Chaowei Fang
Mitigating Undisciplined Over-Smoothing in Transformer for Weakly Supervised Semantic Segmentation
May 04, 2023Abstract:A surge of interest has emerged in weakly supervised semantic segmentation due to its remarkable efficiency in recent years. Existing approaches based on transformers mainly focus on exploring the affinity matrix to boost CAMs with global relationships. While in this work, we first perform a scrupulous examination towards the impact of successive affinity matrices and discover that they possess an inclination toward sparsification as the network approaches convergence, hence disclosing a manifestation of over-smoothing. Besides, it has been observed that enhanced attention maps tend to evince a substantial amount of extraneous background noise in deeper layers. Drawing upon this, we posit a daring conjecture that the undisciplined over-smoothing phenomenon introduces a noteworthy quantity of semantically irrelevant background noise, causing performance degradation. To alleviate this issue, we propose a novel perspective that highlights the objects of interest by investigating the regions of the trait, thereby fostering an extensive comprehension of the successive affinity matrix. Consequently, we suggest an adaptive re-activation mechanism (AReAM) that alleviates the issue of incomplete attention within the object and the unbounded background noise. AReAM accomplishes this by supervising high-level attention with shallow affinity matrices, yielding promising results. Exhaustive experiments conducted on the commonly used dataset manifest that segmentation results can be greatly improved through our proposed AReAM, which imposes restrictions on each affinity matrix in deep layers to make it attentive to semantic regions.
Revisiting Long-tailed Image Classification: Survey and Benchmarks with New Evaluation Metrics
Feb 03, 2023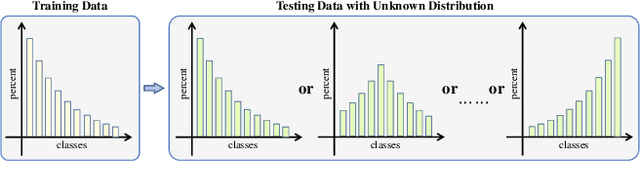
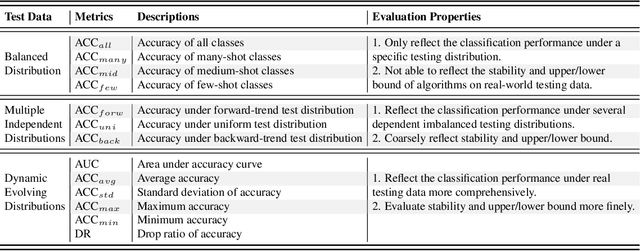
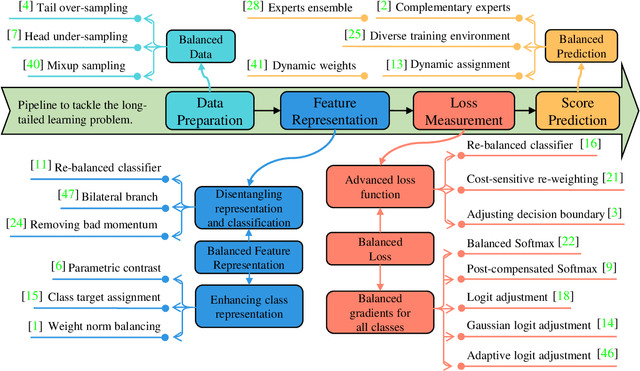
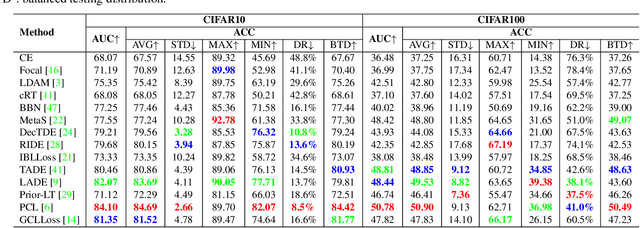
Abstract:Recently, long-tailed image classification harvests lots of research attention, since the data distribution is long-tailed in many real-world situations. Piles of algorithms are devised to address the data imbalance problem by biasing the training process towards less frequent classes. However, they usually evaluate the performance on a balanced testing set or multiple independent testing sets having distinct distributions with the training data. Considering the testing data may have arbitrary distributions, existing evaluation strategies are unable to reflect the actual classification performance objectively. We set up novel evaluation benchmarks based on a series of testing sets with evolving distributions. A corpus of metrics are designed for measuring the accuracy, robustness, and bounds of algorithms for learning with long-tailed distribution. Based on our benchmarks, we re-evaluate the performance of existing methods on CIFAR10 and CIFAR100 datasets, which is valuable for guiding the selection of data rebalancing techniques. We also revisit existing methods and categorize them into four types including data balancing, feature balancing, loss balancing, and prediction balancing, according the focused procedure during the training pipeline.
Compound Batch Normalization for Long-tailed Image Classification
Dec 02, 2022



Abstract:Significant progress has been made in learning image classification neural networks under long-tail data distribution using robust training algorithms such as data re-sampling, re-weighting, and margin adjustment. Those methods, however, ignore the impact of data imbalance on feature normalization. The dominance of majority classes (head classes) in estimating statistics and affine parameters causes internal covariate shifts within less-frequent categories to be overlooked. To alleviate this challenge, we propose a compound batch normalization method based on a Gaussian mixture. It can model the feature space more comprehensively and reduce the dominance of head classes. In addition, a moving average-based expectation maximization (EM) algorithm is employed to estimate the statistical parameters of multiple Gaussian distributions. However, the EM algorithm is sensitive to initialization and can easily become stuck in local minima where the multiple Gaussian components continue to focus on majority classes. To tackle this issue, we developed a dual-path learning framework that employs class-aware split feature normalization to diversify the estimated Gaussian distributions, allowing the Gaussian components to fit with training samples of less-frequent classes more comprehensively. Extensive experiments on commonly used datasets demonstrated that the proposed method outperforms existing methods on long-tailed image classification.
Deep 3D Vessel Segmentation based on Cross Transformer Network
Aug 23, 2022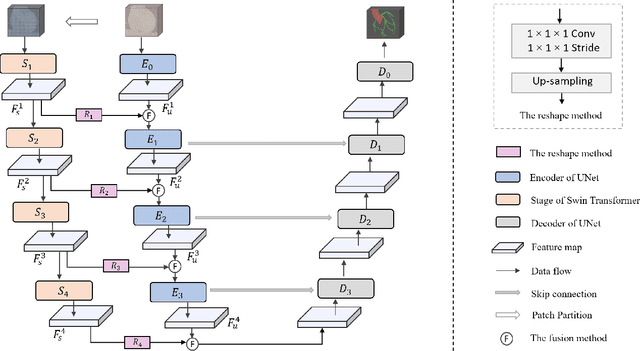
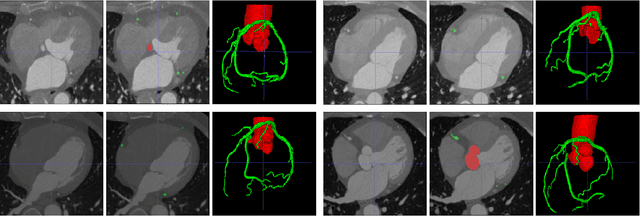
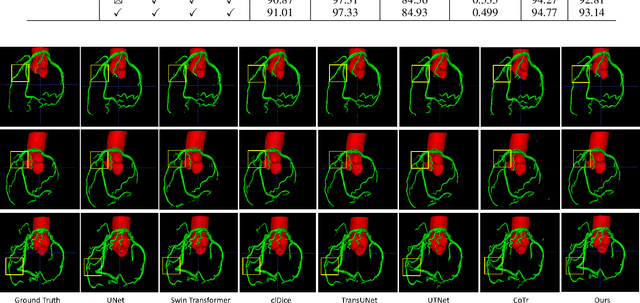
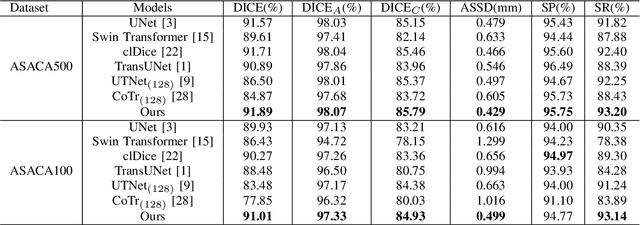
Abstract:The coronary microvascular disease poses a great threat to human health. Computer-aided analysis/diagnosis systems help physicians intervene in the disease at early stages, where 3D vessel segmentation is a fundamental step. However, there is a lack of carefully annotated dataset to support algorithm development and evaluation. On the other hand, the commonly-used U-Net structures often yield disconnected and inaccurate segmentation results, especially for small vessel structures. In this paper, motivated by the data scarcity, we first construct two large-scale vessel segmentation datasets consisting of 100 and 500 computed tomography (CT) volumes with pixel-level annotations by experienced radiologists. To enhance the U-Net, we further propose the cross transformer network (CTN) for fine-grained vessel segmentation. In CTN, a transformer module is constructed in parallel to a U-Net to learn long-distance dependencies between different anatomical regions; and these dependencies are communicated to the U-Net at multiple stages to endow it with global awareness. Experimental results on the two in-house datasets indicate that this hybrid model alleviates unexpected disconnections by considering topological information across regions. Our codes, together with the trained models are made publicly available at https://github.com/qibaolian/ctn.
Computer-aided Tuberculosis Diagnosis with Attribute Reasoning Assistance
Jul 01, 2022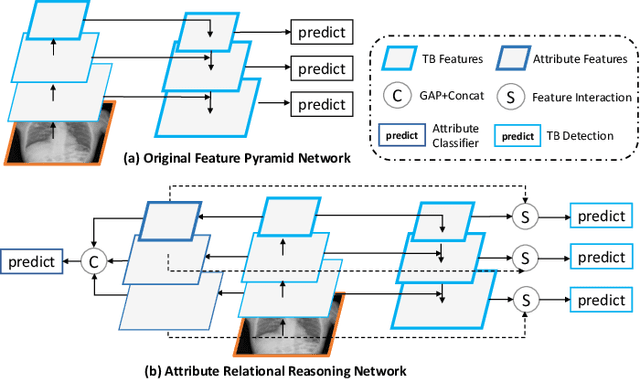
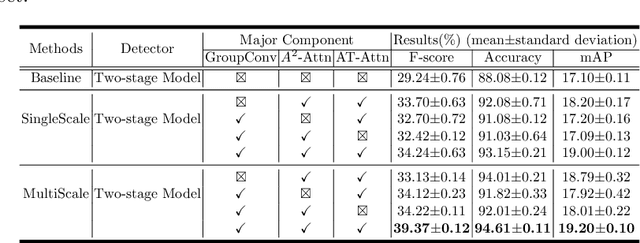
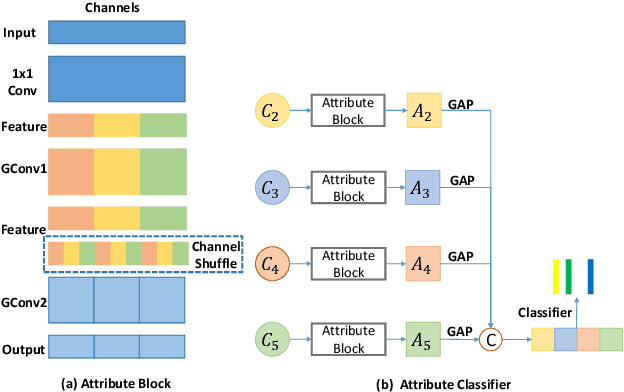
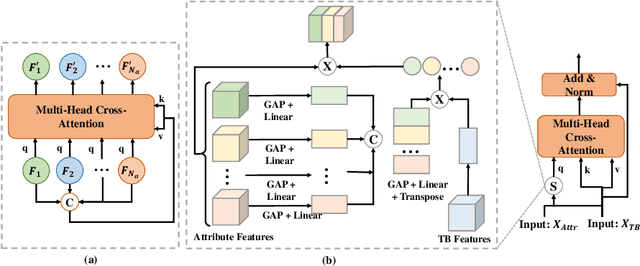
Abstract:Although deep learning algorithms have been intensively developed for computer-aided tuberculosis diagnosis (CTD), they mainly depend on carefully annotated datasets, leading to much time and resource consumption. Weakly supervised learning (WSL), which leverages coarse-grained labels to accomplish fine-grained tasks, has the potential to solve this problem. In this paper, we first propose a new large-scale tuberculosis (TB) chest X-ray dataset, namely the tuberculosis chest X-ray attribute dataset (TBX-Att), and then establish an attribute-assisted weakly-supervised framework to classify and localize TB by leveraging the attribute information to overcome the insufficiency of supervision in WSL scenarios. Specifically, first, the TBX-Att dataset contains 2000 X-ray images with seven kinds of attributes for TB relational reasoning, which are annotated by experienced radiologists. It also includes the public TBX11K dataset with 11200 X-ray images to facilitate weakly supervised detection. Second, we exploit a multi-scale feature interaction model for TB area classification and detection with attribute relational reasoning. The proposed model is evaluated on the TBX-Att dataset and will serve as a solid baseline for future research. The code and data will be available at https://github.com/GangmingZhao/tb-attribute-weak-localization.
Cross-Modality High-Frequency Transformer for MR Image Super-Resolution
Mar 29, 2022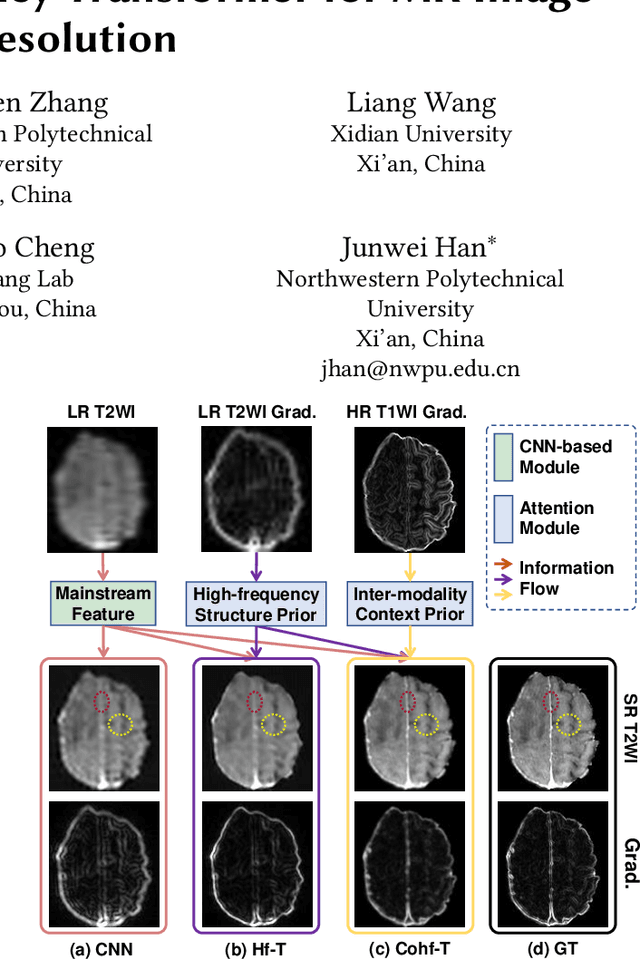
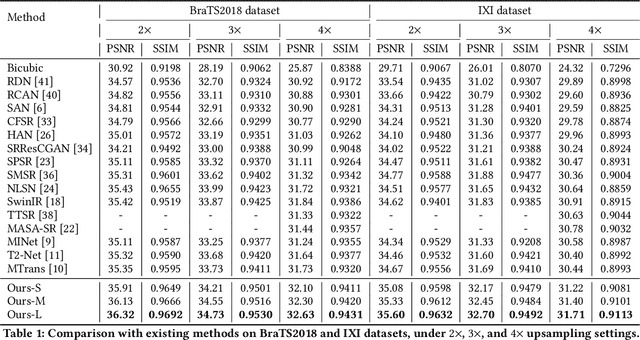
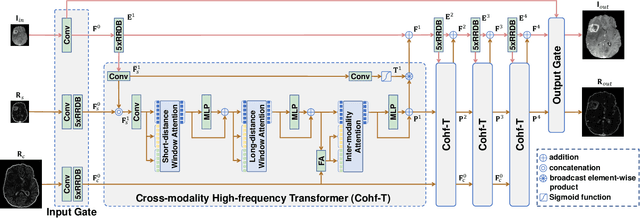
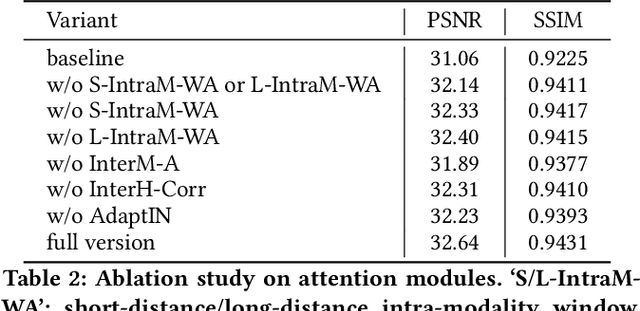
Abstract:Improving the resolution of magnetic resonance (MR) image data is critical to computer-aided diagnosis and brain function analysis. Higher resolution helps to capture more detailed content, but typically induces to lower signal-to-noise ratio and longer scanning time. To this end, MR image super-resolution has become a widely-interested topic in recent times. Existing works establish extensive deep models with the conventional architectures based on convolutional neural networks (CNN). In this work, to further advance this research field, we make an early effort to build a Transformer-based MR image super-resolution framework, with careful designs on exploring valuable domain prior knowledge. Specifically, we consider two-fold domain priors including the high-frequency structure prior and the inter-modality context prior, and establish a novel Transformer architecture, called Cross-modality high-frequency Transformer (Cohf-T), to introduce such priors into super-resolving the low-resolution (LR) MR images. Comprehensive experiments on two datasets indicate that Cohf-T achieves new state-of-the-art performance.
Cross-level Contrastive Learning and Consistency Constraint for Semi-supervised Medical Image Segmentation
Feb 13, 2022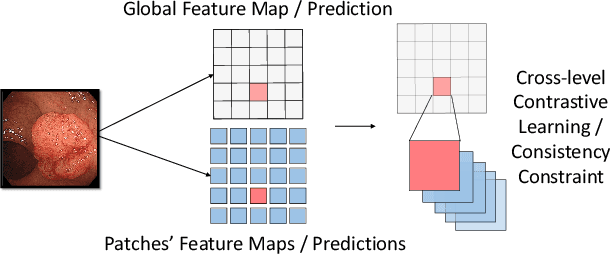
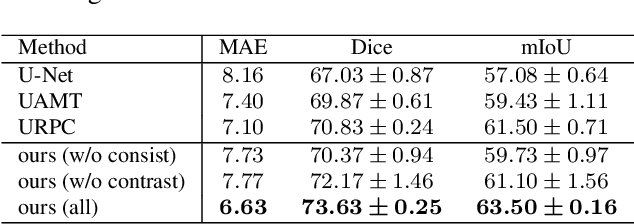
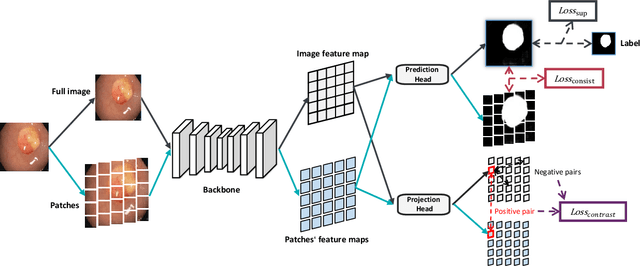
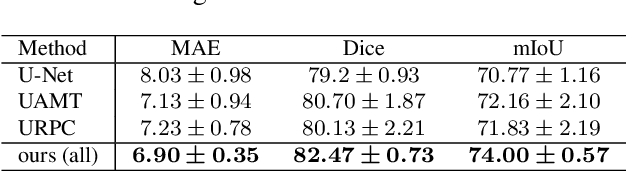
Abstract:Semi-supervised learning (SSL), which aims at leveraging a few labeled images and a large number of unlabeled images for network training, is beneficial for relieving the burden of data annotation in medical image segmentation. According to the experience of medical imaging experts, local attributes such as texture, luster and smoothness are very important factors for identifying target objects like lesions and polyps in medical images. Motivated by this, we propose a cross-level contrastive learning scheme to enhance representation capacity for local features in semi-supervised medical image segmentation. Compared to existing image-wise, patch-wise and point-wise contrastive learning algorithms, our devised method is capable of exploring more complex similarity cues, namely the relational characteristics between global and local patch-wise representations. Additionally, for fully making use of cross-level semantic relations, we devise a novel consistency constraint that compares the predictions of patches against those of the full image. With the help of the cross-level contrastive learning and consistency constraint, the unlabelled data can be effectively explored to improve segmentation performance on two medical image datasets for polyp and skin lesion segmentation respectively. Code of our approach is available.
Incremental Cross-view Mutual Distillation for Self-supervised Medical CT Synthesis
Dec 20, 2021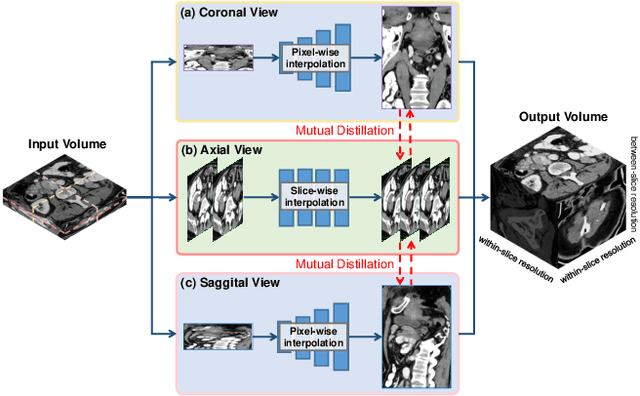
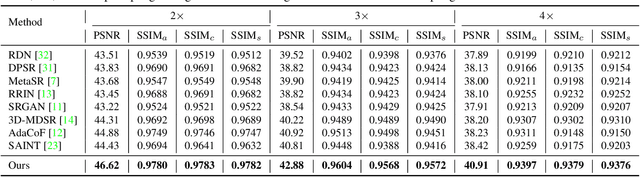
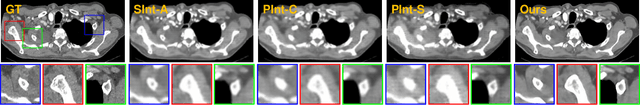
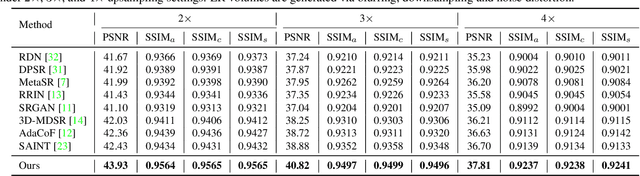
Abstract:Due to the constraints of the imaging device and high cost in operation time, computer tomography (CT) scans are usually acquired with low intra-slice resolution. Improving the intra-slice resolution is beneficial to the disease diagnosis for both human experts and computer-aided systems. To this end, this paper builds a novel medical slice synthesis to increase the between-slice resolution. Considering that the ground-truth intermediate medical slices are always absent in clinical practice, we introduce the incremental cross-view mutual distillation strategy to accomplish this task in the self-supervised learning manner. Specifically, we model this problem from three different views: slice-wise interpolation from axial view and pixel-wise interpolation from coronal and sagittal views. Under this circumstance, the models learned from different views can distill valuable knowledge to guide the learning processes of each other. We can repeat this process to make the models synthesize intermediate slice data with increasing inter-slice resolution. To demonstrate the effectiveness of the proposed approach, we conduct comprehensive experiments on a large-scale CT dataset. Quantitative and qualitative comparison results show that our method outperforms state-of-the-art algorithms by clear margins.
Weakly Supervised Semantic Segmentation via Alternative Self-Dual Teaching
Dec 17, 2021
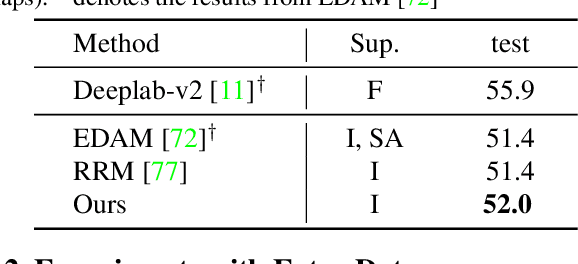
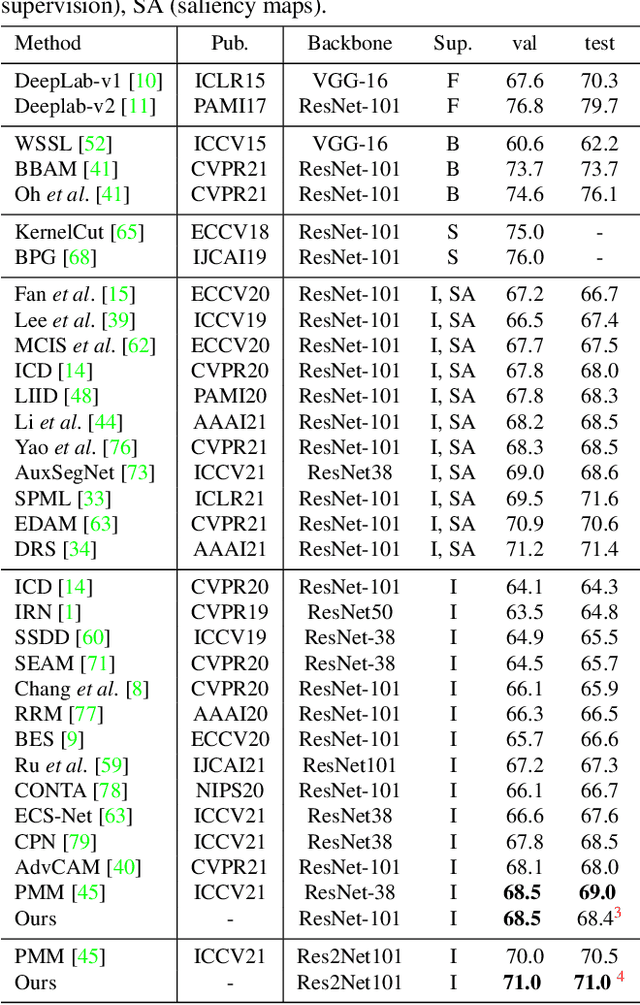
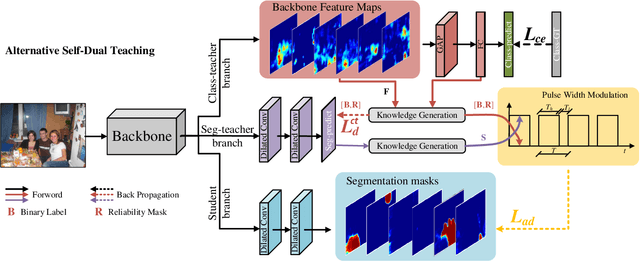

Abstract:Current weakly supervised semantic segmentation (WSSS) frameworks usually contain the separated mask-refinement model and the main semantic region mining model. These approaches would contain redundant feature extraction backbones and biased learning objectives, making them computational complex yet sub-optimal to addressing the WSSS task. To solve this problem, this paper establishes a compact learning framework that embeds the classification and mask-refinement components into a unified deep model. With the shared feature extraction backbone, our model is able to facilitate knowledge sharing between the two components while preserving a low computational complexity. To encourage high-quality knowledge interaction, we propose a novel alternative self-dual teaching (ASDT) mechanism. Unlike the conventional distillation strategy, the knowledge of the two teacher branches in our model is alternatively distilled to the student branch by a Pulse Width Modulation (PWM), which generates PW wave-like selection signal to guide the knowledge distillation process. In this way, the student branch can help prevent the model from falling into local minimum solutions caused by the imperfect knowledge provided of either teacher branch. Comprehensive experiments on the PASCAL VOC 2012 and COCO-Stuff 10K demonstrate the effectiveness of the proposed alternative self-dual teaching mechanism as well as the new state-of-the-art performance of our approach.
MVCNet: Multiview Contrastive Network for Unsupervised Representation Learning for 3D CT Lesions
Aug 18, 2021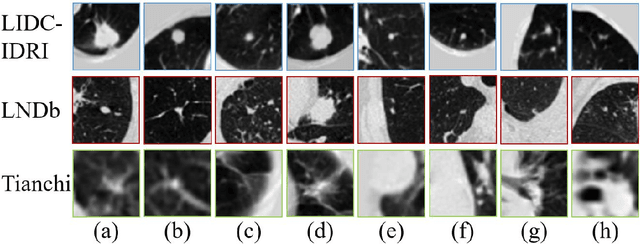
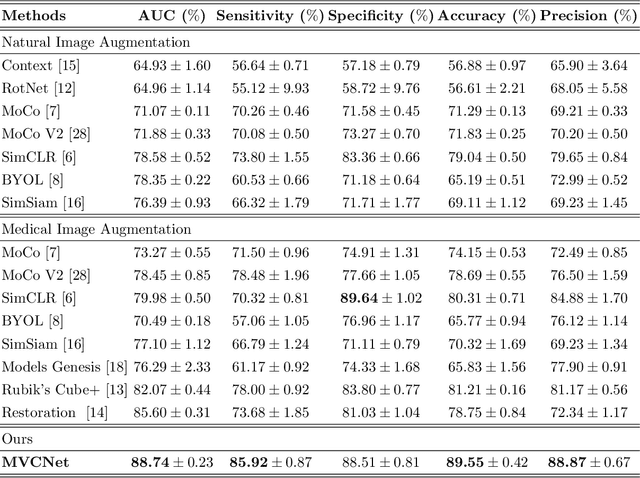
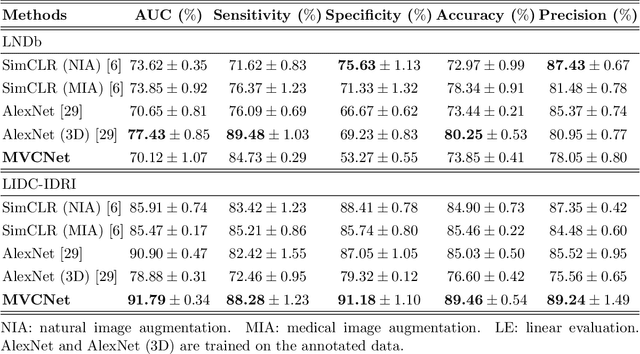
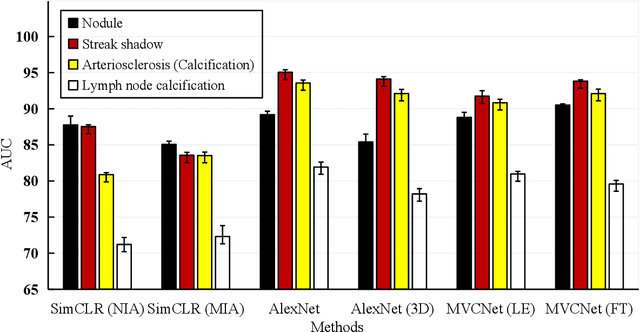
Abstract:\emph{Objective and Impact Statement}. With the renaissance of deep learning, automatic diagnostic systems for computed tomography (CT) have achieved many successful applications. However, they are mostly attributed to careful expert annotations, which are often scarce in practice. This drives our interest to the unsupervised representation learning. \emph{Introduction}. Recent studies have shown that self-supervised learning is an effective approach for learning representations, but most of them rely on the empirical design of transformations and pretext tasks. \emph{Methods}. To avoid the subjectivity associated with these methods, we propose the MVCNet, a novel unsupervised three dimensional (3D) representation learning method working in a transformation-free manner. We view each 3D lesion from different orientations to collect multiple two dimensional (2D) views. Then, an embedding function is learned by minimizing a contrastive loss so that the 2D views of the same 3D lesion are aggregated, and the 2D views of different lesions are separated. We evaluate the representations by training a simple classification head upon the embedding layer. \emph{Results}. Experimental results show that MVCNet achieves state-of-the-art accuracies on the LIDC-IDRI (89.55\%), LNDb (77.69\%) and TianChi (79.96\%) datasets for \emph{unsupervised representation learning}. When fine-tuned on 10\% of the labeled data, the accuracies are comparable to the supervised learning model (89.46\% vs. 85.03\%, 73.85\% vs. 73.44\%, 83.56\% vs. 83.34\% on the three datasets, respectively). \emph{Conclusion}. Results indicate the superiority of MVCNet in \emph{learning representations with limited annotations}.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge