Bin Zhang
OneRec-V2 Technical Report
Aug 28, 2025



Abstract:Recent breakthroughs in generative AI have transformed recommender systems through end-to-end generation. OneRec reformulates recommendation as an autoregressive generation task, achieving high Model FLOPs Utilization. While OneRec-V1 has shown significant empirical success in real-world deployment, two critical challenges hinder its scalability and performance: (1) inefficient computational allocation where 97.66% of resources are consumed by sequence encoding rather than generation, and (2) limitations in reinforcement learning relying solely on reward models. To address these challenges, we propose OneRec-V2, featuring: (1) Lazy Decoder-Only Architecture: Eliminates encoder bottlenecks, reducing total computation by 94% and training resources by 90%, enabling successful scaling to 8B parameters. (2) Preference Alignment with Real-World User Interactions: Incorporates Duration-Aware Reward Shaping and Adaptive Ratio Clipping to better align with user preferences using real-world feedback. Extensive A/B tests on Kuaishou demonstrate OneRec-V2's effectiveness, improving App Stay Time by 0.467%/0.741% while balancing multi-objective recommendations. This work advances generative recommendation scalability and alignment with real-world feedback, representing a step forward in the development of end-to-end recommender systems.
OneRec Technical Report
Jun 16, 2025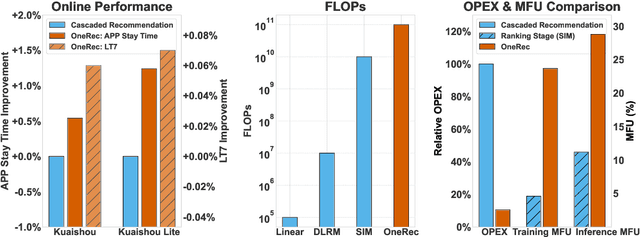

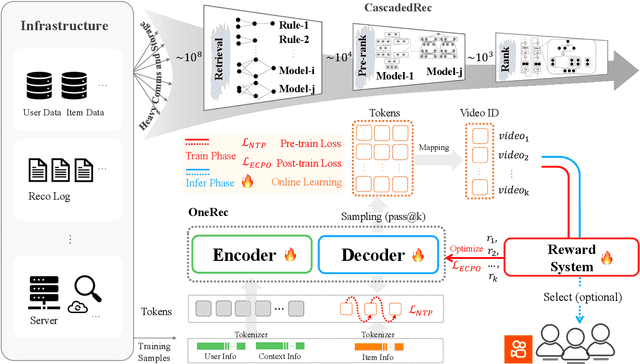
Abstract:Recommender systems have been widely used in various large-scale user-oriented platforms for many years. However, compared to the rapid developments in the AI community, recommendation systems have not achieved a breakthrough in recent years. For instance, they still rely on a multi-stage cascaded architecture rather than an end-to-end approach, leading to computational fragmentation and optimization inconsistencies, and hindering the effective application of key breakthrough technologies from the AI community in recommendation scenarios. To address these issues, we propose OneRec, which reshapes the recommendation system through an end-to-end generative approach and achieves promising results. Firstly, we have enhanced the computational FLOPs of the current recommendation model by 10 $\times$ and have identified the scaling laws for recommendations within certain boundaries. Secondly, reinforcement learning techniques, previously difficult to apply for optimizing recommendations, show significant potential in this framework. Lastly, through infrastructure optimizations, we have achieved 23.7% and 28.8% Model FLOPs Utilization (MFU) on flagship GPUs during training and inference, respectively, aligning closely with the LLM community. This architecture significantly reduces communication and storage overhead, resulting in operating expense that is only 10.6% of traditional recommendation pipelines. Deployed in Kuaishou/Kuaishou Lite APP, it handles 25% of total queries per second, enhancing overall App Stay Time by 0.54% and 1.24%, respectively. Additionally, we have observed significant increases in metrics such as 7-day Lifetime, which is a crucial indicator of recommendation experience. We also provide practical lessons and insights derived from developing, optimizing, and maintaining a production-scale recommendation system with significant real-world impact.
Graph of Verification: Structured Verification of LLM Reasoning with Directed Acyclic Graphs
Jun 14, 2025



Abstract:Verifying the reliability of complex, multi-step reasoning in Large Language Models (LLMs) remains a fundamental challenge, as existing methods often lack both faithfulness and precision. To address this issue, we propose the Graph of Verification (GoV) framework. GoV offers three key contributions: First, it explicitly models the underlying deductive process as a directed acyclic graph (DAG), whether this structure is implicit or explicitly constructed. Second, it enforces a topological order over the DAG to guide stepwise verification. Third, GoV introduces the notion of customizable node blocks, which flexibly define the verification granularity, from atomic propositions to full paragraphs, while ensuring that all requisite premises derived from the graph are provided as contextual input for each verification unit. We evaluate GoV on the Number Triangle Summation task and the ProcessBench benchmark with varying levels of reasoning complexity. Experimental results show that GoV substantially improves verification accuracy, faithfulness, and error localization when compared to conventional end-to-end verification approaches. Our code and data are available at https://github.com/Frevor/Graph-of-Verification.
MoQAE: Mixed-Precision Quantization for Long-Context LLM Inference via Mixture of Quantization-Aware Experts
Jun 09, 2025Abstract:One of the primary challenges in optimizing large language models (LLMs) for long-context inference lies in the high memory consumption of the Key-Value (KV) cache. Existing approaches, such as quantization, have demonstrated promising results in reducing memory usage. However, current quantization methods cannot take both effectiveness and efficiency into account. In this paper, we propose MoQAE, a novel mixed-precision quantization method via mixture of quantization-aware experts. First, we view different quantization bit-width configurations as experts and use the traditional mixture of experts (MoE) method to select the optimal configuration. To avoid the inefficiency caused by inputting tokens one by one into the router in the traditional MoE method, we input the tokens into the router chunk by chunk. Second, we design a lightweight router-only fine-tuning process to train MoQAE with a comprehensive loss to learn the trade-off between model accuracy and memory usage. Finally, we introduce a routing freezing (RF) and a routing sharing (RS) mechanism to further reduce the inference overhead. Extensive experiments on multiple benchmark datasets demonstrate that our method outperforms state-of-the-art KV cache quantization approaches in both efficiency and effectiveness.
ILIF: Temporal Inhibitory Leaky Integrate-and-Fire Neuron for Overactivation in Spiking Neural Networks
May 15, 2025



Abstract:The Spiking Neural Network (SNN) has drawn increasing attention for its energy-efficient, event-driven processing and biological plausibility. To train SNNs via backpropagation, surrogate gradients are used to approximate the non-differentiable spike function, but they only maintain nonzero derivatives within a narrow range of membrane potentials near the firing threshold, referred to as the surrogate gradient support width gamma. We identify a major challenge, termed the dilemma of gamma: a relatively large gamma leads to overactivation, characterized by excessive neuron firing, which in turn increases energy consumption, whereas a small gamma causes vanishing gradients and weakens temporal dependencies. To address this, we propose a temporal Inhibitory Leaky Integrate-and-Fire (ILIF) neuron model, inspired by biological inhibitory mechanisms. This model incorporates interconnected inhibitory units for membrane potential and current, effectively mitigating overactivation while preserving gradient propagation. Theoretical analysis demonstrates ILIF effectiveness in overcoming the gamma dilemma, and extensive experiments on multiple datasets show that ILIF improves energy efficiency by reducing firing rates, stabilizes training, and enhances accuracy. The code is available at github.com/kaisun1/ILIF.
CogniSNN: A First Exploration to Random Graph Architecture based Spiking Neural Networks with Enhanced Expandability and Neuroplasticity
May 09, 2025



Abstract:Despite advances in spiking neural networks (SNNs) in numerous tasks, their architectures remain highly similar to traditional artificial neural networks (ANNs), restricting their ability to mimic natural connections between biological neurons. This paper develops a new modeling paradigm for SNN with random graph architecture (RGA), termed Cognition-aware SNN (CogniSNN). Furthermore, we improve the expandability and neuroplasticity of CogniSNN by introducing a modified spiking residual neural node (ResNode) to counteract network degradation in deeper graph pathways, as well as a critical path-based algorithm that enables CogniSNN to perform continual learning on new tasks leveraging the features of the data and the RGA learned in the old task. Experiments show that CogniSNN with re-designed ResNode performs outstandingly in neuromorphic datasets with fewer parameters, achieving 95.5% precision in the DVS-Gesture dataset with only 5 timesteps. The critical path-based approach decreases 3% to 5% forgetting while maintaining expected performance in learning new tasks that are similar to or distinct from the old ones. This study showcases the potential of RGA-based SNN and paves a new path for biologically inspired networks based on graph theory.
QLLM: Do We Really Need a Mixing Network for Credit Assignment in Multi-Agent Reinforcement Learning?
Apr 17, 2025



Abstract:Credit assignment has remained a fundamental challenge in multi-agent reinforcement learning (MARL). Previous studies have primarily addressed this issue through value decomposition methods under the centralized training with decentralized execution paradigm, where neural networks are utilized to approximate the nonlinear relationship between individual Q-values and the global Q-value. Although these approaches have achieved considerable success in various benchmark tasks, they still suffer from several limitations, including imprecise attribution of contributions, limited interpretability, and poor scalability in high-dimensional state spaces. To address these challenges, we propose a novel algorithm, \textbf{QLLM}, which facilitates the automatic construction of credit assignment functions using large language models (LLMs). Specifically, the concept of \textbf{TFCAF} is introduced, wherein the credit allocation process is represented as a direct and expressive nonlinear functional formulation. A custom-designed \textit{coder-evaluator} framework is further employed to guide the generation, verification, and refinement of executable code by LLMs, significantly mitigating issues such as hallucination and shallow reasoning during inference. Extensive experiments conducted on several standard MARL benchmarks demonstrate that the proposed method consistently outperforms existing state-of-the-art baselines. Moreover, QLLM exhibits strong generalization capability and maintains compatibility with a wide range of MARL algorithms that utilize mixing networks, positioning it as a promising and versatile solution for complex multi-agent scenarios.
Cocktail: Chunk-Adaptive Mixed-Precision Quantization for Long-Context LLM Inference
Mar 30, 2025



Abstract:Recently, large language models (LLMs) have been able to handle longer and longer contexts. However, a context that is too long may cause intolerant inference latency and GPU memory usage. Existing methods propose mixed-precision quantization to the key-value (KV) cache in LLMs based on token granularity, which is time-consuming in the search process and hardware inefficient during computation. This paper introduces a novel approach called Cocktail, which employs chunk-adaptive mixed-precision quantization to optimize the KV cache. Cocktail consists of two modules: chunk-level quantization search and chunk-level KV cache computation. Chunk-level quantization search determines the optimal bitwidth configuration of the KV cache chunks quickly based on the similarity scores between the corresponding context chunks and the query, maintaining the model accuracy. Furthermore, chunk-level KV cache computation reorders the KV cache chunks before quantization, avoiding the hardware inefficiency caused by mixed-precision quantization in inference computation. Extensive experiments demonstrate that Cocktail outperforms state-of-the-art KV cache quantization methods on various models and datasets.
RUNA: Object-level Out-of-Distribution Detection via Regional Uncertainty Alignment of Multimodal Representations
Mar 28, 2025Abstract:Enabling object detectors to recognize out-of-distribution (OOD) objects is vital for building reliable systems. A primary obstacle stems from the fact that models frequently do not receive supervisory signals from unfamiliar data, leading to overly confident predictions regarding OOD objects. Despite previous progress that estimates OOD uncertainty based on the detection model and in-distribution (ID) samples, we explore using pre-trained vision-language representations for object-level OOD detection. We first discuss the limitations of applying image-level CLIP-based OOD detection methods to object-level scenarios. Building upon these insights, we propose RUNA, a novel framework that leverages a dual encoder architecture to capture rich contextual information and employs a regional uncertainty alignment mechanism to distinguish ID from OOD objects effectively. We introduce a few-shot fine-tuning approach that aligns region-level semantic representations to further improve the model's capability to discriminate between similar objects. Our experiments show that RUNA substantially surpasses state-of-the-art methods in object-level OOD detection, particularly in challenging scenarios with diverse and complex object instances.
VisTa: Visual-contextual and Text-augmented Zero-shot Object-level OOD Detection
Mar 28, 2025Abstract:As object detectors are increasingly deployed as black-box cloud services or pre-trained models with restricted access to the original training data, the challenge of zero-shot object-level out-of-distribution (OOD) detection arises. This task becomes crucial in ensuring the reliability of detectors in open-world settings. While existing methods have demonstrated success in image-level OOD detection using pre-trained vision-language models like CLIP, directly applying such models to object-level OOD detection presents challenges due to the loss of contextual information and reliance on image-level alignment. To tackle these challenges, we introduce a new method that leverages visual prompts and text-augmented in-distribution (ID) space construction to adapt CLIP for zero-shot object-level OOD detection. Our method preserves critical contextual information and improves the ability to differentiate between ID and OOD objects, achieving competitive performance across different benchmarks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge