Dustin Tran
Dima
BatchEnsemble: An Alternative Approach to Efficient Ensemble and Lifelong Learning
Feb 20, 2020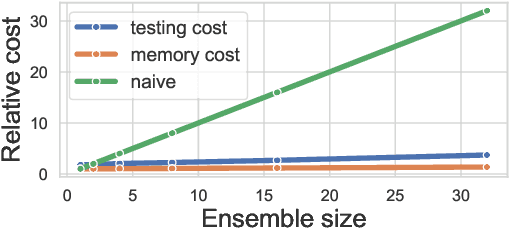

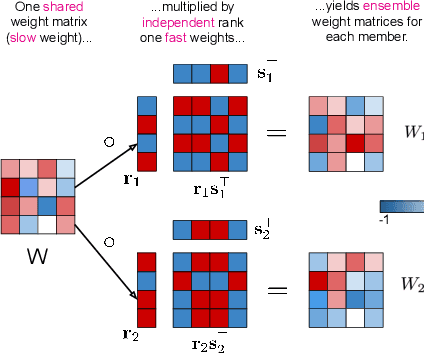
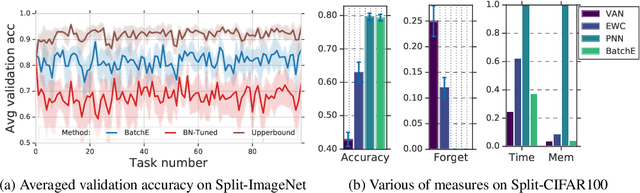
Abstract:Ensembles, where multiple neural networks are trained individually and their predictions are averaged, have been shown to be widely successful for improving both the accuracy and predictive uncertainty of single neural networks. However, an ensemble's cost for both training and testing increases linearly with the number of networks, which quickly becomes untenable. In this paper, we propose BatchEnsemble, an ensemble method whose computational and memory costs are significantly lower than typical ensembles. BatchEnsemble achieves this by defining each weight matrix to be the Hadamard product of a shared weight among all ensemble members and a rank-one matrix per member. Unlike ensembles, BatchEnsemble is not only parallelizable across devices, where one device trains one member, but also parallelizable within a device, where multiple ensemble members are updated simultaneously for a given mini-batch. Across CIFAR-10, CIFAR-100, WMT14 EN-DE/EN-FR translation, and out-of-distribution tasks, BatchEnsemble yields competitive accuracy and uncertainties as typical ensembles; the speedup at test time is 3X and memory reduction is 3X at an ensemble of size 4. We also apply BatchEnsemble to lifelong learning, where on Split-CIFAR-100, BatchEnsemble yields comparable performance to progressive neural networks while having a much lower computational and memory costs. We further show that BatchEnsemble can easily scale up to lifelong learning on Split-ImageNet which involves 100 sequential learning tasks.
On the Discrepancy between Density Estimation and Sequence Generation
Feb 17, 2020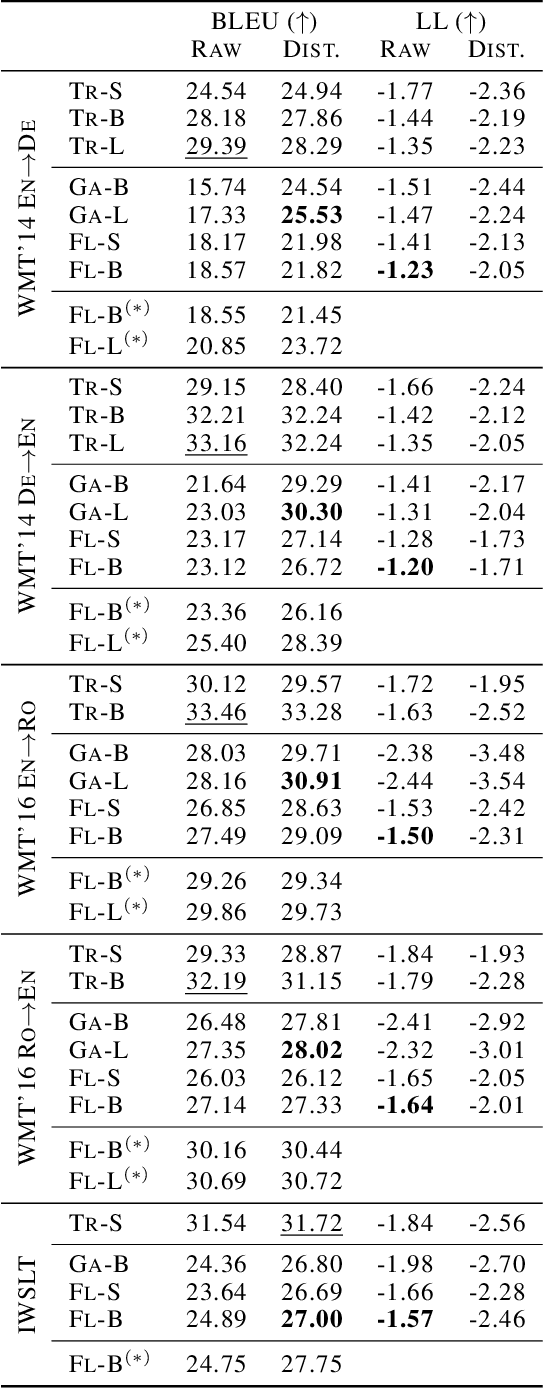
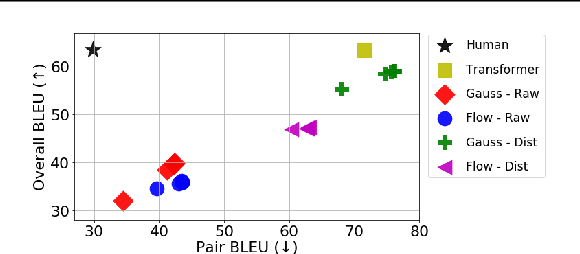

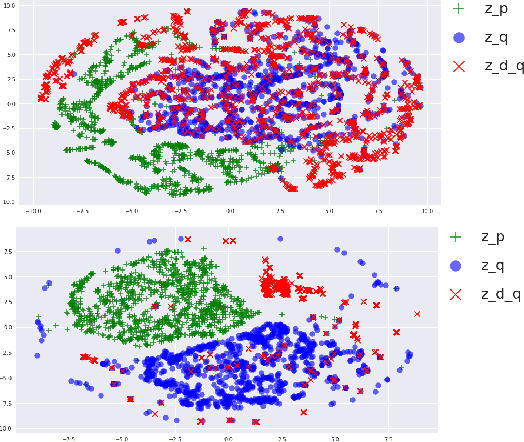
Abstract:Many sequence-to-sequence generation tasks, including machine translation and text-to-speech, can be posed as estimating the density of the output y given the input x: p(y|x). Given this interpretation, it is natural to evaluate sequence-to-sequence models using conditional log-likelihood on a test set. However, the goal of sequence-to-sequence generation (or structured prediction) is to find the best output y^ given an input x, and each task has its own downstream metric R that scores a model output by comparing against a set of references y*: R(y^, y* | x). While we hope that a model that excels in density estimation also performs well on the downstream metric, the exact correlation has not been studied for sequence generation tasks. In this paper, by comparing several density estimators on five machine translation tasks, we find that the correlation between rankings of models based on log-likelihood and BLEU varies significantly depending on the range of the model families being compared. First, log-likelihood is highly correlated with BLEU when we consider models within the same family (e.g. autoregressive models, or latent variable models with the same parameterization of the prior). However, we observe no correlation between rankings of models across different families: (1) among non-autoregressive latent variable models, a flexible prior distribution is better at density estimation but gives worse generation quality than a simple prior, and (2) autoregressive models offer the best translation performance overall, while latent variable models with a normalizing flow prior give the highest held-out log-likelihood across all datasets. Therefore, we recommend using a simple prior for the latent variable non-autoregressive model when fast generation speed is desired.
Analyzing the Role of Model Uncertainty for Electronic Health Records
Jun 10, 2019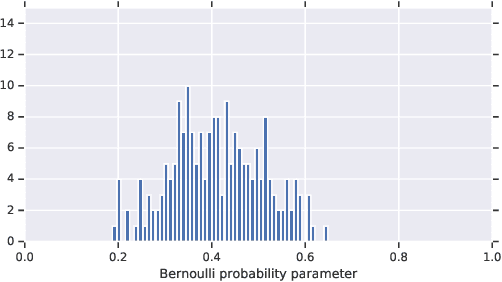
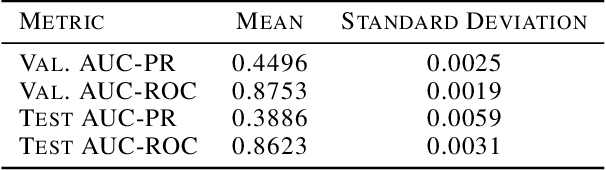
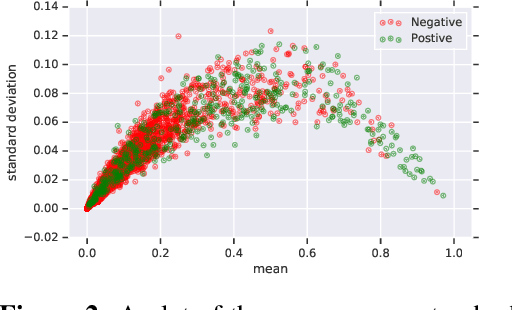
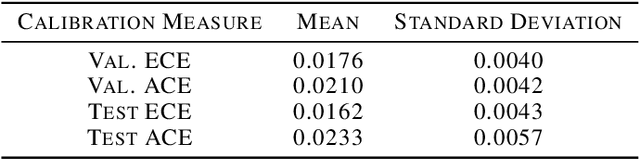
Abstract:In medicine, both ethical and monetary costs of incorrect predictions can be significant, and the complexity of the problems often necessitates increasingly complex models. Recent work has shown that changing just the random seed is enough for otherwise well-tuned deep neural networks to vary in their individual predicted probabilities. In light of this, we investigate the role of model uncertainty methods in the medical domain. Using RNN ensembles and various Bayesian RNNs, we show that population-level metrics, such as AUC-PR, AUC-ROC, log-likelihood, and calibration error, do not capture model uncertainty. Meanwhile, the presence of significant variability in patient-specific predictions and optimal decisions motivates the need for capturing model uncertainty. Understanding the uncertainty for individual patients is an area with clear clinical impact, such as determining when a model decision is likely to be brittle. We further show that RNNs with only Bayesian embeddings can be a more efficient way to capture model uncertainty compared to ensembles, and we analyze how model uncertainty is impacted across individual input features and patient subgroups.
Discrete Flows: Invertible Generative Models of Discrete Data
May 24, 2019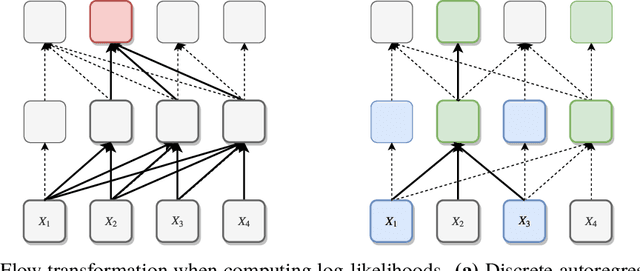

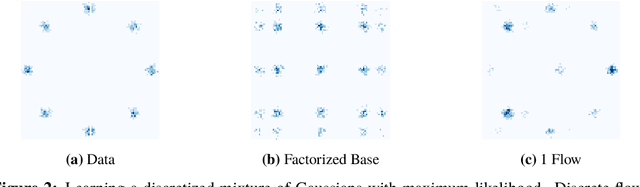
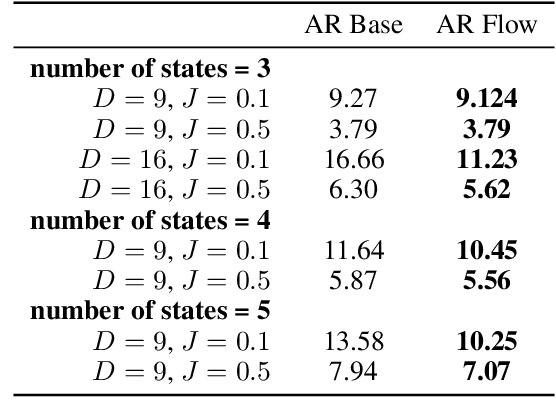
Abstract:While normalizing flows have led to significant advances in modeling high-dimensional continuous distributions, their applicability to discrete distributions remains unknown. In this paper, we show that flows can in fact be extended to discrete events---and under a simple change-of-variables formula not requiring log-determinant-Jacobian computations. Discrete flows have numerous applications. We consider two flow architectures: discrete autoregressive flows that enable bidirectionality, allowing, for example, tokens in text to depend on both left-to-right and right-to-left contexts in an exact language model; and discrete bipartite flows that enable efficient non-autoregressive generation as in RealNVP. Empirically, we find that discrete autoregressive flows outperform autoregressive baselines on synthetic discrete distributions, an addition task, and Potts models; and bipartite flows can obtain competitive performance with autoregressive baselines on character-level language modeling for Penn Tree Bank and text8.
Measuring Calibration in Deep Learning
Apr 02, 2019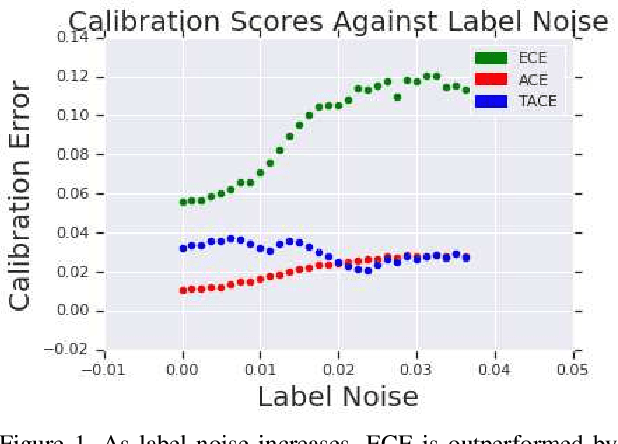
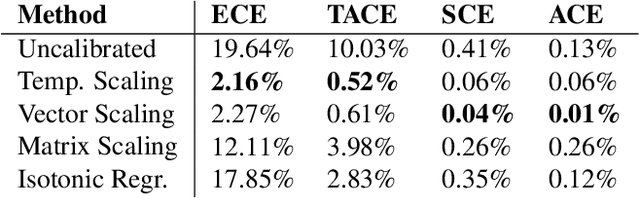
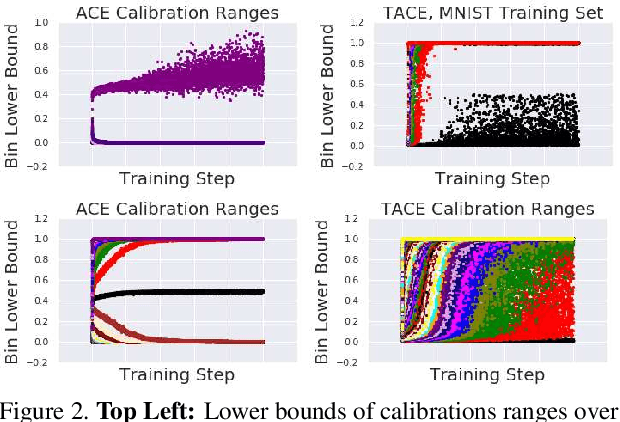
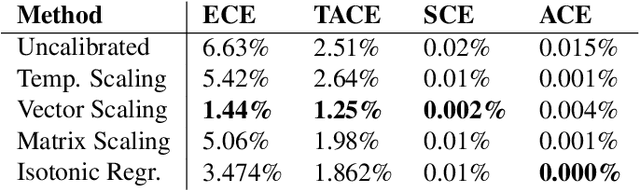
Abstract:The reliability of a machine learning model's confidence in its predictions is critical for highrisk applications. Calibration-the idea that a model's predicted probabilities of outcomes reflect true probabilities of those outcomes-formalizes this notion. While analyzing the calibration of deep neural networks, we've identified core problems with the way calibration is currently measured. We design the Thresholded Adaptive Calibration Error (TACE) metric to resolve these pathologies and show that it outperforms other metrics, especially in settings where predictions beyond the maximum prediction that is chosen as the output class matter. There are many cases where what a practitioner cares about is the calibration of a specific prediction, and so we introduce a dynamic programming based Prediction Specific Calibration Error (PSCE) that smoothly considers the calibration of nearby predictions to give an estimate of the calibration error of a specific prediction.
NeuTra-lizing Bad Geometry in Hamiltonian Monte Carlo Using Neural Transport
Mar 09, 2019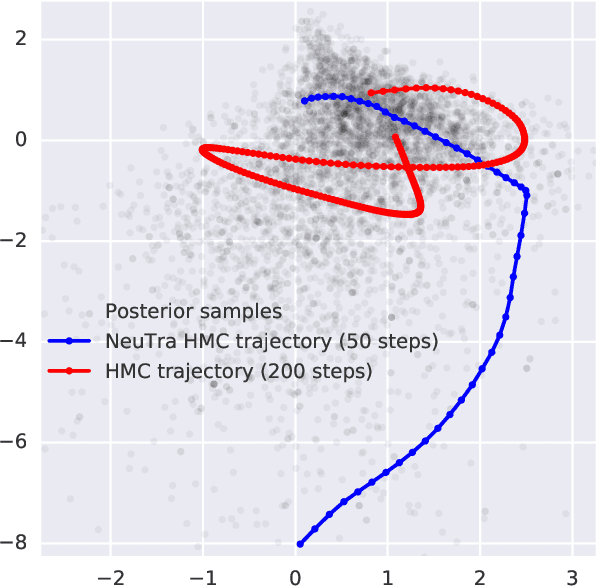
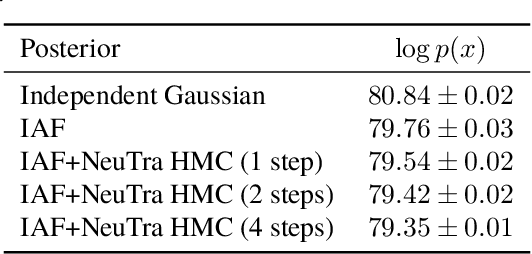

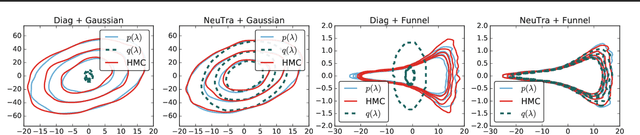
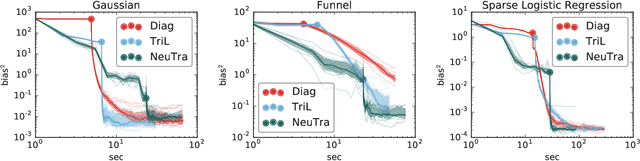
Abstract:Hamiltonian Monte Carlo is a powerful algorithm for sampling from difficult-to-normalize posterior distributions. However, when the geometry of the posterior is unfavorable, it may take many expensive evaluations of the target distribution and its gradient to converge and mix. We propose neural transport (NeuTra) HMC, a technique for learning to correct this sort of unfavorable geometry using inverse autoregressive flows (IAF), a powerful neural variational inference technique. The IAF is trained to minimize the KL divergence from an isotropic Gaussian to the warped posterior, and then HMC sampling is performed in the warped space. We evaluate NeuTra HMC on a variety of synthetic and real problems, and find that it significantly outperforms vanilla HMC both in time to reach the stationary distribution and asymptotic effective-sample-size rates.
Bayesian Layers: A Module for Neural Network Uncertainty
Dec 11, 2018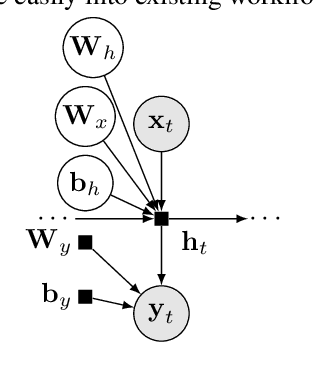
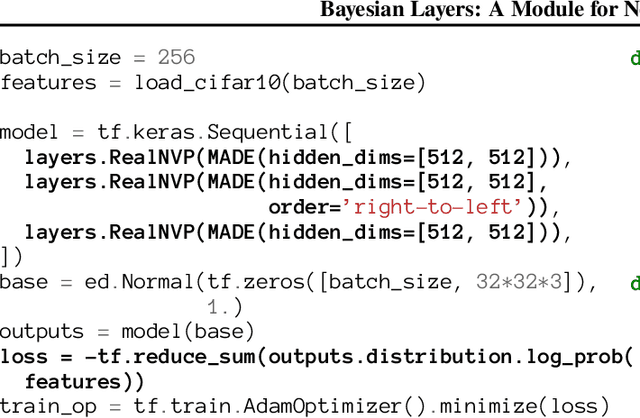
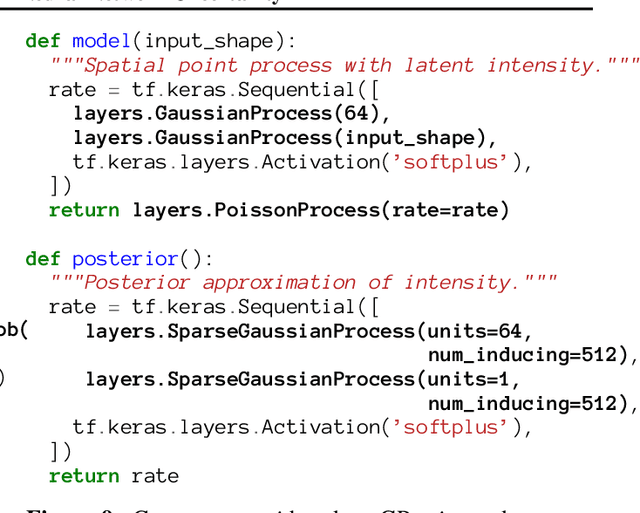


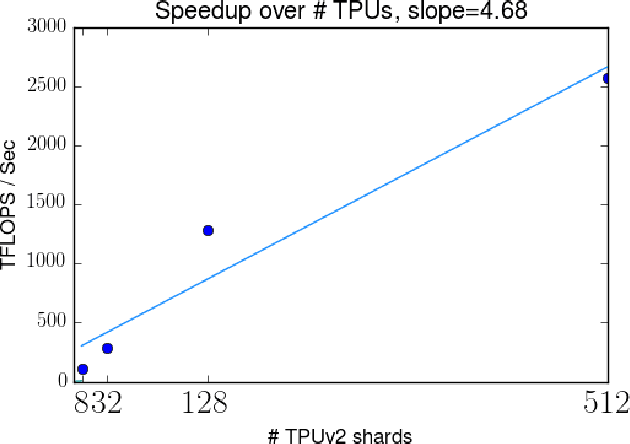
Abstract:We describe Bayesian Layers, a module designed for fast experimentation with neural network uncertainty. It extends neural network libraries with layers capturing uncertainty over weights (Bayesian neural nets), pre-activation units (dropout), activations ("stochastic output layers"), and the function itself (Gaussian processes). With reversible layers, one can also propagate uncertainty from input to output such as for flow-based distributions and constant-memory backpropagation. Bayesian Layers are a drop-in replacement for other layers, maintaining core features that one typically desires for experimentation. As demonstration, we fit a 10-billion parameter "Bayesian Transformer" on 512 TPUv2 cores, which replaces attention layers with their Bayesian counterpart.
Simple, Distributed, and Accelerated Probabilistic Programming
Nov 29, 2018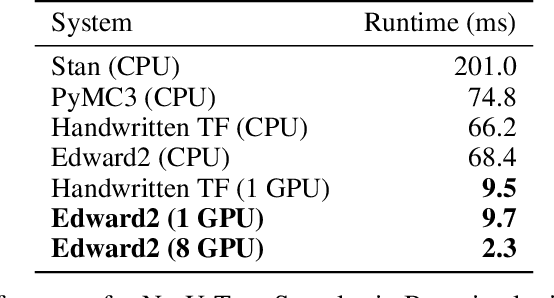
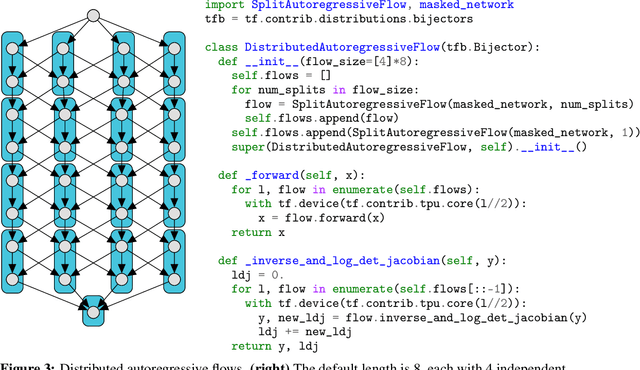
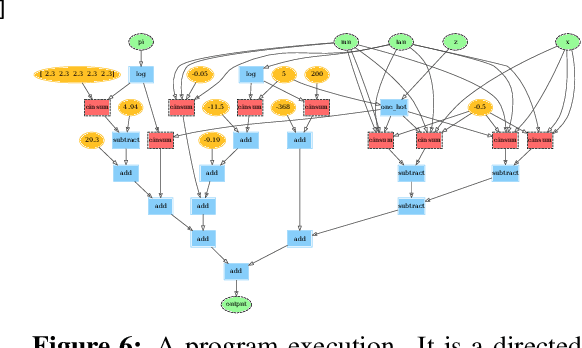
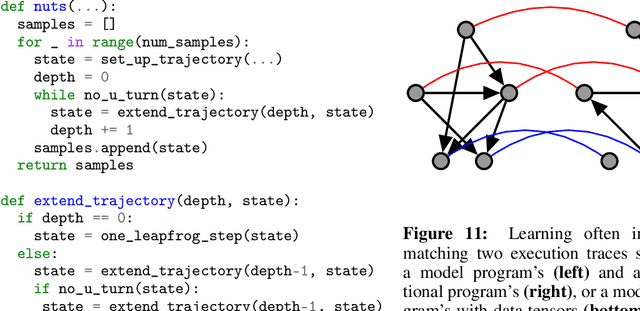
Abstract:We describe a simple, low-level approach for embedding probabilistic programming in a deep learning ecosystem. In particular, we distill probabilistic programming down to a single abstraction---the random variable. Our lightweight implementation in TensorFlow enables numerous applications: a model-parallel variational auto-encoder (VAE) with 2nd-generation tensor processing units (TPUv2s); a data-parallel autoregressive model (Image Transformer) with TPUv2s; and multi-GPU No-U-Turn Sampler (NUTS). For both a state-of-the-art VAE on 64x64 ImageNet and Image Transformer on 256x256 CelebA-HQ, our approach achieves an optimal linear speedup from 1 to 256 TPUv2 chips. With NUTS, we see a 100x speedup on GPUs over Stan and 37x over PyMC3.
Autoconj: Recognizing and Exploiting Conjugacy Without a Domain-Specific Language
Nov 29, 2018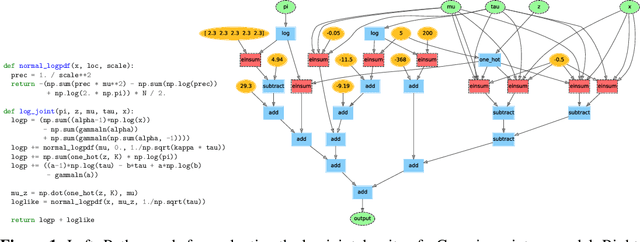
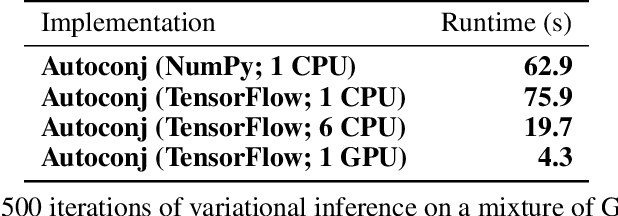

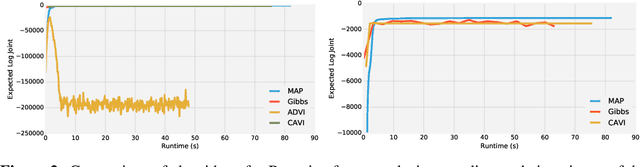
Abstract:Deriving conditional and marginal distributions using conjugacy relationships can be time consuming and error prone. In this paper, we propose a strategy for automating such derivations. Unlike previous systems which focus on relationships between pairs of random variables, our system (which we call Autoconj) operates directly on Python functions that compute log-joint distribution functions. Autoconj provides support for conjugacy-exploiting algorithms in any Python embedded PPL. This paves the way for accelerating development of novel inference algorithms and structure-exploiting modeling strategies.
Mesh-TensorFlow: Deep Learning for Supercomputers
Nov 05, 2018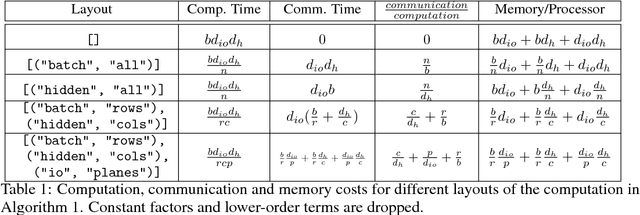
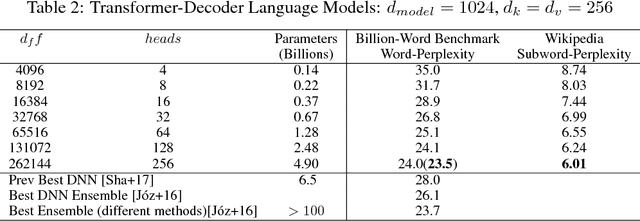
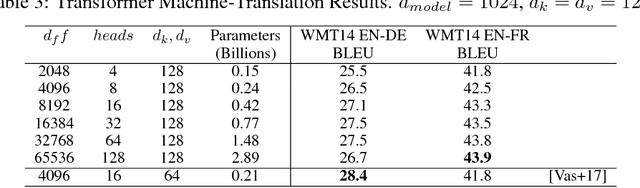
Abstract:Batch-splitting (data-parallelism) is the dominant distributed Deep Neural Network (DNN) training strategy, due to its universal applicability and its amenability to Single-Program-Multiple-Data (SPMD) programming. However, batch-splitting suffers from problems including the inability to train very large models (due to memory constraints), high latency, and inefficiency at small batch sizes. All of these can be solved by more general distribution strategies (model-parallelism). Unfortunately, efficient model-parallel algorithms tend to be complicated to discover, describe, and to implement, particularly on large clusters. We introduce Mesh-TensorFlow, a language for specifying a general class of distributed tensor computations. Where data-parallelism can be viewed as splitting tensors and operations along the "batch" dimension, in Mesh-TensorFlow, the user can specify any tensor-dimensions to be split across any dimensions of a multi-dimensional mesh of processors. A Mesh-TensorFlow graph compiles into a SPMD program consisting of parallel operations coupled with collective communication primitives such as Allreduce. We use Mesh-TensorFlow to implement an efficient data-parallel, model-parallel version of the Transformer sequence-to-sequence model. Using TPU meshes of up to 512 cores, we train Transformer models with up to 5 billion parameters, surpassing state of the art results on WMT'14 English-to-French translation task and the one-billion-word language modeling benchmark. Mesh-Tensorflow is available at https://github.com/tensorflow/mesh .
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge