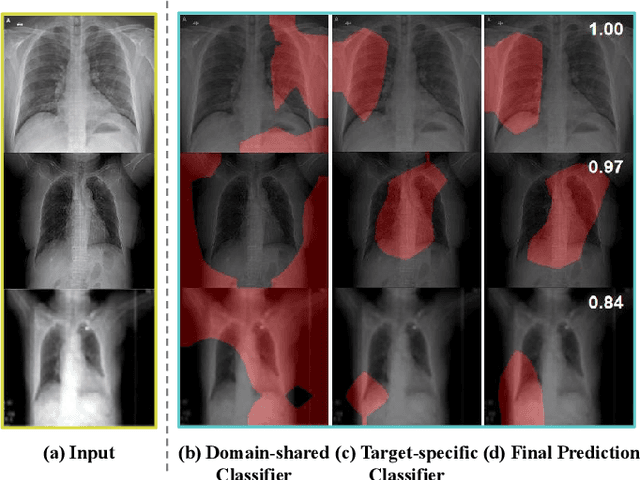
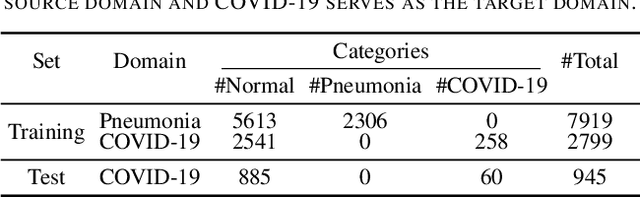
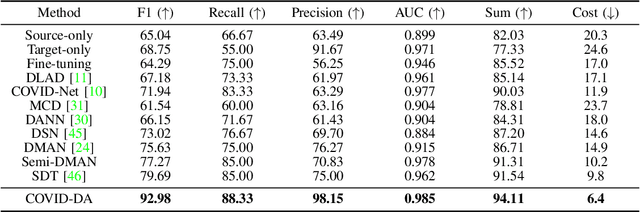
Abstract:The outbreak of novel coronavirus disease 2019 (COVID-19) has already infected millions of people and is still rapidly spreading all over the globe. Most COVID-19 patients suffer from lung infection, so one important diagnostic method is to screen chest radiography images, e.g., X-Ray or CT images. However, such examinations are time-consuming and labor-intensive, leading to limited diagnostic efficiency. To solve this issue, AI-based technologies, such as deep learning, have been used recently as effective computer-aided means to improve diagnostic efficiency. However, one practical and critical difficulty is the limited availability of annotated COVID-19 data, due to the prohibitive annotation costs and urgent work of doctors to fight against the pandemic. This makes the learning of deep diagnosis models very challenging. To address this, motivated by that typical pneumonia has similar characteristics with COVID-19 and many pneumonia datasets are publicly available, we propose to conduct domain knowledge adaptation from typical pneumonia to COVID-19. There are two main challenges: 1) the discrepancy of data distributions between domains; 2) the task difference between the diagnosis of typical pneumonia and COVID-19. To address them, we propose a new deep domain adaptation method for COVID-19 diagnosis, namely COVID-DA. Specifically, we alleviate the domain discrepancy via feature adversarial adaptation and handle the task difference issue via a novel classifier separation scheme. In this way, COVID-DA is able to diagnose COVID-19 effectively with only a small number of COVID-19 annotations. Extensive experiments verify the effectiveness of COVID-DA and its great potential for real-world applications.
Real-Time Semantic Segmentation via Auto Depth, Downsampling Joint Decision and Feature Aggregation
Mar 31, 2020
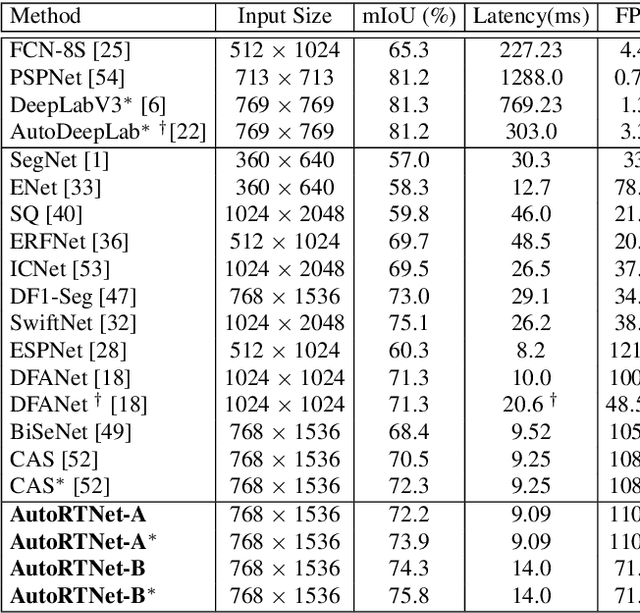

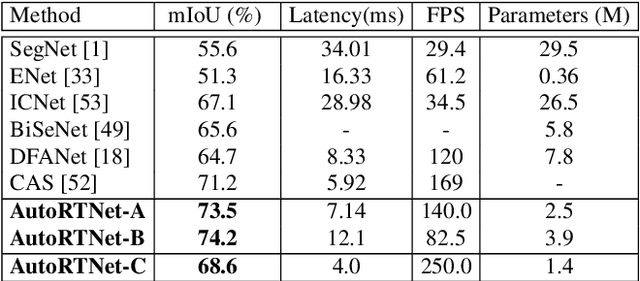
Abstract:To satisfy the stringent requirements on computational resources in the field of real-time semantic segmentation, most approaches focus on the hand-crafted design of light-weight segmentation networks. Recently, Neural Architecture Search (NAS) has been used to search for the optimal building blocks of networks automatically, but the network depth, downsampling strategy, and feature aggregation way are still set in advance by trial and error. In this paper, we propose a joint search framework, called AutoRTNet, to automate the design of these strategies. Specifically, we propose hyper-cells to jointly decide the network depth and downsampling strategy, and an aggregation cell to achieve automatic multi-scale feature aggregation. Experimental results show that AutoRTNet achieves 73.9% mIoU on the Cityscapes test set and 110.0 FPS on an NVIDIA TitanXP GPU card with 768x1536 input images.
Disturbance-immune Weight Sharing for Neural Architecture Search
Mar 29, 2020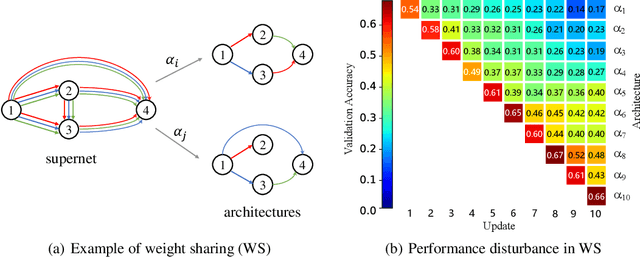
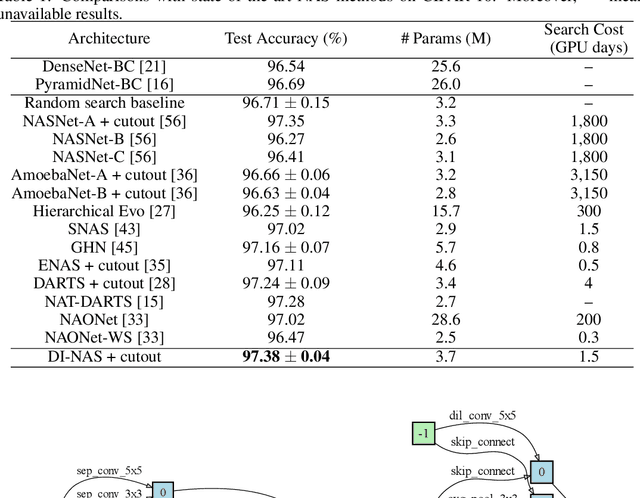
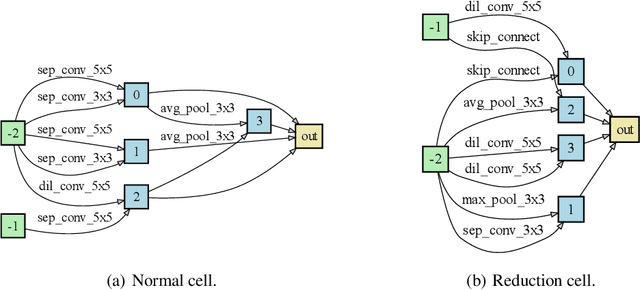
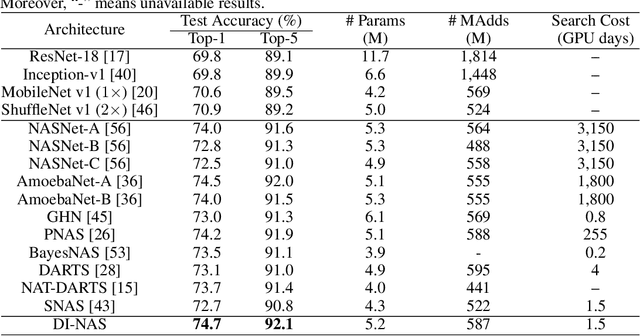
Abstract:Neural architecture search (NAS) has gained increasing attention in the community of architecture design. One of the key factors behind the success lies in the training efficiency created by the weight sharing (WS) technique. However, WS-based NAS methods often suffer from a performance disturbance (PD) issue. That is, the training of subsequent architectures inevitably disturbs the performance of previously trained architectures due to the partially shared weights. This leads to inaccurate performance estimation for the previous architectures, which makes it hard to learn a good search strategy. To alleviate the performance disturbance issue, we propose a new disturbance-immune update strategy for model updating. Specifically, to preserve the knowledge learned by previous architectures, we constrain the training of subsequent architectures in an orthogonal space via orthogonal gradient descent. Equipped with this strategy, we propose a novel disturbance-immune training scheme for NAS. We theoretically analyze the effectiveness of our strategy in alleviating the PD risk. Extensive experiments on CIFAR-10 and ImageNet verify the superiority of our method.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge