"Image": models, code, and papers
Uncertainty driven probabilistic voxel selection for image registration
Oct 02, 2020
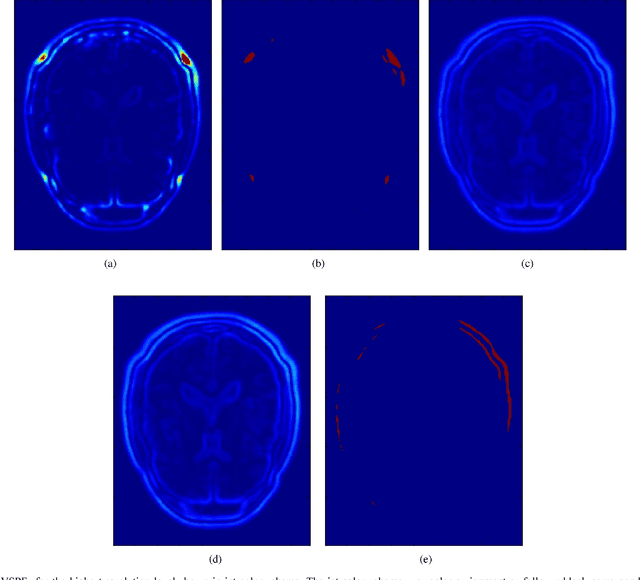
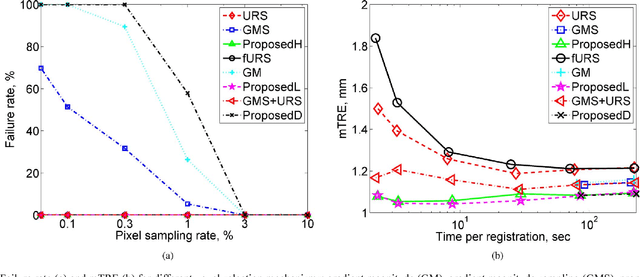
This paper presents a novel probabilistic voxel selection strategy for medical image registration in time-sensitive contexts, where the goal is aggressive voxel sampling (e.g. using less than 1% of the total number) while maintaining registration accuracy and low failure rate. We develop a Bayesian framework whereby, first, a voxel sampling probability field (VSPF) is built based on the uncertainty on the transformation parameters. We then describe a practical, multi-scale registration algorithm, where, at each optimization iteration, different voxel subsets are sampled based on the VSPF. The approach maximizes accuracy without committing to a particular fixed subset of voxels. The probabilistic sampling scheme developed is shown to manage the tradeoff between the robustness of traditional random voxel selection (by permitting more exploration) and the accuracy of fixed voxel selection (by permitting a greater proportion of informative voxels).
Disentangled Image Generation Through Structured Noise Injection
May 05, 2020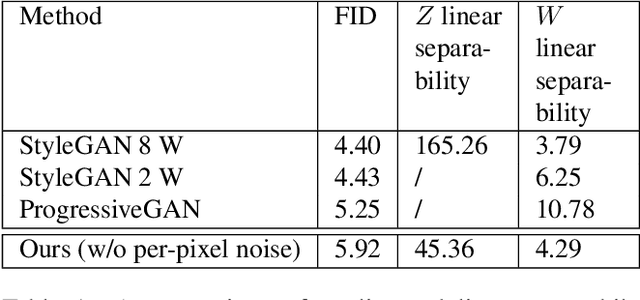
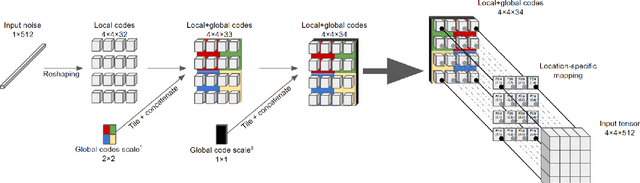
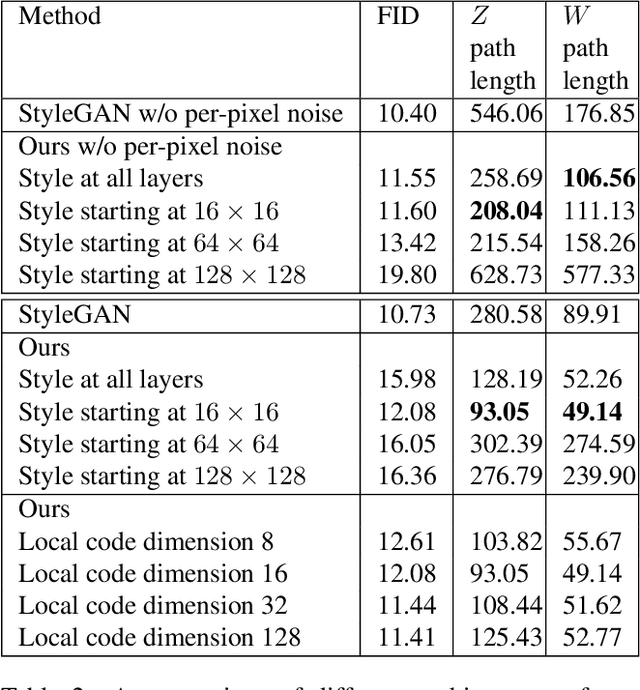
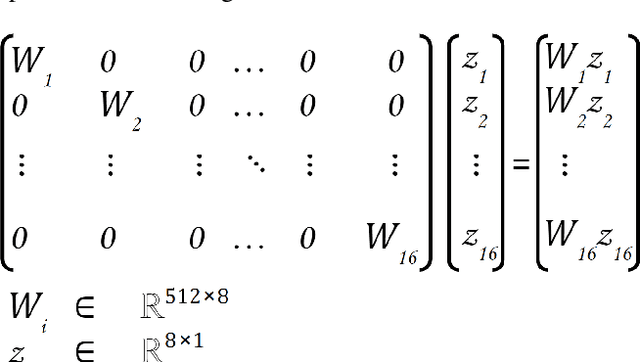
We explore different design choices for injecting noise into generative adversarial networks (GANs) with the goal of disentangling the latent space. Instead of traditional approaches, we propose feeding multiple noise codes through separate fully-connected layers respectively. The aim is restricting the influence of each noise code to specific parts of the generated image. We show that disentanglement in the first layer of the generator network leads to disentanglement in the generated image. Through a grid-based structure, we achieve several aspects of disentanglement without complicating the network architecture and without requiring labels. We achieve spatial disentanglement, scale-space disentanglement, and disentanglement of the foreground object from the background style allowing fine-grained control over the generated images. Examples include changing facial expressions in face images, changing beak length in bird images, and changing car dimensions in car images. This empirically leads to better disentanglement scores than state-of-the-art methods on the FFHQ dataset.
Diverse and Styled Image Captioning Using SVD-Based Mixture of Recurrent Experts
Jul 07, 2020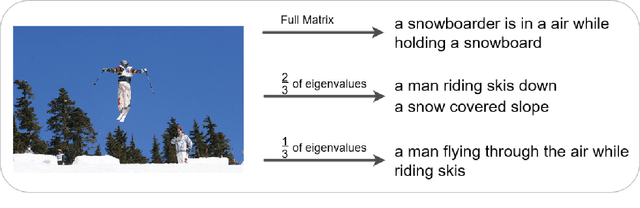
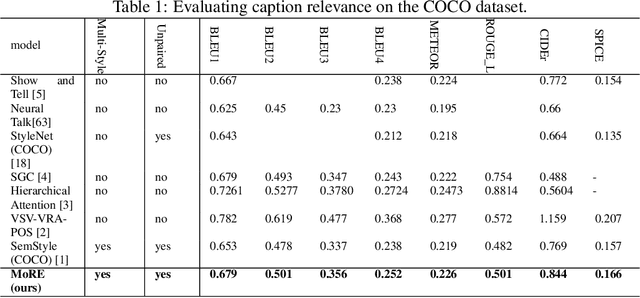
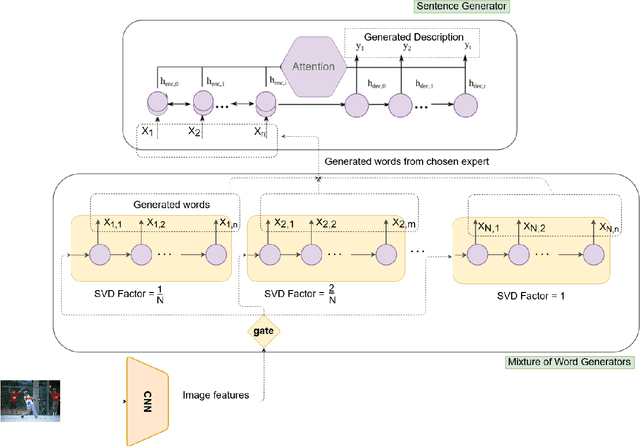

With great advances in vision and natural language processing, the generation of image captions becomes a need. In a recent paper, Mathews, Xie and He [1], extended a new model to generate styled captions by separating semantics and style. In continuation of this work, here a new captioning model is developed including an image encoder to extract the features, a mixture of recurrent networks to embed the set of extracted features to a set of words, and a sentence generator that combines the obtained words as a stylized sentence. The resulted system that entitled as Mixture of Recurrent Experts (MoRE), uses a new training algorithm that derives singular value decomposition (SVD) from weighting matrices of Recurrent Neural Networks (RNNs) to increase the diversity of captions. Each decomposition step depends on a distinctive factor based on the number of RNNs in MoRE. Since the used sentence generator gives a stylized language corpus without paired images, our captioning model can do the same. Besides, the styled and diverse captions are extracted without training on a densely labeled or styled dataset. To validate this captioning model, we use Microsoft COCO which is a standard factual image caption dataset. We show that the proposed captioning model can generate a diverse and stylized image captions without the necessity of extra-labeling. The results also show better descriptions in terms of content accuracy.
Quality-aware Cine Cardiac MRI Reconstruction and Analysis from Undersampled k-space Data
Sep 16, 2021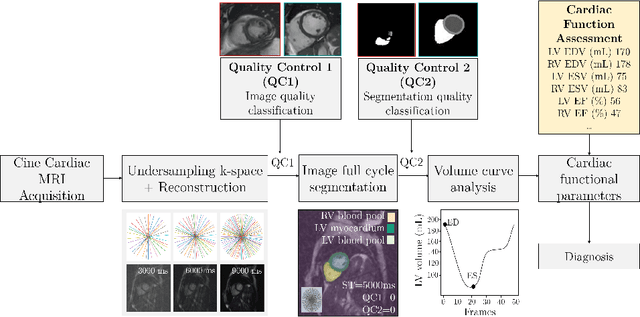
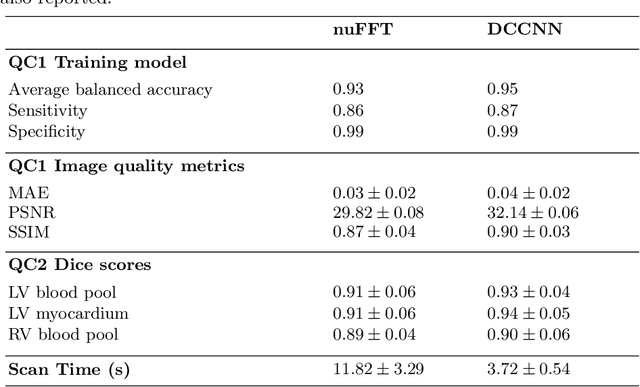
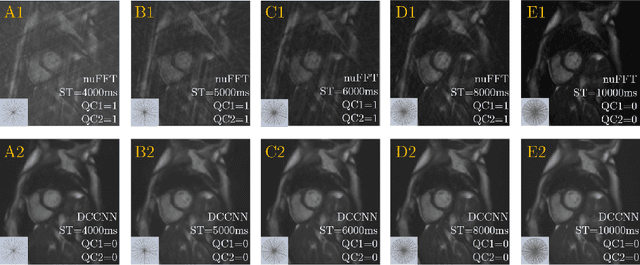
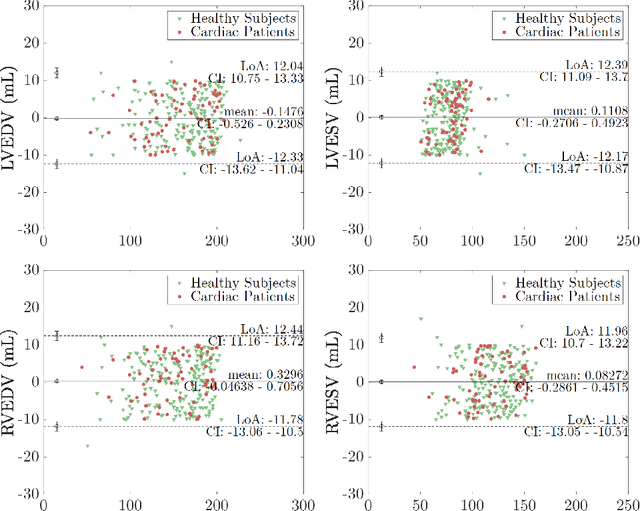
Cine cardiac MRI is routinely acquired for the assessment of cardiac health, but the imaging process is slow and typically requires several breath-holds to acquire sufficient k-space profiles to ensure good image quality. Several undersampling-based reconstruction techniques have been proposed during the last decades to speed up cine cardiac MRI acquisition. However, the undersampling factor is commonly fixed to conservative values before acquisition to ensure diagnostic image quality, potentially leading to unnecessarily long scan times. In this paper, we propose an end-to-end quality-aware cine short-axis cardiac MRI framework that combines image acquisition and reconstruction with downstream tasks such as segmentation, volume curve analysis and estimation of cardiac functional parameters. The goal is to reduce scan time by acquiring only a fraction of k-space data to enable the reconstruction of images that can pass quality control checks and produce reliable estimates of cardiac functional parameters. The framework consists of a deep learning model for the reconstruction of 2D+t cardiac cine MRI images from undersampled data, an image quality-control step to detect good quality reconstructions, followed by a deep learning model for bi-ventricular segmentation, a quality-control step to detect good quality segmentations and automated calculation of cardiac functional parameters. To demonstrate the feasibility of the proposed approach, we perform simulations using a cohort of selected participants from the UK Biobank (n=270), 200 healthy subjects and 70 patients with cardiomyopathies. Our results show that we can produce quality-controlled images in a scan time reduced from 12 to 4 seconds per slice, enabling reliable estimates of cardiac functional parameters such as ejection fraction within 5% mean absolute error.
A deep convolutional neural network for classification of Aedes albopictus mosquitoes
Oct 29, 2021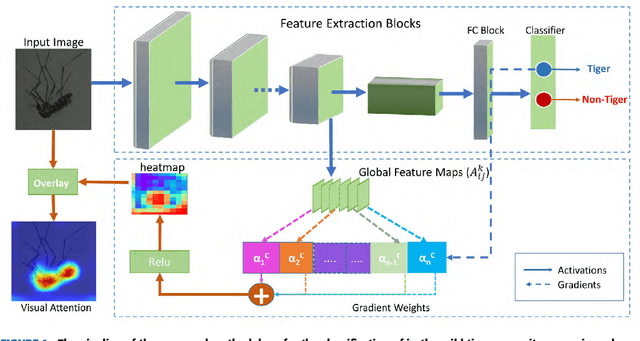
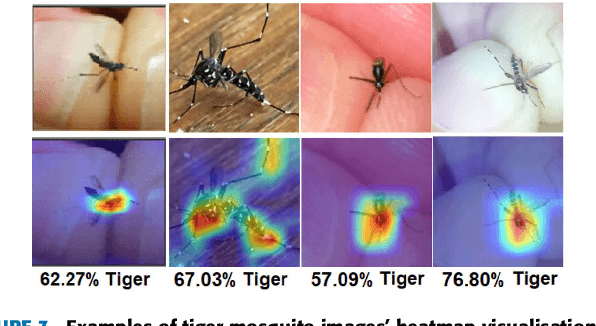
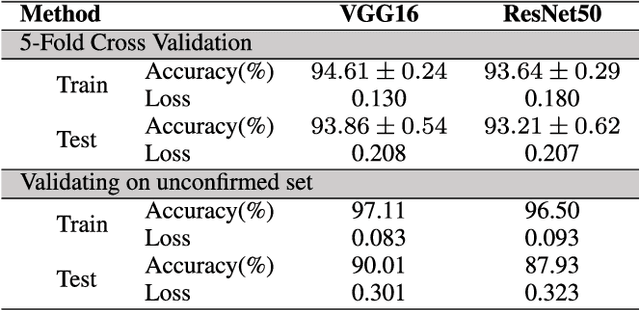
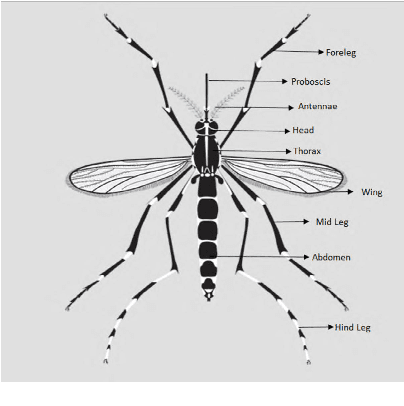

Monitoring the spread of disease-carrying mosquitoes is a first and necessary step to control severe diseases such as dengue, chikungunya, Zika or yellow fever. Previous citizen science projects have been able to obtain large image datasets with linked geo-tracking information. As the number of international collaborators grows, the manual annotation by expert entomologists of the large amount of data gathered by these users becomes too time demanding and unscalable, posing a strong need for automated classification of mosquito species from images. We introduce the application of two Deep Convolutional Neural Networks in a comparative study to automate this classification task. We use the transfer learning principle to train two state-of-the-art architectures on the data provided by the Mosquito Alert project, obtaining testing accuracy of 94%. In addition, we applied explainable models based on the Grad-CAM algorithm to visualise the most discriminant regions of the classified images, which coincide with the white band stripes located at the legs, abdomen, and thorax of mosquitoes of the Aedes albopictus species. The model allows us to further analyse the classification errors. Visual Grad-CAM models show that they are linked to poor acquisition conditions and strong image occlusions.
Bounding Boxes Are All We Need: Street View Image Classification via Context Encoding of Detected Buildings
Oct 03, 2020
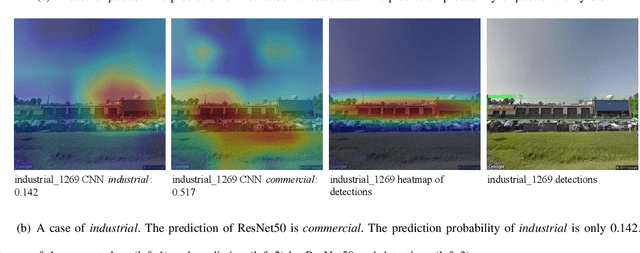
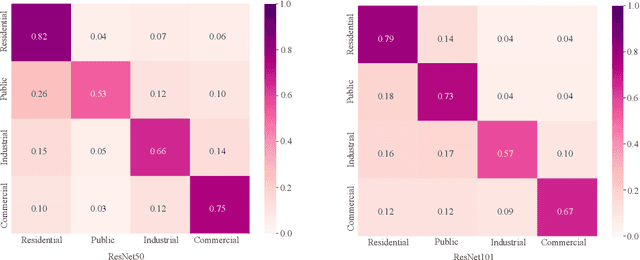
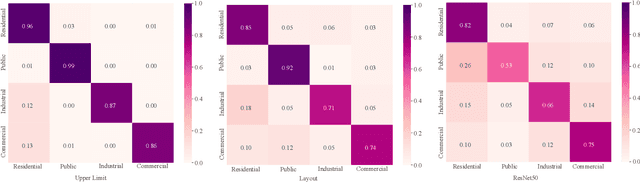

Street view images have been increasingly used in tasks like urban land use classification and urban functional zone portraying. Street view image classification is difficult because the class labels such as commercial area, are concepts with higher abstract level compared to general visual tasks. Therefore, classification models using only visual features often fail to achieve satisfactory performance. We believe that the efficient representation of significant objects and their context relations in street view images are the keys to solve this problem. In this paper, a novel approach based on a detector-encoder-classifier framework is proposed. Different from common image-level end-to-end models, our approach does not use visual features of the whole image directly. The proposed framework obtains the bounding boxes of buildings in street view images from a detector. Their contextual information such as building classes and positions are then encoded into metadata and finally classified by a recurrent neural network (RNN). To verify our approach, we made a dataset of 19,070 street view images and 38,857 buildings based on the BIC_GSV dataset through a combination of automatic label acquisition and expert annotation. The dataset can be used not only for street view image classification aiming at urban land use analysis, but also for multi-class building detection. Experiments show that the proposed approach achieves a 12.65% performance improvement on macro-precision and 12% on macro-recall over the models based on end-to-end convolutional neural network (CNN). Our code and dataset are available at https://github.com/kyle-one/Context-Encoding-of-Detected-Buildings/
UNITER: Learning UNiversal Image-TExt Representations
Sep 25, 2019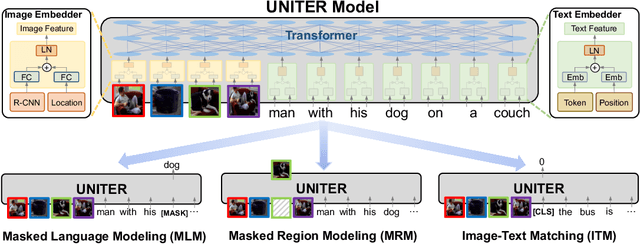

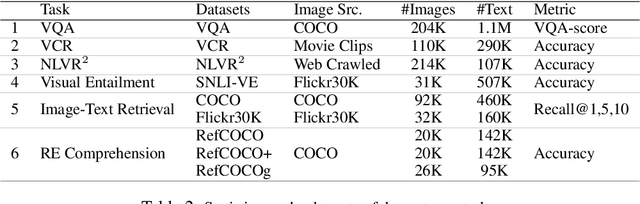
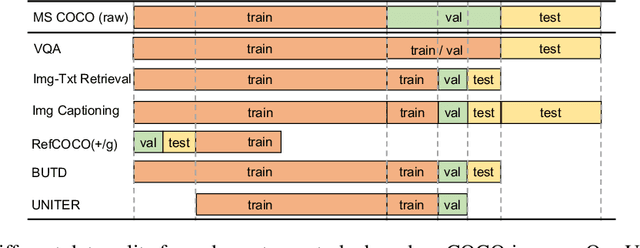
Joint image-text embedding is the bedrock for most Vision-and-Language (V+L) tasks, where multimodality inputs are jointly processed for visual and textual understanding. In this paper, we introduce UNITER, a UNiversal Image-TExt Representation, learned through large-scale pre-training over four image-text datasets (COCO, Visual Genome, Conceptual Captions, and SBU Captions), which can power heterogeneous downstream V+L tasks with joint multimodal embeddings. We design three pre-training tasks: Masked Language Modeling (MLM), Image-Text Matching (ITM), and Masked Region Modeling (MRM, with three variants). Different from concurrent work on multimodal pre-training that apply joint random masking to both modalities, we use conditioned masking on pre-training tasks (i.e., masked language/region modeling is conditioned on full observation of image/text). Comprehensive analysis shows that conditioned masking yields better performance than unconditioned masking. We also conduct a thorough ablation study to find an optimal setting for the combination of pre-training tasks. Extensive experiments show that UNITER achieves new state of the art across six V+L tasks (over nine datasets), including Visual Question Answering, Image-Text Retrieval, Referring Expression Comprehension, Visual Commonsense Reasoning, Visual Entailment, and NLVR2.
VisualEchoes: Spatial Image Representation Learning through Echolocation
May 04, 2020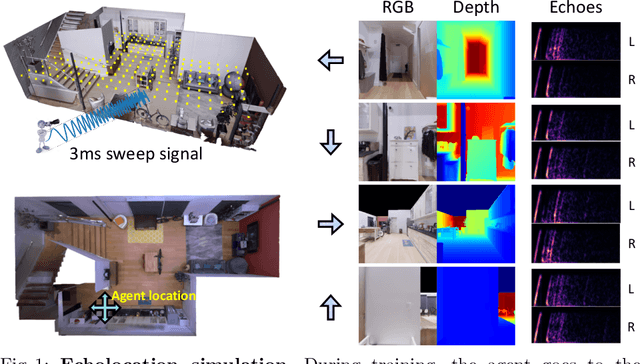
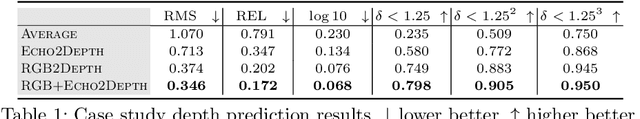
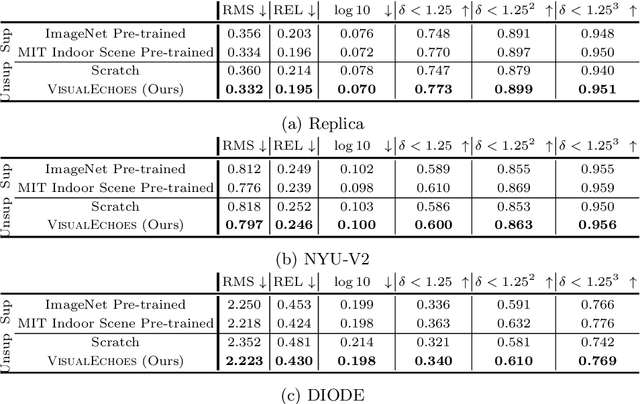
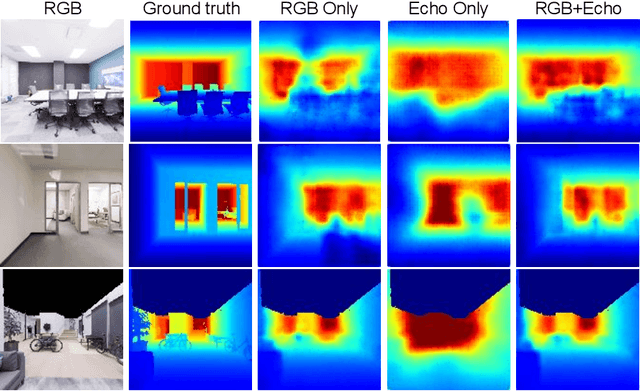
Several animal species (e.g., bats, dolphins, and whales) and even visually impaired humans have the remarkable ability to perform echolocation: a biological sonar used to perceive spatial layout and locate objects in the world. We explore the spatial cues contained in echoes and how they can benefit vision tasks that require spatial reasoning. First we capture echo responses in photo-realistic 3D indoor scene environments. Then we propose a novel interaction-based representation learning framework that learns useful visual features via echolocation. We show that the learned image features are useful for multiple downstream vision tasks requiring spatial reasoning---monocular depth estimation, surface normal estimation, and visual navigation. Our work opens a new path for representation learning for embodied agents, where supervision comes from interacting with the physical world. Our experiments demonstrate that our image features learned from echoes are comparable or even outperform heavily supervised pre-training methods for multiple fundamental spatial tasks.
Fast Strain Estimation and Frame Selection in Ultrasound Elastography using Machine Learning
Oct 16, 2021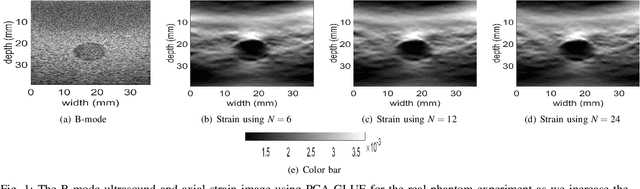
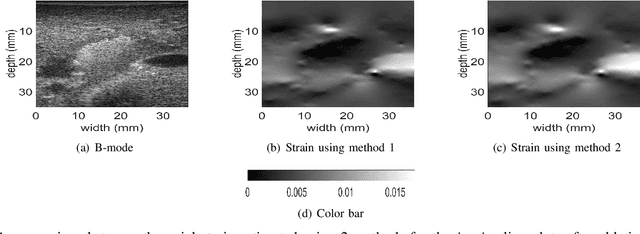
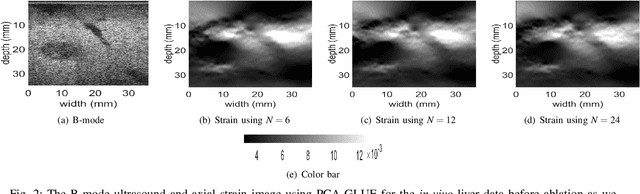
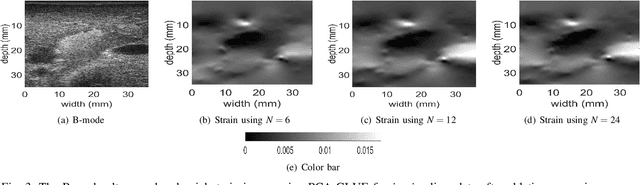
Ultrasound Elastography aims to determine the mechanical properties of the tissue by monitoring tissue deformation due to internal or external forces. Tissue deformations are estimated from ultrasound radio frequency (RF) signals and are often referred to as time delay estimation (TDE). Given two RF frames I1 and I2, we can compute a displacement image which shows the change in the position of each sample in I1 to a new position in I2. Two important challenges in TDE include high computational complexity and the difficulty in choosing suitable RF frames. Selecting suitable frames is of high importance because many pairs of RF frames either do not have acceptable deformation for extracting informative strain images or are decorrelated and deformation cannot be reliably estimated. Herein, we introduce a method that learns 12 displacement modes in quasi-static elastography by performing Principal Component Analysis (PCA) on displacement fields of a large training database. In the inference stage, we use dynamic programming (DP) to compute an initial displacement estimate of around 1% of the samples, and then decompose this sparse displacement into a linear combination of the 12 displacement modes. Our method assumes that the displacement of the whole image could also be described by this linear combination of principal components. We then use the GLobal Ultrasound Elastography (GLUE) method to fine-tune the result yielding the exact displacement image. Our method, which we call PCA-GLUE, is more than 10 times faster than DP in calculating the initial displacement map while giving the same result. Our second contribution in this paper is determining the suitability of the frame pair I1 and I2 for strain estimation, which we achieve by using the weight vector that we calculated for PCA-GLUE as an input to a multi-layer perceptron (MLP) classifier.
DeepMTS: Deep Multi-task Learning for Survival Prediction in Patients with Advanced Nasopharyngeal Carcinoma using Pretreatment PET/CT
Sep 16, 2021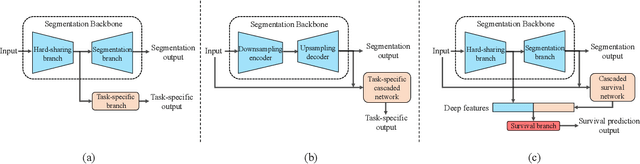
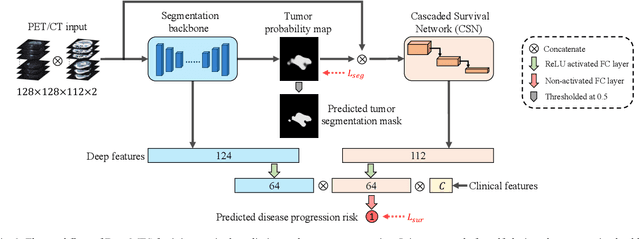
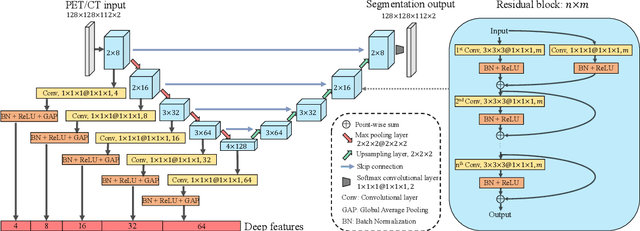
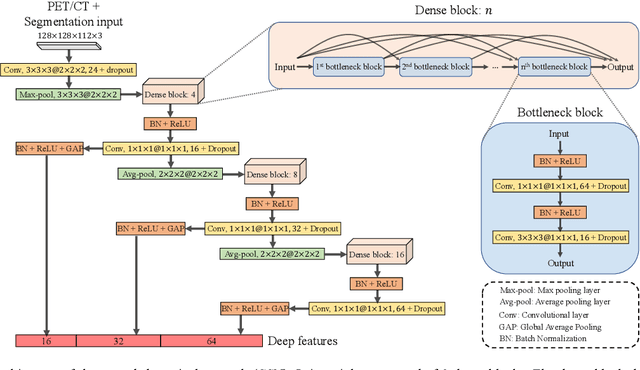
Nasopharyngeal Carcinoma (NPC) is a worldwide malignant epithelial cancer. Survival prediction is a major concern for NPC patients, as it provides early prognostic information that is needed to guide treatments. Recently, deep learning, which leverages Deep Neural Networks (DNNs) to learn deep representations of image patterns, has been introduced to the survival prediction in various cancers including NPC. It has been reported that image-derived end-to-end deep survival models have the potential to outperform clinical prognostic indicators and traditional radiomics-based survival models in prognostic performance. However, deep survival models, especially 3D models, require large image training data to avoid overfitting. Unfortunately, medical image data is usually scarce, especially for Positron Emission Tomography/Computed Tomography (PET/CT) due to the high cost of PET/CT scanning. Compared to Magnetic Resonance Imaging (MRI) or Computed Tomography (CT) providing only anatomical information of tumors, PET/CT that provides both anatomical (from CT) and metabolic (from PET) information is promising to achieve more accurate survival prediction. However, we have not identified any 3D end-to-end deep survival model that applies to small PET/CT data of NPC patients. In this study, we introduced the concept of multi-task leaning into deep survival models to address the overfitting problem resulted from small data. Tumor segmentation was incorporated as an auxiliary task to enhance the model's efficiency of learning from scarce PET/CT data. Based on this idea, we proposed a 3D end-to-end Deep Multi-Task Survival model (DeepMTS) for joint survival prediction and tumor segmentation. Our DeepMTS can jointly learn survival prediction and tumor segmentation using PET/CT data of only 170 patients with advanced NPC.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge