Chi-Wing Fu
Point Cloud Upsampling via Disentangled Refinement
Jun 09, 2021
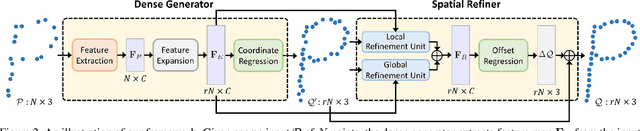
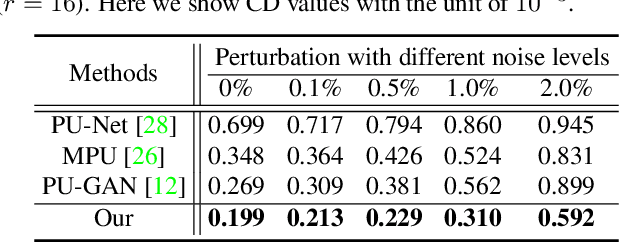
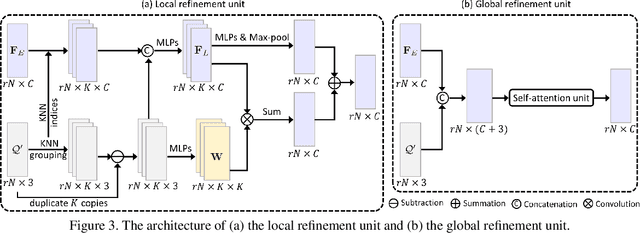
Abstract:Point clouds produced by 3D scanning are often sparse, non-uniform, and noisy. Recent upsampling approaches aim to generate a dense point set, while achieving both distribution uniformity and proximity-to-surface, and possibly amending small holes, all in a single network. After revisiting the task, we propose to disentangle the task based on its multi-objective nature and formulate two cascaded sub-networks, a dense generator and a spatial refiner. The dense generator infers a coarse but dense output that roughly describes the underlying surface, while the spatial refiner further fine-tunes the coarse output by adjusting the location of each point. Specifically, we design a pair of local and global refinement units in the spatial refiner to evolve a coarse feature map. Also, in the spatial refiner, we regress a per-point offset vector to further adjust the coarse outputs in fine-scale. Extensive qualitative and quantitative results on both synthetic and real-scanned datasets demonstrate the superiority of our method over the state-of-the-arts.
SE-SSD: Self-Ensembling Single-Stage Object Detector From Point Cloud
Apr 20, 2021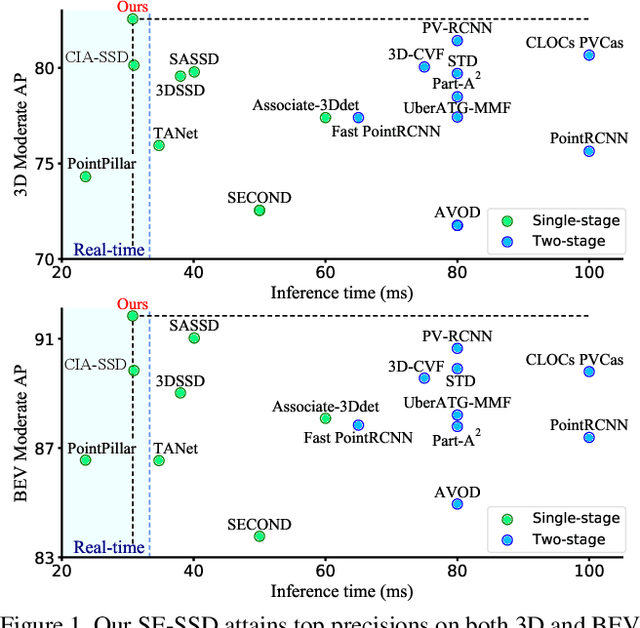
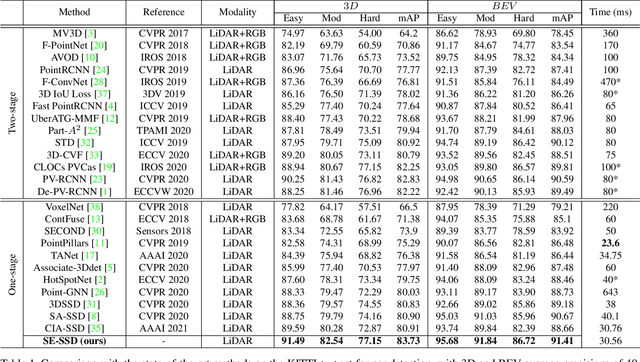
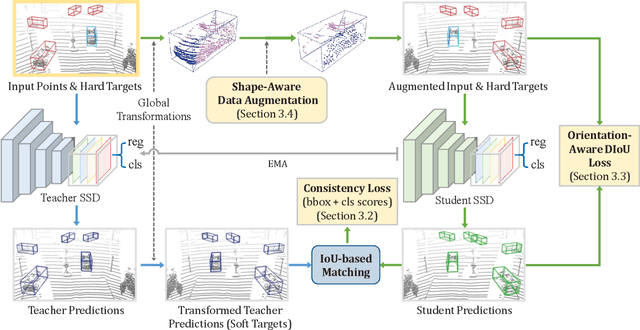
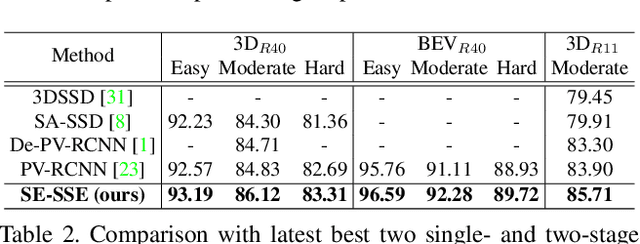
Abstract:We present Self-Ensembling Single-Stage object Detector (SE-SSD) for accurate and efficient 3D object detection in outdoor point clouds. Our key focus is on exploiting both soft and hard targets with our formulated constraints to jointly optimize the model, without introducing extra computation in the inference. Specifically, SE-SSD contains a pair of teacher and student SSDs, in which we design an effective IoU-based matching strategy to filter soft targets from the teacher and formulate a consistency loss to align student predictions with them. Also, to maximize the distilled knowledge for ensembling the teacher, we design a new augmentation scheme to produce shape-aware augmented samples to train the student, aiming to encourage it to infer complete object shapes. Lastly, to better exploit hard targets, we design an ODIoU loss to supervise the student with constraints on the predicted box centers and orientations. Our SE-SSD attains top performance compared with all prior published works. Also, it attains top precisions for car detection in the KITTI benchmark (ranked 1st and 2nd on the BEV and 3D leaderboards, respectively) with an ultra-high inference speed. The code is available at https://github.com/Vegeta2020/SE-SSD.
One Thing One Click: A Self-Training Approach for Weakly Supervised 3D Semantic Segmentation
Apr 16, 2021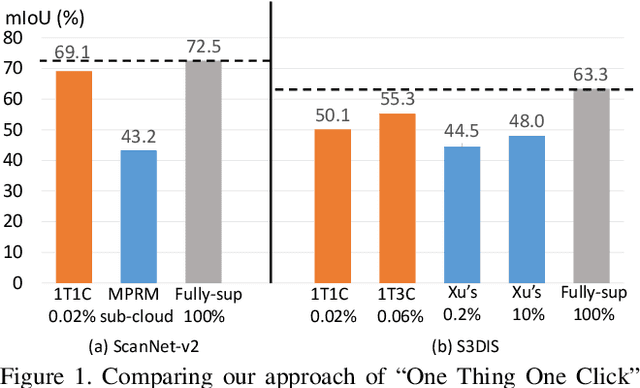
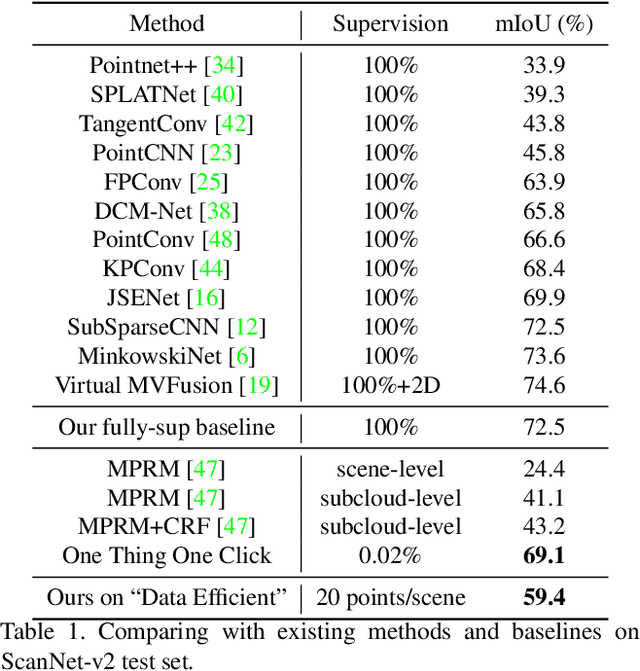
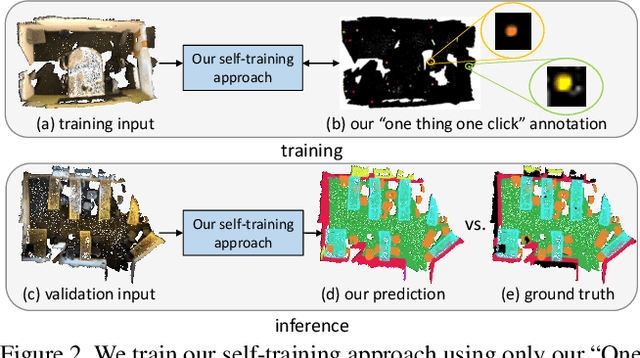
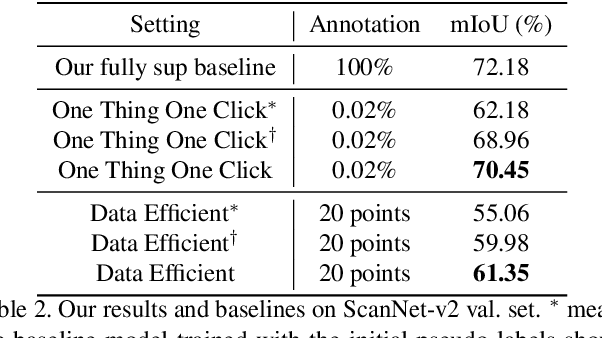
Abstract:Point cloud semantic segmentation often requires largescale annotated training data, but clearly, point-wise labels are too tedious to prepare. While some recent methods propose to train a 3D network with small percentages of point labels, we take the approach to an extreme and propose "One Thing One Click," meaning that the annotator only needs to label one point per object. To leverage these extremely sparse labels in network training, we design a novel self-training approach, in which we iteratively conduct the training and label propagation, facilitated by a graph propagation module. Also, we adopt a relation network to generate per-category prototype and explicitly model the similarity among graph nodes to generate pseudo labels to guide the iterative training. Experimental results on both ScanNet-v2 and S3DIS show that our self-training approach, with extremely-sparse annotations, outperforms all existing weakly supervised methods for 3D semantic segmentation by a large margin, and our results are also comparable to those of the fully supervised counterparts.
3D-to-2D Distillation for Indoor Scene Parsing
Apr 07, 2021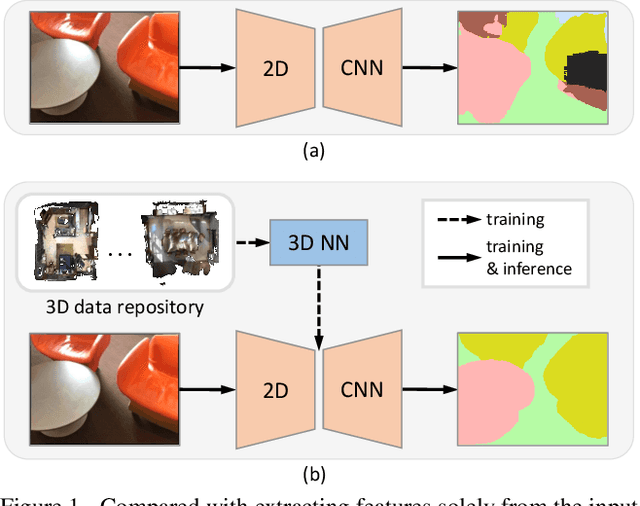

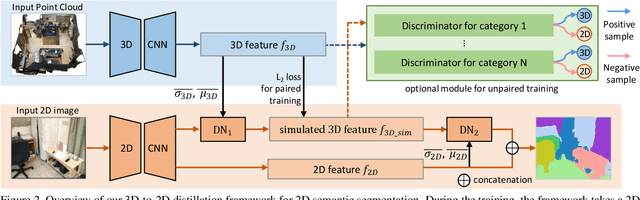
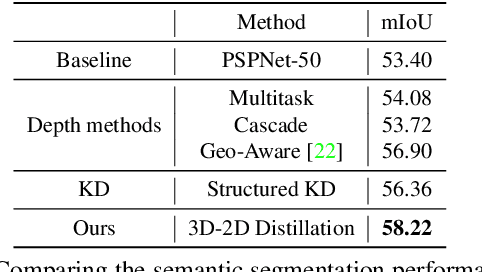
Abstract:Indoor scene semantic parsing from RGB images is very challenging due to occlusions, object distortion, and viewpoint variations. Going beyond prior works that leverage geometry information, typically paired depth maps, we present a new approach, a 3D-to-2D distillation framework, that enables us to leverage 3D features extracted from large-scale 3D data repository (e.g., ScanNet-v2) to enhance 2D features extracted from RGB images. Our work has three novel contributions. First, we distill 3D knowledge from a pretrained 3D network to supervise a 2D network to learn simulated 3D features from 2D features during the training, so the 2D network can infer without requiring 3D data. Second, we design a two-stage dimension normalization scheme to calibrate the 2D and 3D features for better integration. Third, we design a semantic-aware adversarial training model to extend our framework for training with unpaired 3D data. Extensive experiments on various datasets, ScanNet-V2, S3DIS, and NYU-v2, demonstrate the superiority of our approach. Also, experimental results show that our 3D-to-2D distillation improves the model generalization.
One Point is All You Need: Directional Attention Point for Feature Learning
Dec 14, 2020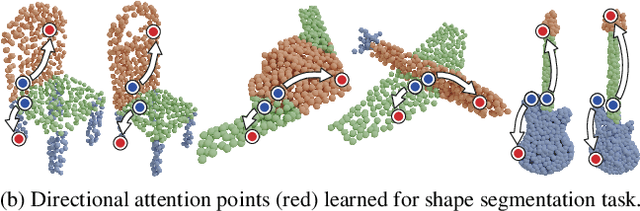
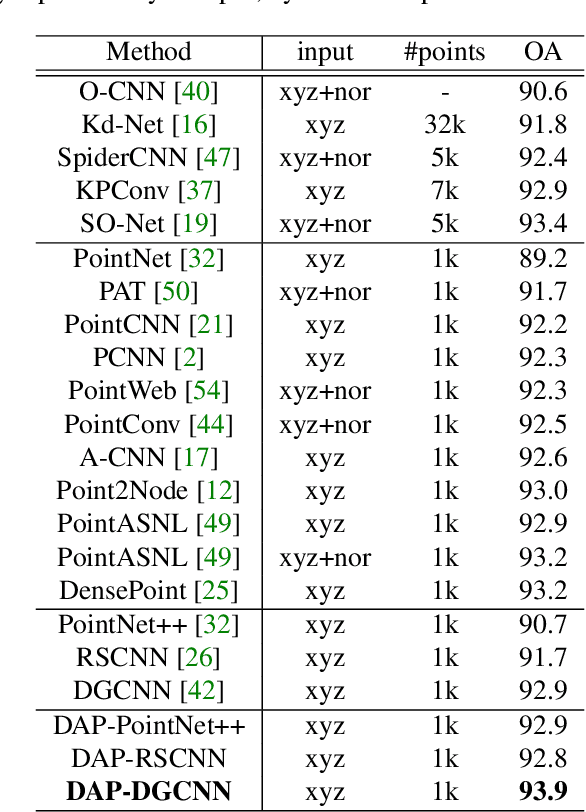
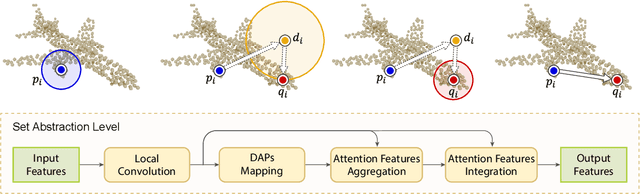
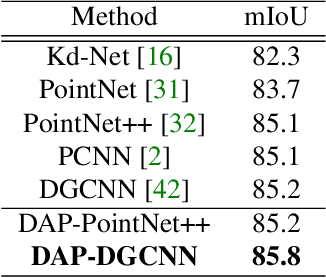
Abstract:We present a novel attention-based mechanism for learning enhanced point features for tasks such as point cloud classification and segmentation. Our key message is that if the right attention point is selected, then "one point is all you need" -- not a sequence as in a recurrent model and not a pre-selected set as in all prior works. Also, where the attention point is should be learned, from data and specific to the task at hand. Our mechanism is characterized by a new and simple convolution, which combines the feature at an input point with the feature at its associated attention point. We call such a point a directional attention point (DAP), since it is found by adding to the original point an offset vector that is learned by maximizing the task performance in training. We show that our attention mechanism can be easily incorporated into state-of-the-art point cloud classification and segmentation networks. Extensive experiments on common benchmarks such as ModelNet40, ShapeNetPart, and S3DIS demonstrate that our DAP-enabled networks consistently outperform the respective original networks, as well as all other competitive alternatives, including those employing pre-selected sets of attention points.
CIA-SSD: Confident IoU-Aware Single-Stage Object Detector From Point Cloud
Dec 05, 2020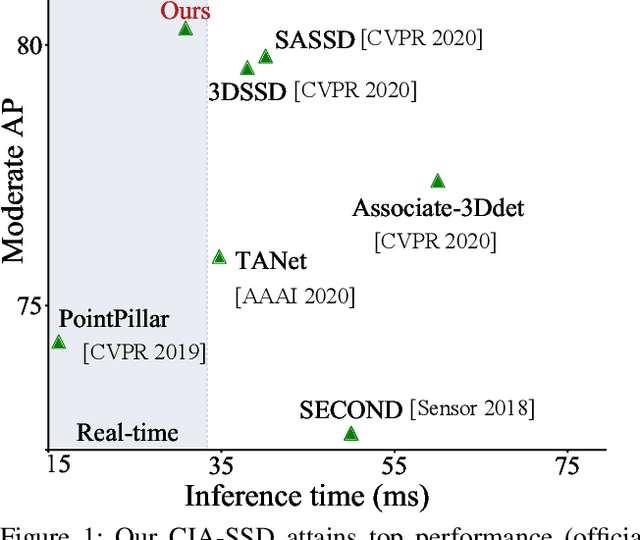
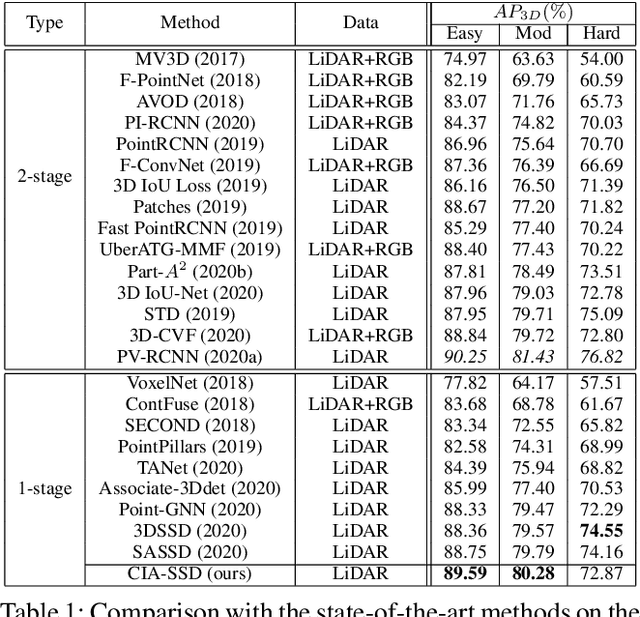
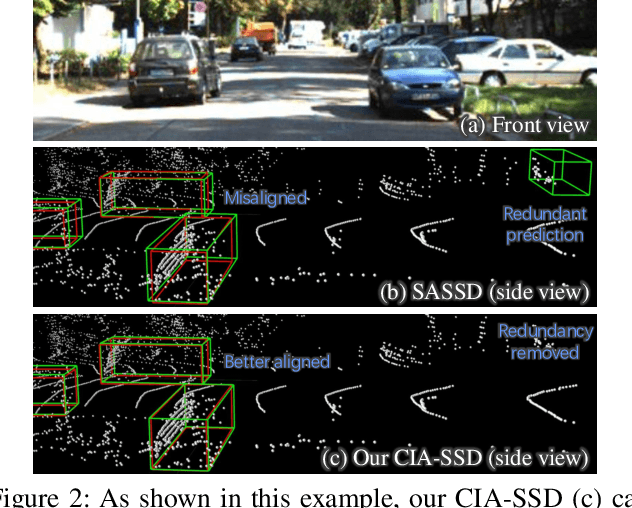
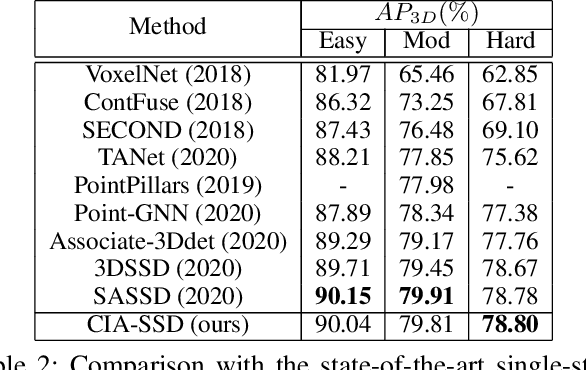
Abstract:Existing single-stage detectors for locating objects in point clouds often treat object localization and category classification as separate tasks, so the localization accuracy and classification confidence may not well align. To address this issue, we present a new single-stage detector named the Confident IoU-Aware Single-Stage object Detector (CIA-SSD). First, we design the lightweight Spatial-Semantic Feature Aggregation module to adaptively fuse high-level abstract semantic features and low-level spatial features for accurate predictions of bounding boxes and classification confidence. Also, the predicted confidence is further rectified with our designed IoU-aware confidence rectification module to make the confidence more consistent with the localization accuracy. Based on the rectified confidence, we further formulate the Distance-variant IoU-weighted NMS to obtain smoother regressions and avoid redundant predictions. We experiment CIA-SSD on 3D car detection in the KITTI test set and show that it attains top performance in terms of the official ranking metric (moderate AP 80.28%) and above 32 FPS inference speed, outperforming all prior single-stage detectors. The code is available at https://github.com/Vegeta2020/CIA-SSD.
DoFE: Domain-oriented Feature Embedding for Generalizable Fundus Image Segmentation on Unseen Datasets
Oct 13, 2020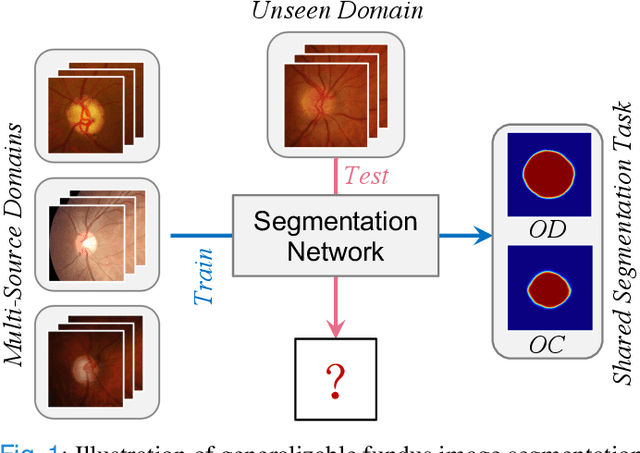
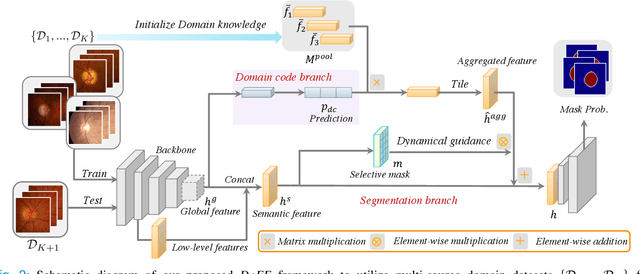
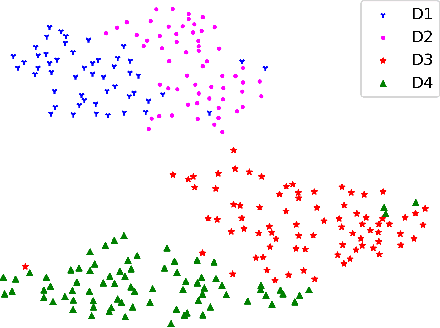
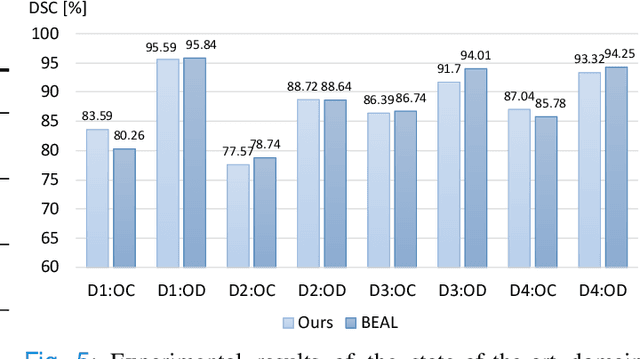
Abstract:Deep convolutional neural networks have significantly boosted the performance of fundus image segmentation when test datasets have the same distribution as the training datasets. However, in clinical practice, medical images often exhibit variations in appearance for various reasons, e.g., different scanner vendors and image quality. These distribution discrepancies could lead the deep networks to over-fit on the training datasets and lack generalization ability on the unseen test datasets. To alleviate this issue, we present a novel Domain-oriented Feature Embedding (DoFE) framework to improve the generalization ability of CNNs on unseen target domains by exploring the knowledge from multiple source domains. Our DoFE framework dynamically enriches the image features with additional domain prior knowledge learned from multi-source domains to make the semantic features more discriminative. Specifically, we introduce a Domain Knowledge Pool to learn and memorize the prior information extracted from multi-source domains. Then the original image features are augmented with domain-oriented aggregated features, which are induced from the knowledge pool based on the similarity between the input image and multi-source domain images. We further design a novel domain code prediction branch to infer this similarity and employ an attention-guided mechanism to dynamically combine the aggregated features with the semantic features. We comprehensively evaluate our DoFE framework on two fundus image segmentation tasks, including the optic cup and disc segmentation and vessel segmentation. Our DoFE framework generates satisfying segmentation results on unseen datasets and surpasses other domain generalization and network regularization methods.
Learning from Extrinsic and Intrinsic Supervisions for Domain Generalization
Jul 18, 2020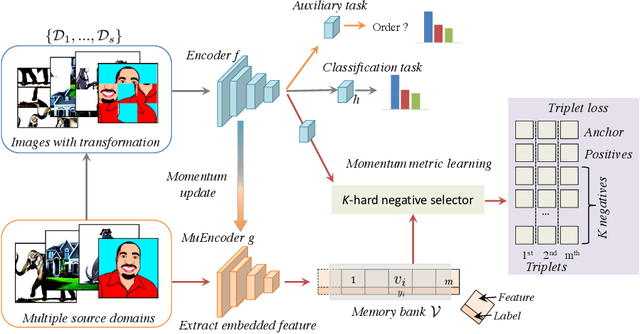
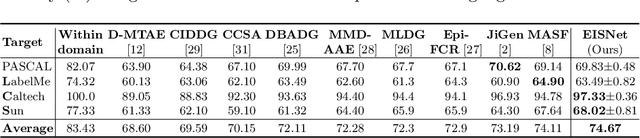
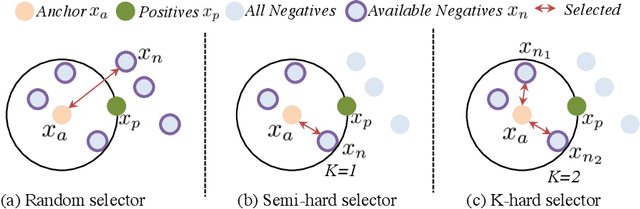
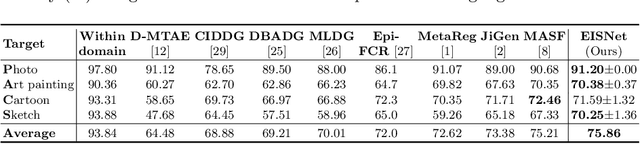
Abstract:The generalization capability of neural networks across domains is crucial for real-world applications. We argue that a generalized object recognition system should well understand the relationships among different images and also the images themselves at the same time. To this end, we present a new domain generalization framework that learns how to generalize across domains simultaneously from extrinsic relationship supervision and intrinsic self-supervision for images from multi-source domains. To be specific, we formulate our framework with feature embedding using a multi-task learning paradigm. Besides conducting the common supervised recognition task, we seamlessly integrate a momentum metric learning task and a self-supervised auxiliary task to collectively utilize the extrinsic supervision and intrinsic supervision. Also, we develop an effective momentum metric learning scheme with K-hard negative mining to boost the network to capture image relationship for domain generalization. We demonstrate the effectiveness of our approach on two standard object recognition benchmarks VLCS and PACS, and show that our methods achieve state-of-the-art performance.
TilinGNN: Learning to Tile with Self-Supervised Graph Neural Network
Jul 05, 2020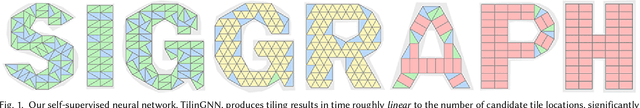
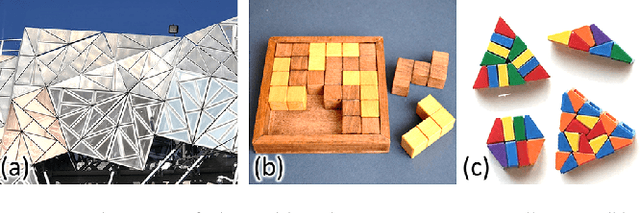
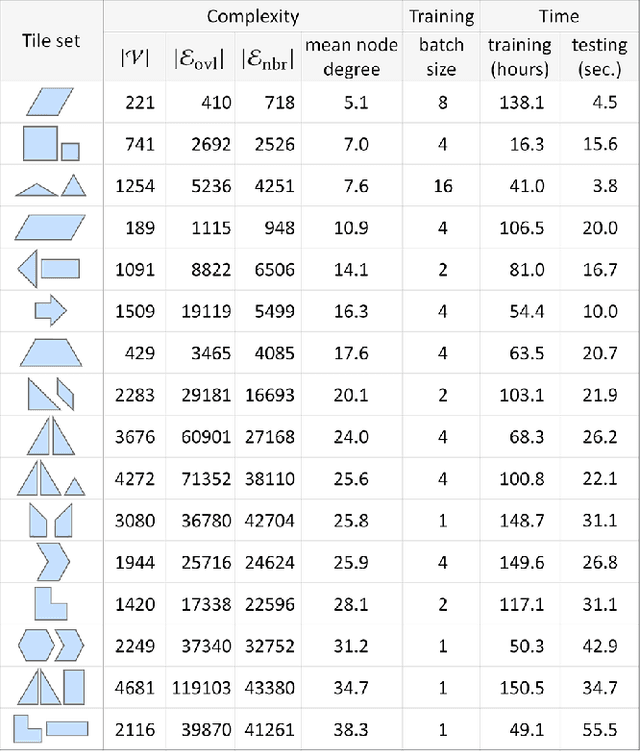

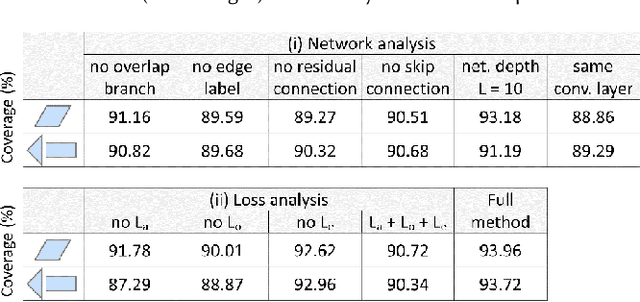
Abstract:We introduce the first neural optimization framework to solve a classical instance of the tiling problem. Namely, we seek a non-periodic tiling of an arbitrary 2D shape using one or more types of tiles: the tiles maximally fill the shape's interior without overlaps or holes. To start, we reformulate tiling as a graph problem by modeling candidate tile locations in the target shape as graph nodes and connectivity between tile locations as edges. Further, we build a graph convolutional neural network, coined TilinGNN, to progressively propagate and aggregate features over graph edges and predict tile placements. TilinGNN is trained by maximizing the tiling coverage on target shapes, while avoiding overlaps and holes between the tiles. Importantly, our network is self-supervised, as we articulate these criteria as loss terms defined on the network outputs, without the need of ground-truth tiling solutions. After training, the runtime of TilinGNN is roughly linear to the number of candidate tile locations, significantly outperforming traditional combinatorial search. We conducted various experiments on a variety of shapes to showcase the speed and versatility of TilinGNN. We also present comparisons to alternative methods and manual solutions, robustness analysis, and ablation studies to demonstrate the quality of our approach.
PointGroup: Dual-Set Point Grouping for 3D Instance Segmentation
Apr 03, 2020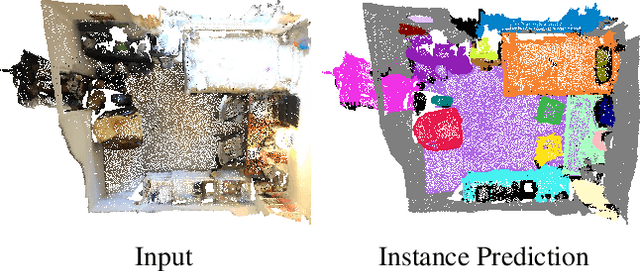
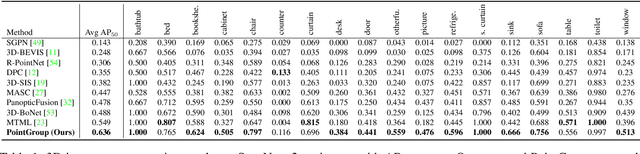
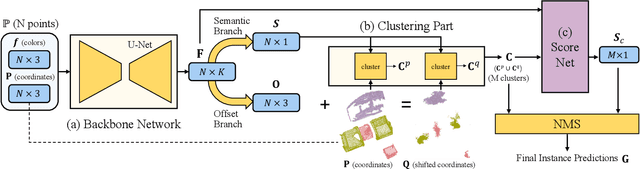
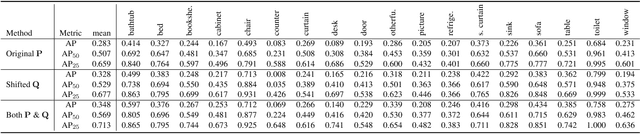
Abstract:Instance segmentation is an important task for scene understanding. Compared to the fully-developed 2D, 3D instance segmentation for point clouds have much room to improve. In this paper, we present PointGroup, a new end-to-end bottom-up architecture, specifically focused on better grouping the points by exploring the void space between objects. We design a two-branch network to extract point features and predict semantic labels and offsets, for shifting each point towards its respective instance centroid. A clustering component is followed to utilize both the original and offset-shifted point coordinate sets, taking advantage of their complementary strength. Further, we formulate the ScoreNet to evaluate the candidate instances, followed by the Non-Maximum Suppression (NMS) to remove duplicates. We conduct extensive experiments on two challenging datasets, ScanNet v2 and S3DIS, on which our method achieves the highest performance, 63.6% and 64.0%, compared to 54.9% and 54.4% achieved by former best solutions in terms of mAP with IoU threshold 0.5.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge