Greg S. Corrado
A deep learning system for differential diagnosis of skin diseases
Sep 11, 2019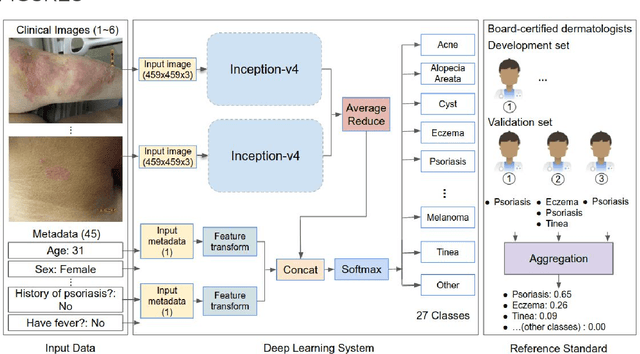
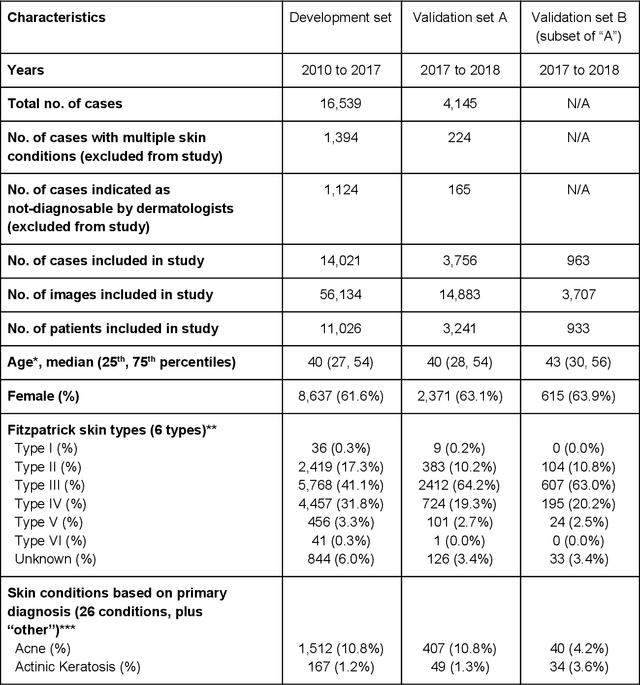
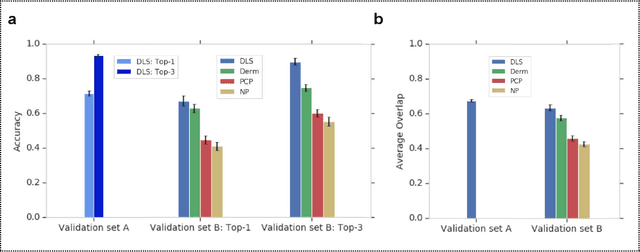
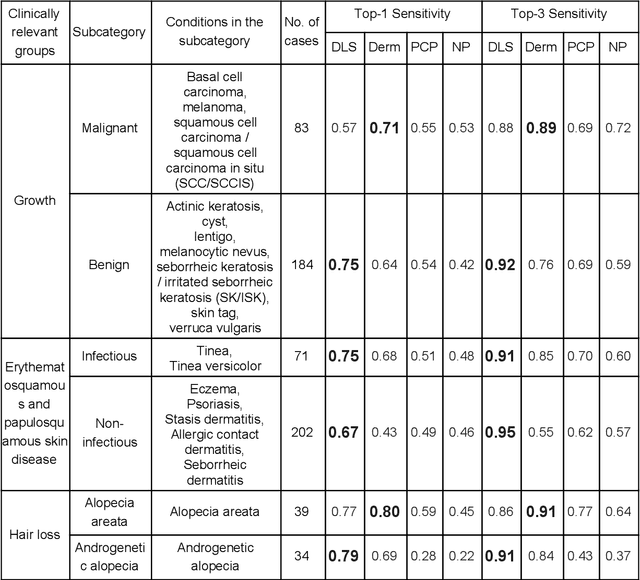
Abstract:Skin conditions affect an estimated 1.9 billion people worldwide. A shortage of dermatologists causes long wait times and leads patients to seek dermatologic care from general practitioners. However, the diagnostic accuracy of general practitioners has been reported to be only 0.24-0.70 (compared to 0.77-0.96 for dermatologists), resulting in referral errors, delays in care, and errors in diagnosis and treatment. In this paper, we developed a deep learning system (DLS) to provide a differential diagnosis of skin conditions for clinical cases (skin photographs and associated medical histories). The DLS distinguishes between 26 skin conditions that represent roughly 80% of the volume of skin conditions seen in primary care. The DLS was developed and validated using de-identified cases from a teledermatology practice serving 17 clinical sites via a temporal split: the first 14,021 cases for development and the last 3,756 cases for validation. On the validation set, where a panel of three board-certified dermatologists defined the reference standard for every case, the DLS achieved 0.71 and 0.93 top-1 and top-3 accuracies respectively. For a random subset of the validation set (n=963 cases), 18 clinicians reviewed the cases for comparison. On this subset, the DLS achieved a 0.67 top-1 accuracy, non-inferior to board-certified dermatologists (0.63, p<0.001), and higher than primary care physicians (PCPs, 0.45) and nurse practitioners (NPs, 0.41). The top-3 accuracy showed a similar trend: 0.90 DLS, 0.75 dermatologists, 0.60 PCPs, and 0.55 NPs. These results highlight the potential of the DLS to augment general practitioners to accurately diagnose skin conditions by suggesting differential diagnoses that may not have been considered. Future work will be needed to prospectively assess the clinical impact of using this tool in actual clinical workflows.
Detecting Anemia from Retinal Fundus Images
Apr 12, 2019
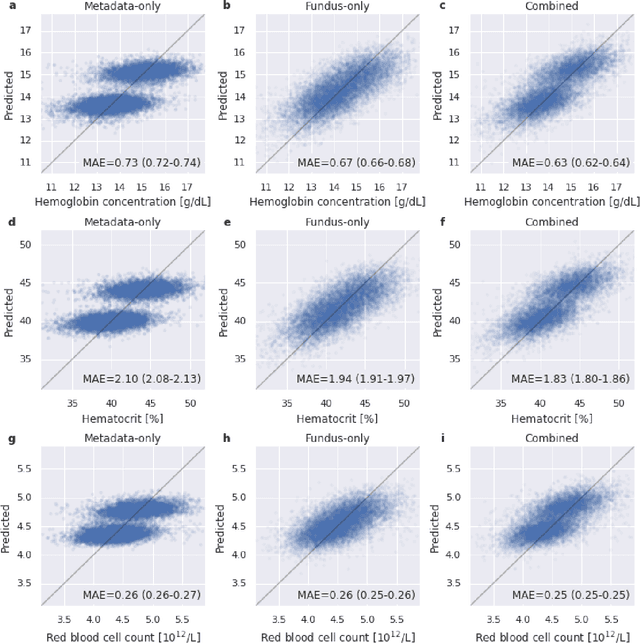
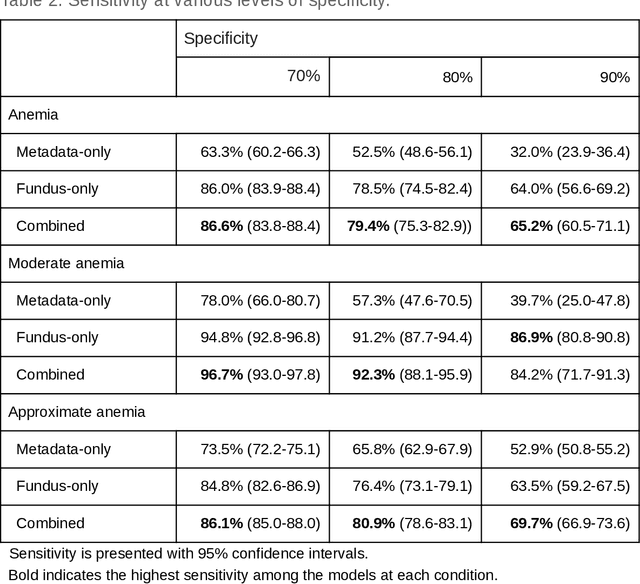
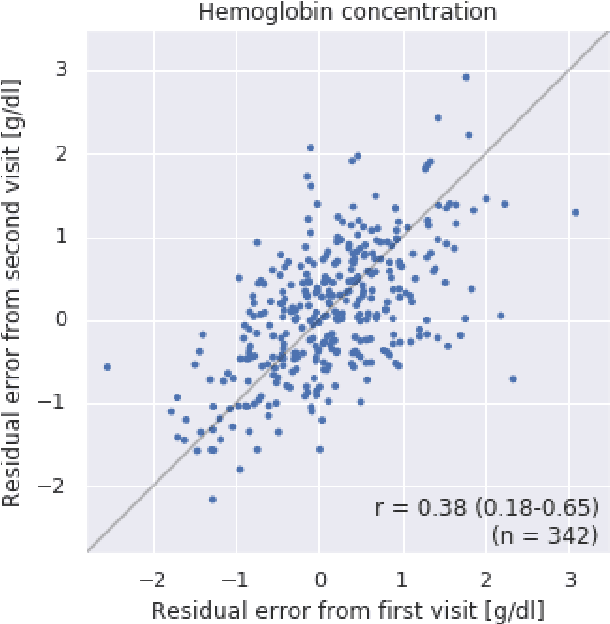
Abstract:Despite its high prevalence, anemia is often undetected due to the invasiveness and cost of screening and diagnostic tests. Though some non-invasive approaches have been developed, they are less accurate than invasive methods, resulting in an unmet need for more accurate non-invasive methods. Here, we show that deep learning-based algorithms can detect anemia and quantify several related blood measurements using retinal fundus images both in isolation and in combination with basic metadata such as patient demographics. On a validation dataset of 11,388 patients from the UK Biobank, our algorithms achieved a mean absolute error of 0.63 g/dL (95% confidence interval (CI) 0.62-0.64) in quantifying hemoglobin concentration and an area under receiver operating characteristic curve (AUC) of 0.88 (95% CI 0.86-0.89) in detecting anemia. This work shows the potential of automated non-invasive anemia screening based on fundus images, particularly in diabetic patients, who may have regular retinal imaging and are at increased risk of further morbidity and mortality from anemia.
Predicting Progression of Age-related Macular Degeneration from Fundus Images using Deep Learning
Apr 10, 2019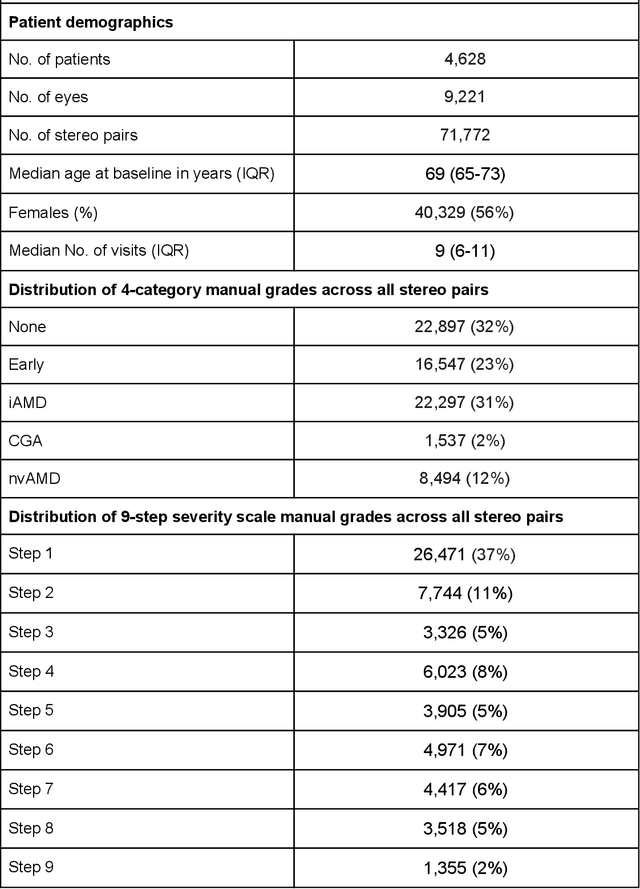
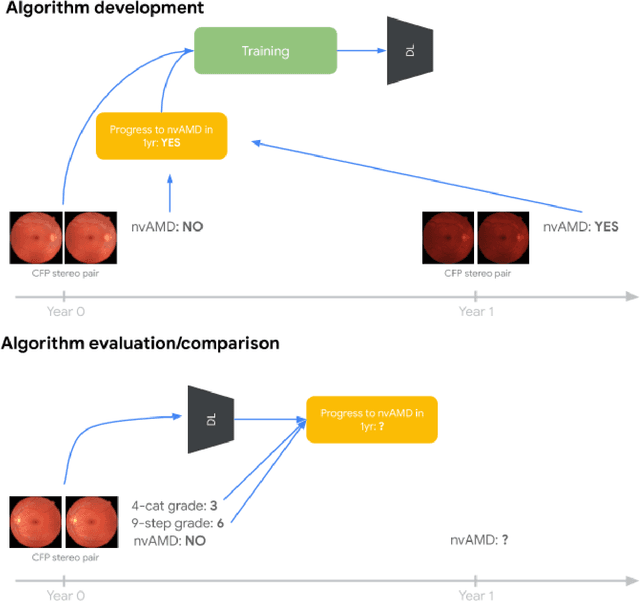
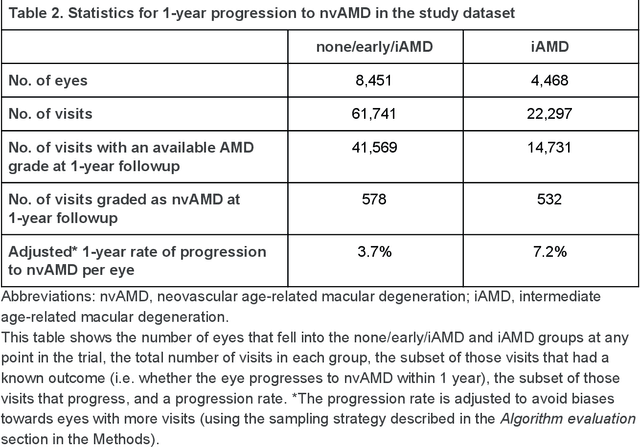
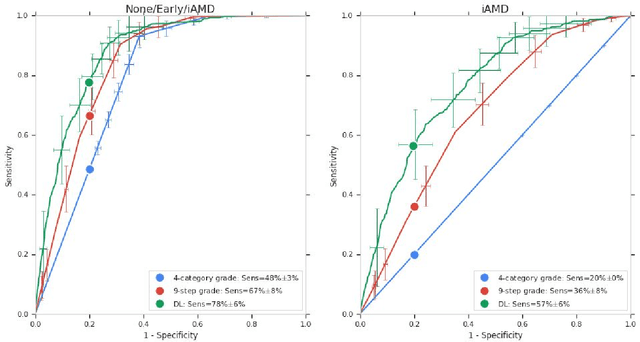
Abstract:Background: Patients with neovascular age-related macular degeneration (AMD) can avoid vision loss via certain therapy. However, methods to predict the progression to neovascular age-related macular degeneration (nvAMD) are lacking. Purpose: To develop and validate a deep learning (DL) algorithm to predict 1-year progression of eyes with no, early, or intermediate AMD to nvAMD, using color fundus photographs (CFP). Design: Development and validation of a DL algorithm. Methods: We trained a DL algorithm to predict 1-year progression to nvAMD, and used 10-fold cross-validation to evaluate this approach on two groups of eyes in the Age-Related Eye Disease Study (AREDS): none/early/intermediate AMD, and intermediate AMD (iAMD) only. We compared the DL algorithm to the manually graded 4-category and 9-step scales in the AREDS dataset. Main outcome measures: Performance of the DL algorithm was evaluated using the sensitivity at 80% specificity for progression to nvAMD. Results: The DL algorithm's sensitivity for predicting progression to nvAMD from none/early/iAMD (78+/-6%) was higher than manual grades from the 9-step scale (67+/-8%) or the 4-category scale (48+/-3%). For predicting progression specifically from iAMD, the DL algorithm's sensitivity (57+/-6%) was also higher compared to the 9-step grades (36+/-8%) and the 4-category grades (20+/-0%). Conclusions: Our DL algorithm performed better in predicting progression to nvAMD than manual grades. Future investigations are required to test the application of this DL algorithm in a real-world clinical setting.
Similar Image Search for Histopathology: SMILY
Feb 06, 2019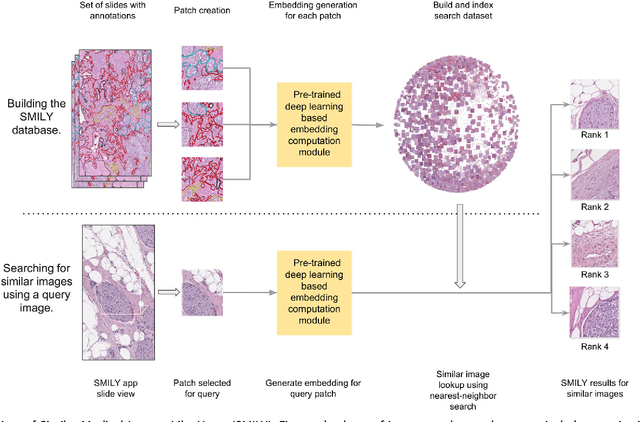
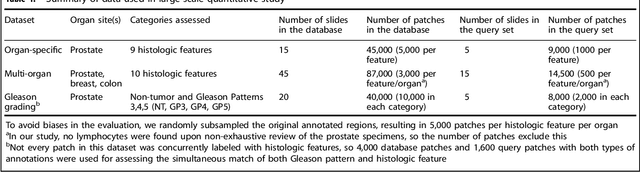
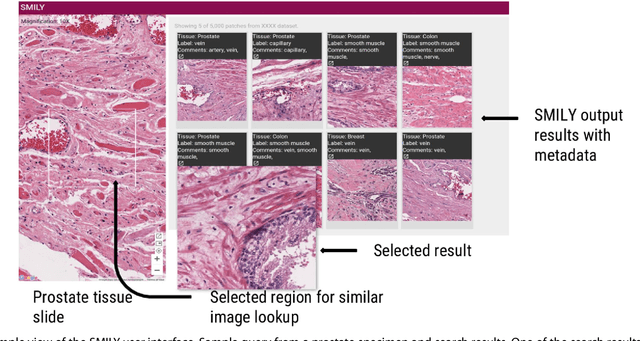
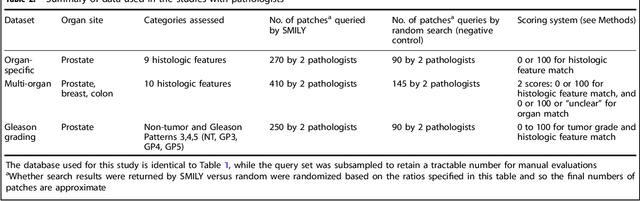
Abstract:The increasing availability of large institutional and public histopathology image datasets is enabling the searching of these datasets for diagnosis, research, and education. Though these datasets typically have associated metadata such as diagnosis or clinical notes, even carefully curated datasets rarely contain annotations of the location of regions of interest on each image. Because pathology images are extremely large (up to 100,000 pixels in each dimension), further laborious visual search of each image may be needed to find the feature of interest. In this paper, we introduce a deep learning based reverse image search tool for histopathology images: Similar Medical Images Like Yours (SMILY). We assessed SMILY's ability to retrieve search results in two ways: using pathologist-provided annotations, and via prospective studies where pathologists evaluated the quality of SMILY search results. As a negative control in the second evaluation, pathologists were blinded to whether search results were retrieved by SMILY or randomly. In both types of assessments, SMILY was able to retrieve search results with similar histologic features, organ site, and prostate cancer Gleason grade compared with the original query. SMILY may be a useful general-purpose tool in the pathologist's arsenal, to improve the efficiency of searching large archives of histopathology images, without the need to develop and implement specific tools for each application.
Deep Learning to Assess Glaucoma Risk and Associated Features in Fundus Images
Dec 21, 2018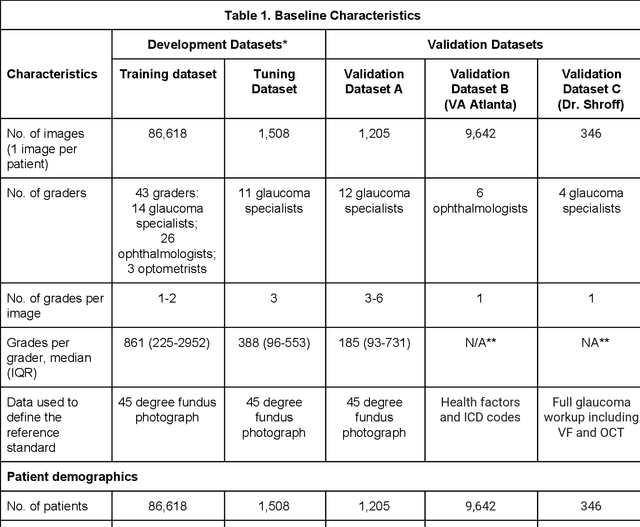
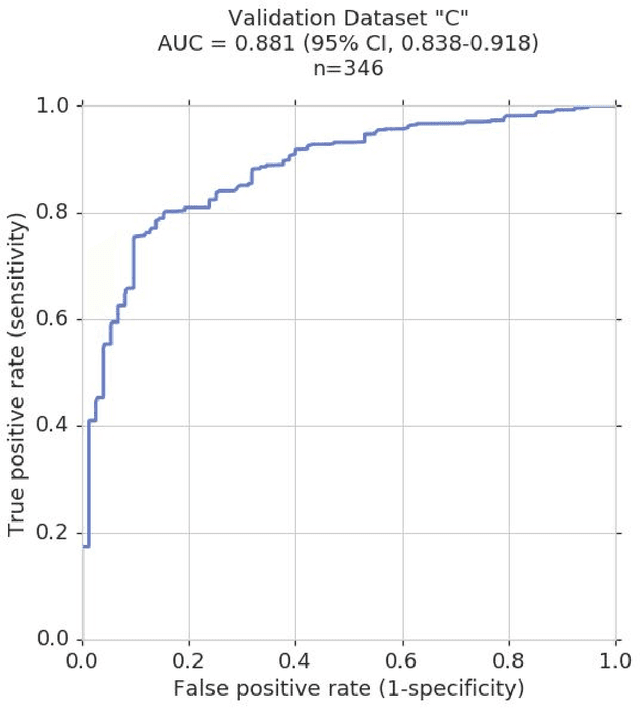
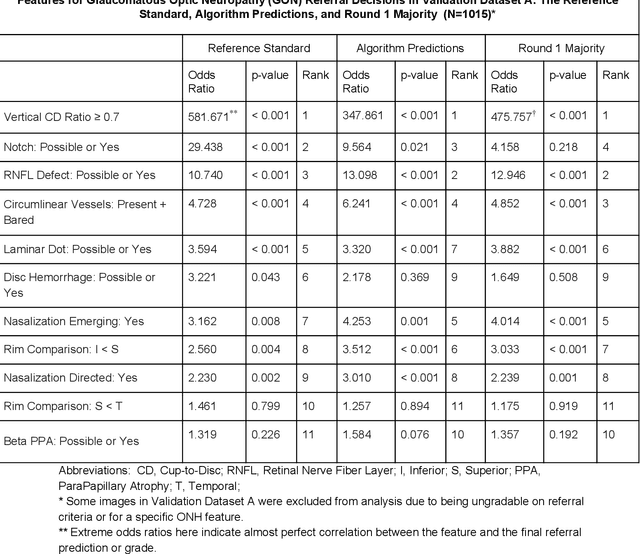
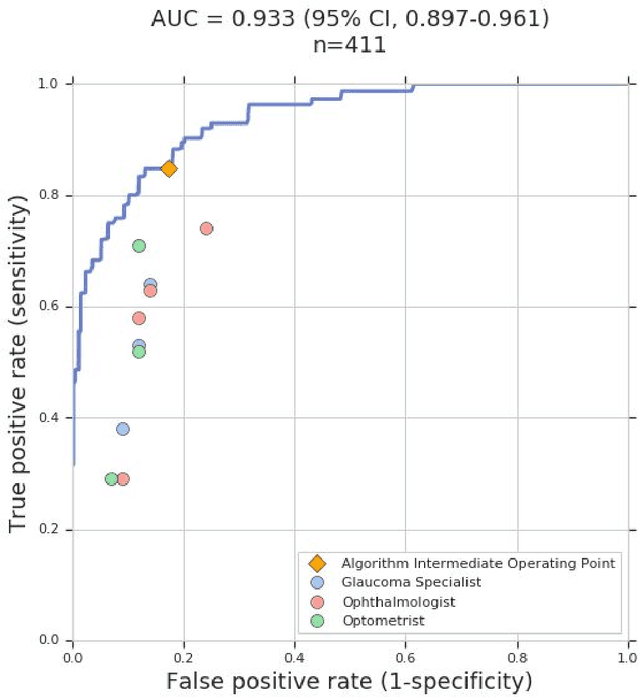
Abstract:Glaucoma is the leading cause of preventable, irreversible blindness world-wide. The disease can remain asymptomatic until severe, and an estimated 50%-90% of people with glaucoma remain undiagnosed. Thus, glaucoma screening is recommended for early detection and treatment. A cost-effective tool to detect glaucoma could expand healthcare access to a much larger patient population, but such a tool is currently unavailable. We trained a deep learning (DL) algorithm using a retrospective dataset of 58,033 images, assessed for gradability, glaucomatous optic nerve head (ONH) features, and referable glaucoma risk. The resultant algorithm was validated using 2 separate datasets. For referable glaucoma risk, the algorithm had an AUC of 0.940 (95%CI, 0.922-0.955) in validation dataset "A" (1,205 images, 1 image/patient; 19% referable where images were adjudicated by panels of fellowship-trained glaucoma specialists) and 0.858 (95% CI, 0.836-0.878) in validation dataset "B" (17,593 images from 9,643 patients; 9.2% referable where images were from the Atlanta Veterans Affairs Eye Clinic diabetic teleretinal screening program using clinical referral decisions as the reference standard). Additionally, we found that the presence of vertical cup-to-disc ratio >= 0.7, neuroretinal rim notching, retinal nerve fiber layer defect, and bared circumlinear vessels contributed most to referable glaucoma risk assessment by both glaucoma specialists and the algorithm. Algorithm AUCs ranged between 0.608-0.977 for glaucomatous ONH features. The DL algorithm was significantly more sensitive than 6 of 10 graders, including 2 of 3 glaucoma specialists, with comparable or higher specificity relative to all graders. A DL algorithm trained on fundus images alone can detect referable glaucoma risk with higher sensitivity and comparable specificity to eye care providers.
Microscope 2.0: An Augmented Reality Microscope with Real-time Artificial Intelligence Integration
Dec 04, 2018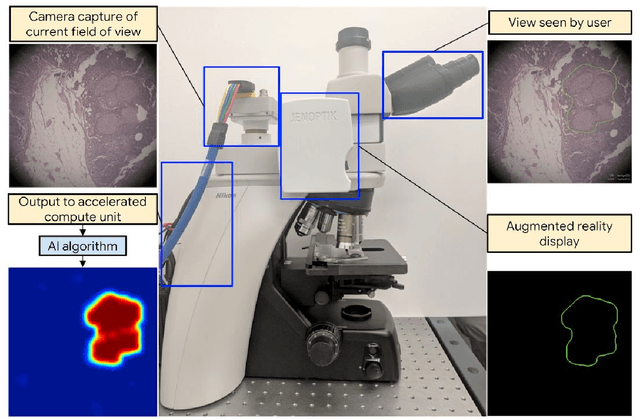
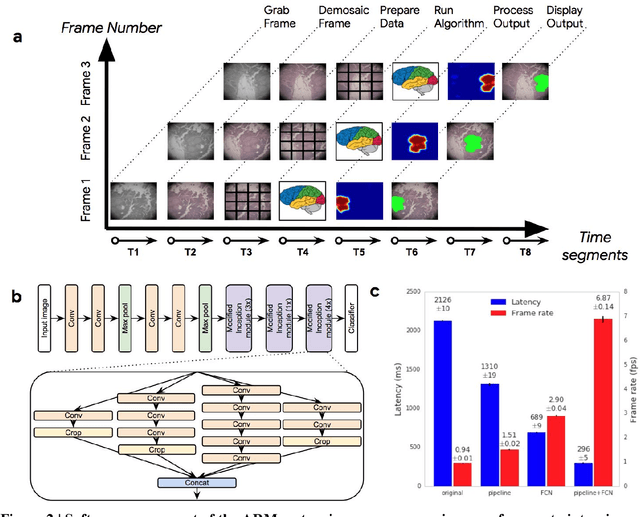
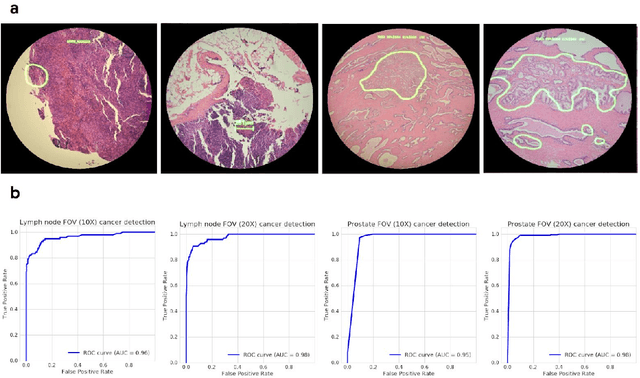
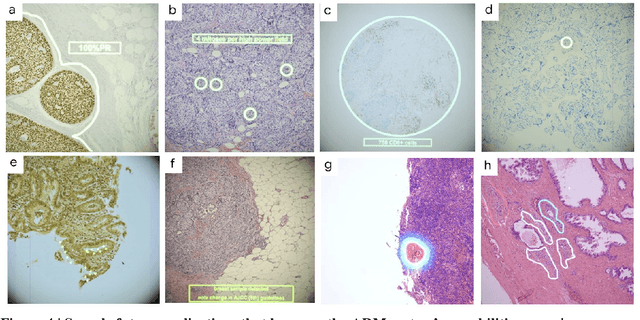
Abstract:The brightfield microscope is instrumental in the visual examination of both biological and physical samples at sub-millimeter scales. One key clinical application has been in cancer histopathology, where the microscopic assessment of the tissue samples is used for the diagnosis and staging of cancer and thus guides clinical therapy. However, the interpretation of these samples is inherently subjective, resulting in significant diagnostic variability. Moreover, in many regions of the world, access to pathologists is severely limited due to lack of trained personnel. In this regard, Artificial Intelligence (AI) based tools promise to improve the access and quality of healthcare. However, despite significant advances in AI research, integration of these tools into real-world cancer diagnosis workflows remains challenging because of the costs of image digitization and difficulties in deploying AI solutions. Here we propose a cost-effective solution to the integration of AI: the Augmented Reality Microscope (ARM). The ARM overlays AI-based information onto the current view of the sample through the optical pathway in real-time, enabling seamless integration of AI into the regular microscopy workflow. We demonstrate the utility of ARM in the detection of lymph node metastases in breast cancer and the identification of prostate cancer with a latency that supports real-time workflows. We anticipate that ARM will remove barriers towards the use of AI in microscopic analysis and thus improve the accuracy and efficiency of cancer diagnosis. This approach is applicable to other microscopy tasks and AI algorithms in the life sciences and beyond.
Development and Validation of a Deep Learning Algorithm for Improving Gleason Scoring of Prostate Cancer
Nov 15, 2018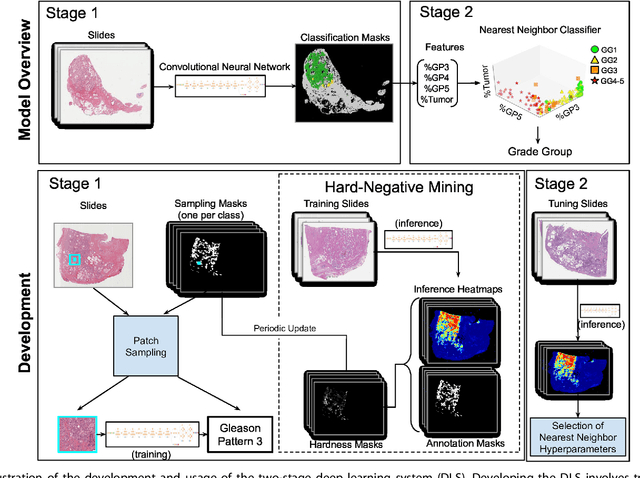
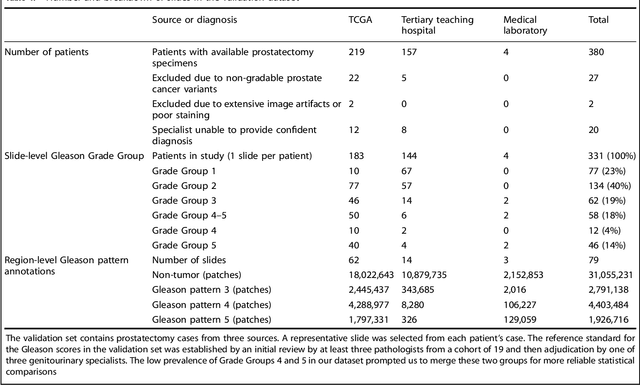
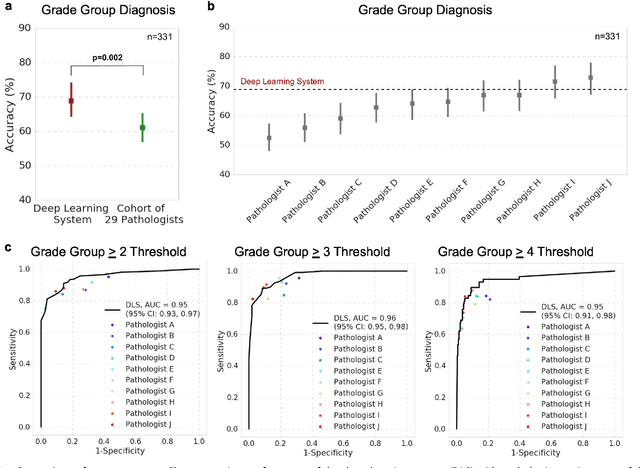
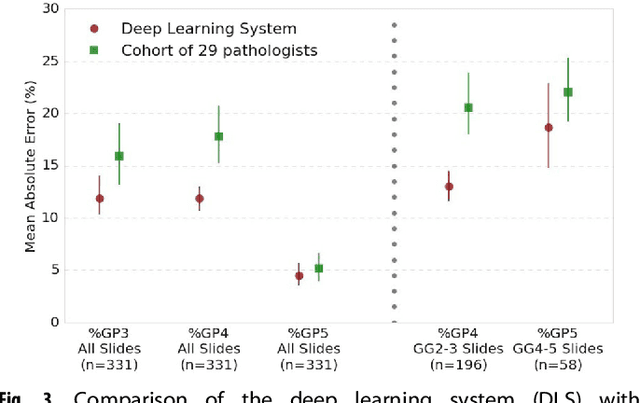
Abstract:For prostate cancer patients, the Gleason score is one of the most important prognostic factors, potentially determining treatment independent of the stage. However, Gleason scoring is based on subjective microscopic examination of tumor morphology and suffers from poor reproducibility. Here we present a deep learning system (DLS) for Gleason scoring whole-slide images of prostatectomies. Our system was developed using 112 million pathologist-annotated image patches from 1,226 slides, and evaluated on an independent validation dataset of 331 slides, where the reference standard was established by genitourinary specialist pathologists. On the validation dataset, the mean accuracy among 29 general pathologists was 0.61. The DLS achieved a significantly higher diagnostic accuracy of 0.70 (p=0.002) and trended towards better patient risk stratification in correlations to clinical follow-up data. Our approach could improve the accuracy of Gleason scoring and subsequent therapy decisions, particularly where specialist expertise is unavailable. The DLS also goes beyond the current Gleason system to more finely characterize and quantitate tumor morphology, providing opportunities for refinement of the Gleason system itself.
Grader variability and the importance of reference standards for evaluating machine learning models for diabetic retinopathy
Jul 03, 2018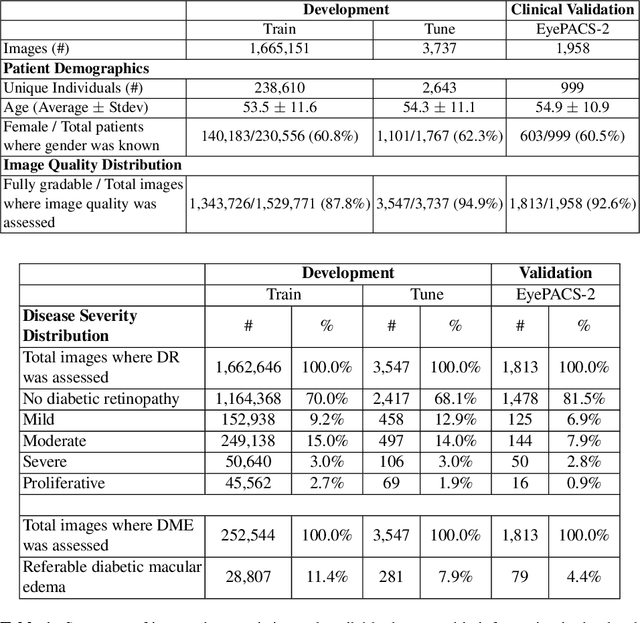
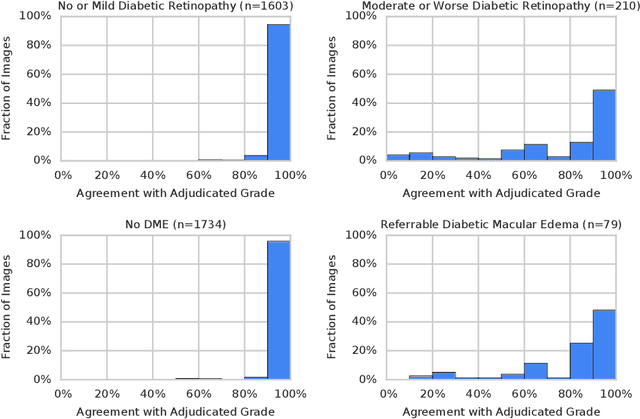
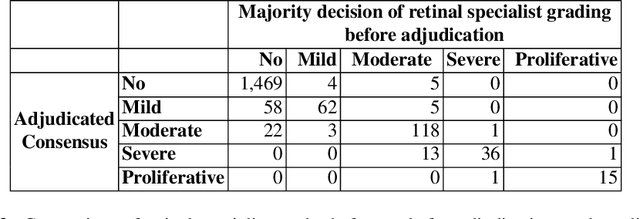
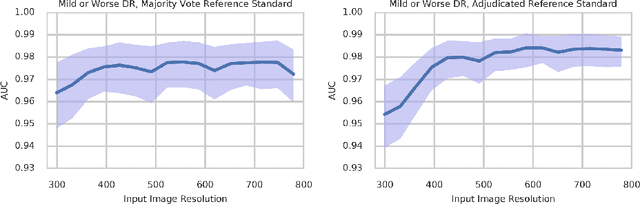
Abstract:Diabetic retinopathy (DR) and diabetic macular edema are common complications of diabetes which can lead to vision loss. The grading of DR is a fairly complex process that requires the detection of fine features such as microaneurysms, intraretinal hemorrhages, and intraretinal microvascular abnormalities. Because of this, there can be a fair amount of grader variability. There are different methods of obtaining the reference standard and resolving disagreements between graders, and while it is usually accepted that adjudication until full consensus will yield the best reference standard, the difference between various methods of resolving disagreements has not been examined extensively. In this study, we examine the variability in different methods of grading, definitions of reference standards, and their effects on building deep learning models for the detection of diabetic eye disease. We find that a small set of adjudicated DR grades allows substantial improvements in algorithm performance. The resulting algorithm's performance was on par with that of individual U.S. board-certified ophthalmologists and retinal specialists.
Deep learning for predicting refractive error from retinal fundus images
Dec 21, 2017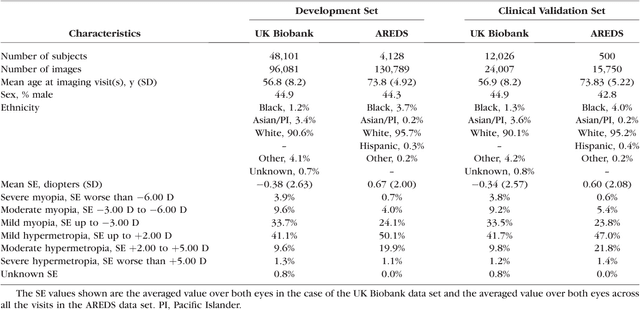
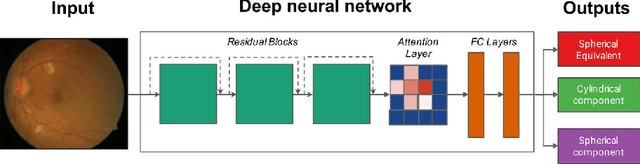
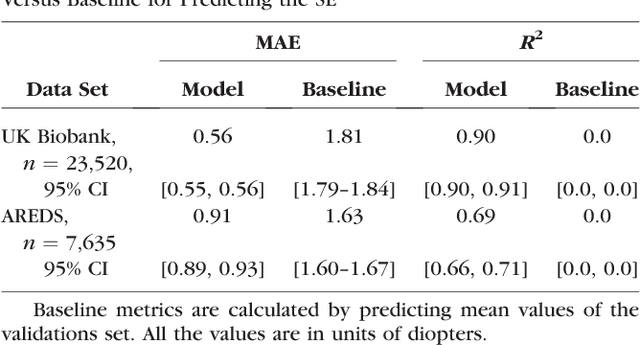
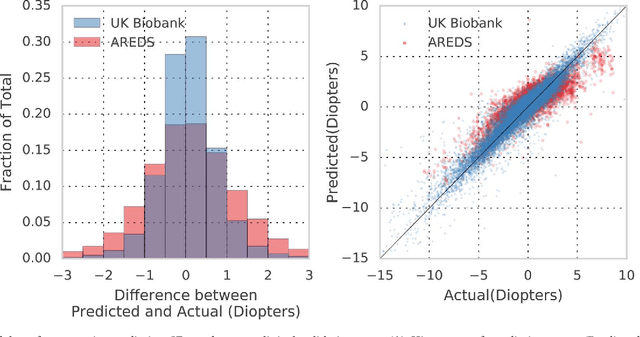
Abstract:Refractive error, one of the leading cause of visual impairment, can be corrected by simple interventions like prescribing eyeglasses. We trained a deep learning algorithm to predict refractive error from the fundus photographs from participants in the UK Biobank cohort, which were 45 degree field of view images and the AREDS clinical trial, which contained 30 degree field of view images. Our model use the "attention" method to identify features that are correlated with refractive error. Mean absolute error (MAE) of the algorithm's prediction compared to the refractive error obtained in the AREDS and UK Biobank. The resulting algorithm had a MAE of 0.56 diopters (95% CI: 0.55-0.56) for estimating spherical equivalent on the UK Biobank dataset and 0.91 diopters (95% CI: 0.89-0.92) for the AREDS dataset. The baseline expected MAE (obtained by simply predicting the mean of this population) was 1.81 diopters (95% CI: 1.79-1.84) for UK Biobank and 1.63 (95% CI: 1.60-1.67) for AREDS. Attention maps suggested that the foveal region was one of the most important areas used by the algorithm to make this prediction, though other regions also contribute to the prediction. The ability to estimate refractive error with high accuracy from retinal fundus photos has not been previously known and demonstrates that deep learning can be applied to make novel predictions from medical images. Given that several groups have recently shown that it is feasible to obtain retinal fundus photos using mobile phones and inexpensive attachments, this work may be particularly relevant in regions of the world where autorefractors may not be readily available.
Detecting Cancer Metastases on Gigapixel Pathology Images
Mar 08, 2017
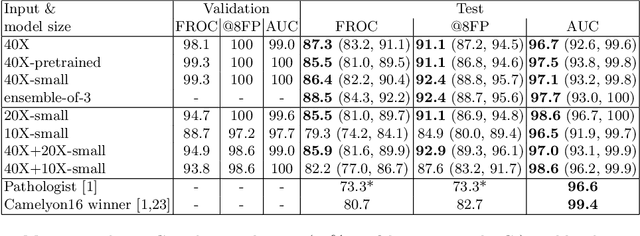
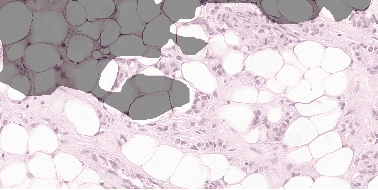
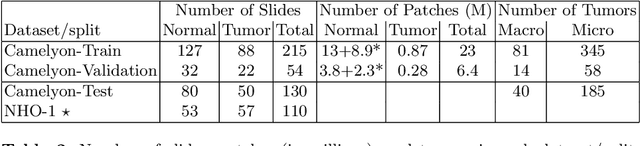
Abstract:Each year, the treatment decisions for more than 230,000 breast cancer patients in the U.S. hinge on whether the cancer has metastasized away from the breast. Metastasis detection is currently performed by pathologists reviewing large expanses of biological tissues. This process is labor intensive and error-prone. We present a framework to automatically detect and localize tumors as small as 100 x 100 pixels in gigapixel microscopy images sized 100,000 x 100,000 pixels. Our method leverages a convolutional neural network (CNN) architecture and obtains state-of-the-art results on the Camelyon16 dataset in the challenging lesion-level tumor detection task. At 8 false positives per image, we detect 92.4% of the tumors, relative to 82.7% by the previous best automated approach. For comparison, a human pathologist attempting exhaustive search achieved 73.2% sensitivity. We achieve image-level AUC scores above 97% on both the Camelyon16 test set and an independent set of 110 slides. In addition, we discover that two slides in the Camelyon16 training set were erroneously labeled normal. Our approach could considerably reduce false negative rates in metastasis detection.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge