"Image": models, code, and papers
ACTION++: Improving Semi-supervised Medical Image Segmentation with Adaptive Anatomical Contrast
Apr 07, 2023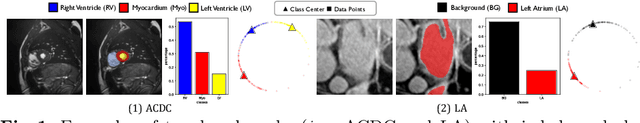
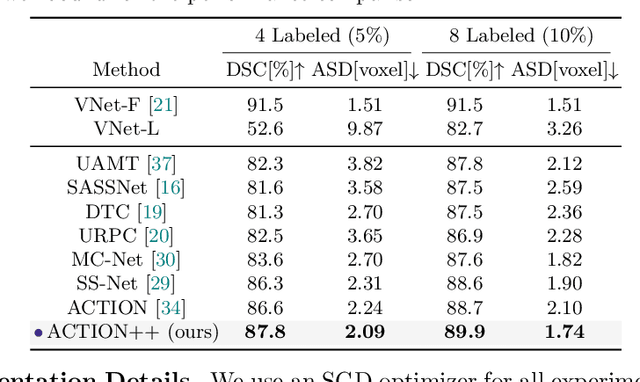
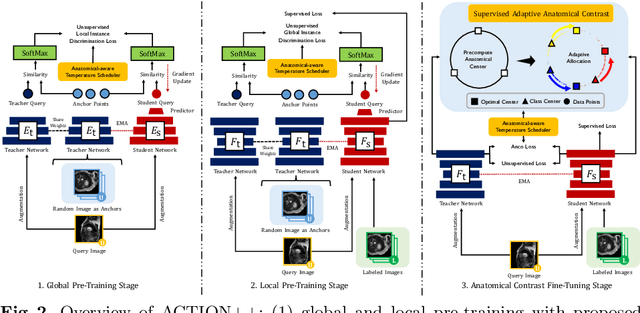
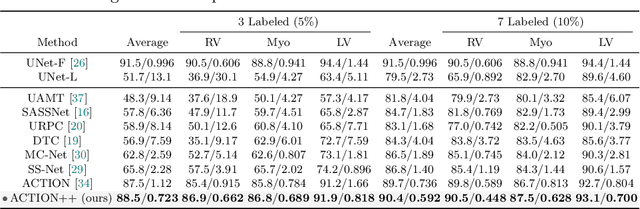
Medical data often exhibits long-tail distributions with heavy class imbalance, which naturally leads to difficulty in classifying the minority classes (i.e., boundary regions or rare objects). Recent work has significantly improved semi-supervised medical image segmentation in long-tailed scenarios by equipping them with unsupervised contrastive criteria. However, it remains unclear how well they will perform in the labeled portion of data where class distribution is also highly imbalanced. In this work, we present ACTION++, an improved contrastive learning framework with adaptive anatomical contrast for semi-supervised medical segmentation. Specifically, we propose an adaptive supervised contrastive loss, where we first compute the optimal locations of class centers uniformly distributed on the embedding space (i.e., off-line), and then perform online contrastive matching training by encouraging different class features to adaptively match these distinct and uniformly distributed class centers. Moreover, we argue that blindly adopting a constant temperature $\tau$ in the contrastive loss on long-tailed medical data is not optimal, and propose to use a dynamic $\tau$ via a simple cosine schedule to yield better separation between majority and minority classes. Empirically, we evaluate ACTION++ on ACDC and LA benchmarks and show that it achieves state-of-the-art across two semi-supervised settings. Theoretically, we analyze the performance of adaptive anatomical contrast and confirm its superiority in label efficiency.
Real-time myocardial landmark tracking for MRI-guided cardiac radio-ablation using Gaussian Processes
Jun 19, 2023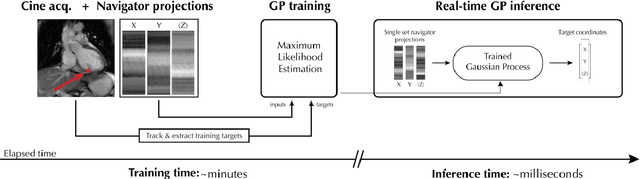
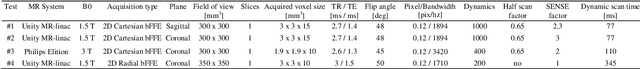
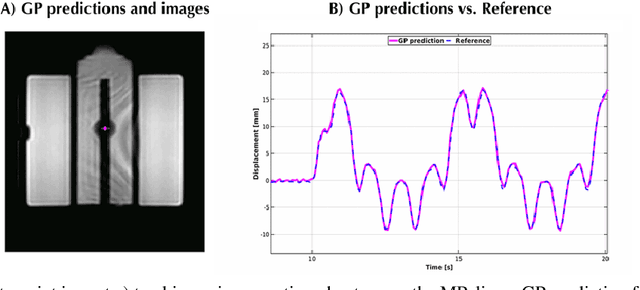
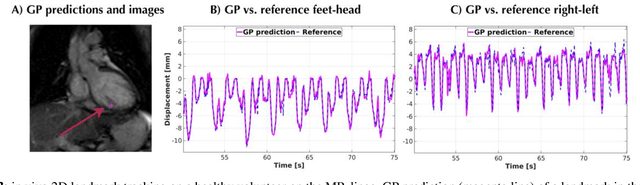
The high speed of cardiorespiratory motion introduces a unique challenge for cardiac stereotactic radio-ablation (STAR) treatments with the MR-linac. Such treatments require tracking myocardial landmarks with a maximum latency of 100 ms, which includes the acquisition of the required data. The aim of this study is to present a new method that allows to track myocardial landmarks from few readouts of MRI data, thereby achieving a latency sufficient for STAR treatments. We present a tracking framework that requires only few readouts of k-space data as input, which can be acquired at least an order of magnitude faster than MR-images. Combined with the real-time tracking speed of a probabilistic machine learning framework called Gaussian Processes, this allows to track myocardial landmarks with a sufficiently low latency for cardiac STAR guidance, including both the acquisition of required data, and the tracking inference. The framework is demonstrated in 2D on a motion phantom, and in vivo on volunteers and a ventricular tachycardia (arrhythmia) patient. Moreover, the feasibility of an extension to 3D was demonstrated by in silico 3D experiments with a digital motion phantom. The framework was compared with template matching - a reference, image-based, method - and linear regression methods. Results indicate an order of magnitude lower total latency (<10 ms) for the proposed framework in comparison with alternative methods. The root-mean-square-distances and mean end-point-distance with the reference tracking method was less than 0.8 mm for all experiments, showing excellent (sub-voxel) agreement. The high accuracy in combination with a total latency of less than 10 ms - including data acquisition and processing - make the proposed method a suitable candidate for tracking during STAR treatments.
Towards Better Explanations for Object Detection
Jun 06, 2023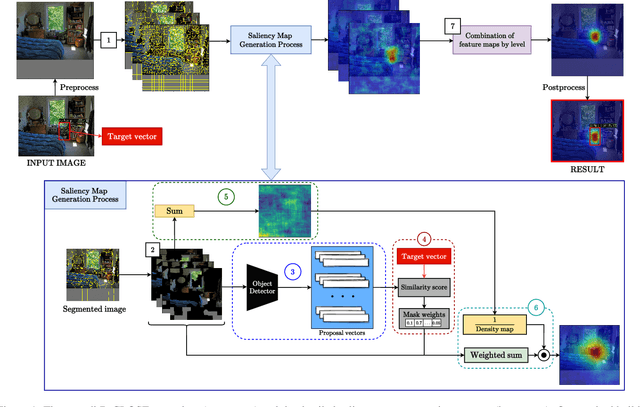

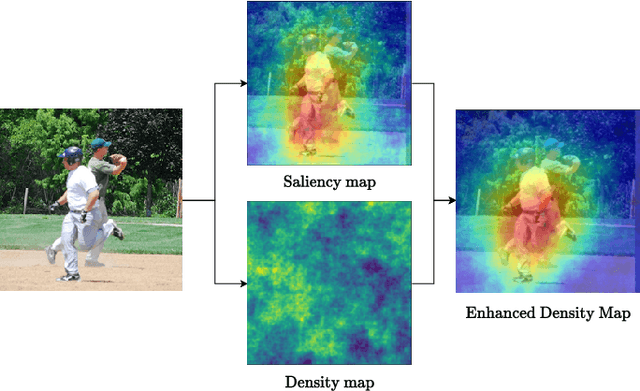
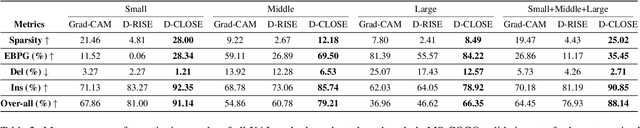
Recent advances in Artificial Intelligence (AI) technology have promoted their use in almost every field. The growing complexity of deep neural networks (DNNs) makes it increasingly difficult and important to explain the inner workings and decisions of the network. However, most current techniques for explaining DNNs focus mainly on interpreting classification tasks. This paper proposes a method to explain the decision for any object detection model called D-CLOSE. To closely track the model's behavior, we used multiple levels of segmentation on the image and a process to combine them. We performed tests on the MS-COCO dataset with the YOLOX model, which shows that our method outperforms D-RISE and can give a better quality and less noise explanation.
Single-Shot Global Localization via Graph-Theoretic Correspondence Matching
Jun 06, 2023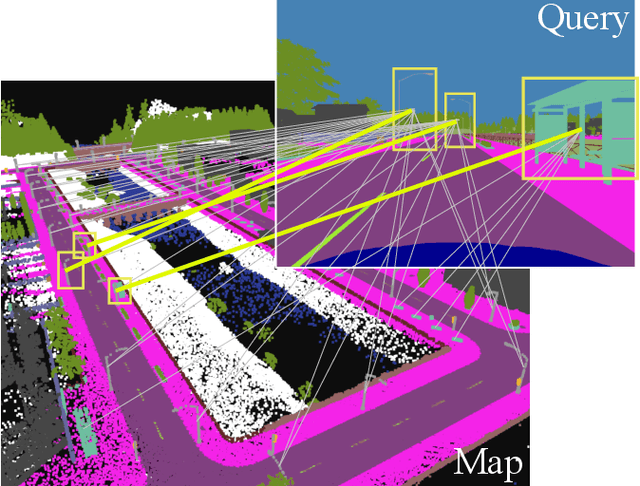
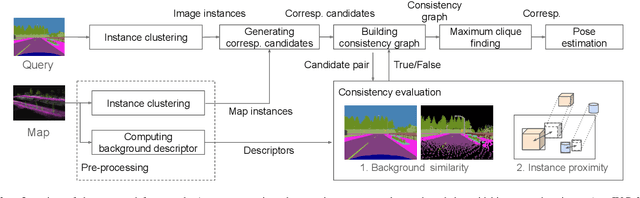
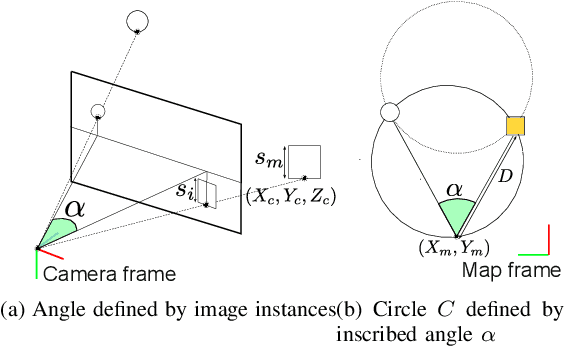
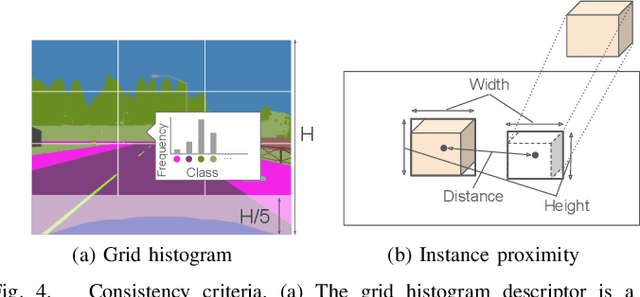
This paper describes a method of global localization based on graph-theoretic association of instances between a query and the prior map. The proposed framework employs correspondence matching based on the maximum clique problem (MCP). The framework is potentially applicable to other map and/or query modalities thanks to the graph-based abstraction of the problem, while many of existing global localization methods rely on a query and the dataset in the same modality. We implement it with a semantically labeled 3D point cloud map, and a semantic segmentation image as a query. Leveraging the graph-theoretic framework, the proposed method realizes global localization exploiting only the map and the query. The method shows promising results on multiple large-scale simulated maps of urban scenes.
DermSynth3D: Synthesis of in-the-wild Annotated Dermatology Images
May 22, 2023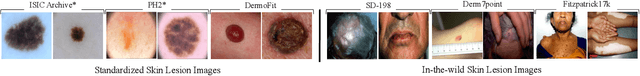
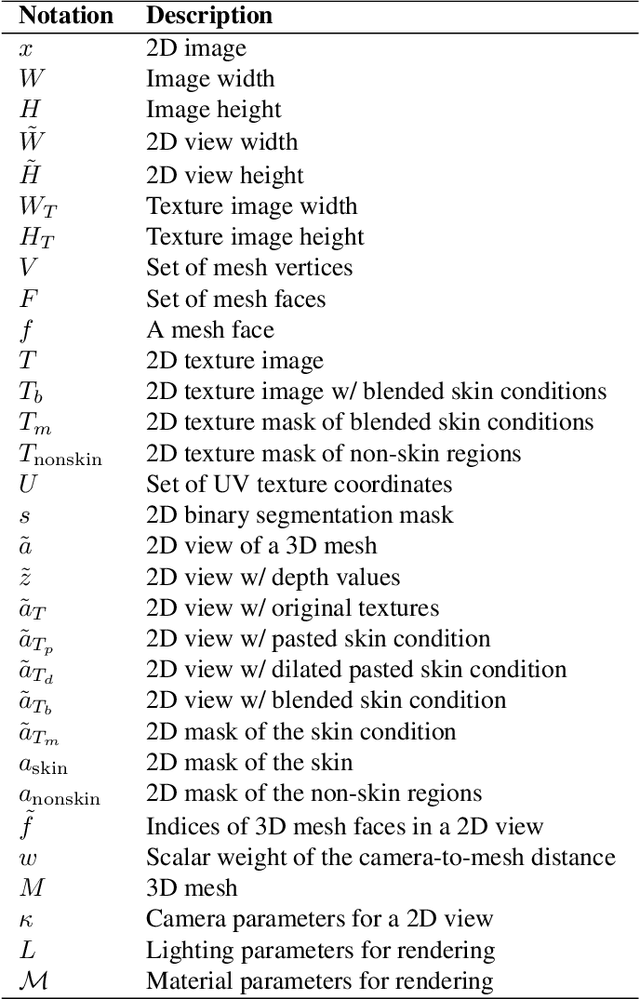
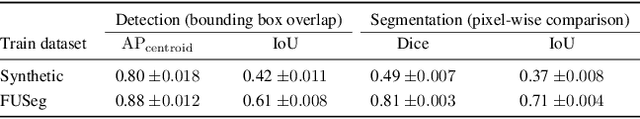

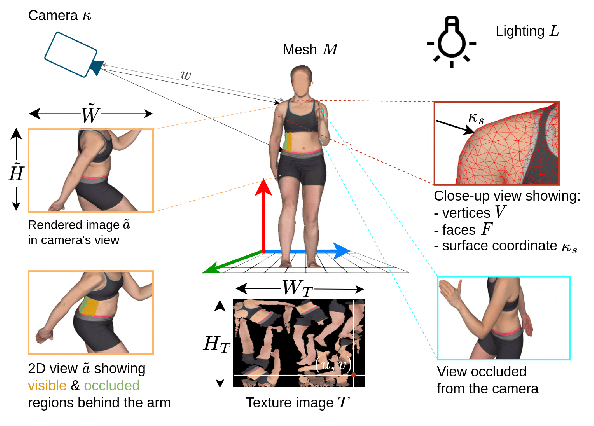

In recent years, deep learning (DL) has shown great potential in the field of dermatological image analysis. However, existing datasets in this domain have significant limitations, including a small number of image samples, limited disease conditions, insufficient annotations, and non-standardized image acquisitions. To address these shortcomings, we propose a novel framework called DermSynth3D. DermSynth3D blends skin disease patterns onto 3D textured meshes of human subjects using a differentiable renderer and generates 2D images from various camera viewpoints under chosen lighting conditions in diverse background scenes. Our method adheres to top-down rules that constrain the blending and rendering process to create 2D images with skin conditions that mimic in-the-wild acquisitions, ensuring more meaningful results. The framework generates photo-realistic 2D dermoscopy images and the corresponding dense annotations for semantic segmentation of the skin, skin conditions, body parts, bounding boxes around lesions, depth maps, and other 3D scene parameters, such as camera position and lighting conditions. DermSynth3D allows for the creation of custom datasets for various dermatology tasks. We demonstrate the effectiveness of data generated using DermSynth3D by training DL models on synthetic data and evaluating them on various dermatology tasks using real 2D dermatological images. We make our code publicly available at https://github.com/sfu-mial/DermSynth3D
Improvement of Color Image Analysis Using a New Hybrid Face Recognition Algorithm based on Discrete Wavelets and Chebyshev Polynomials
Mar 23, 2023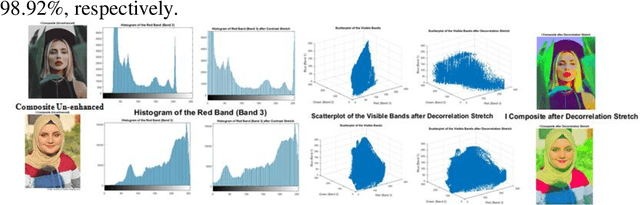
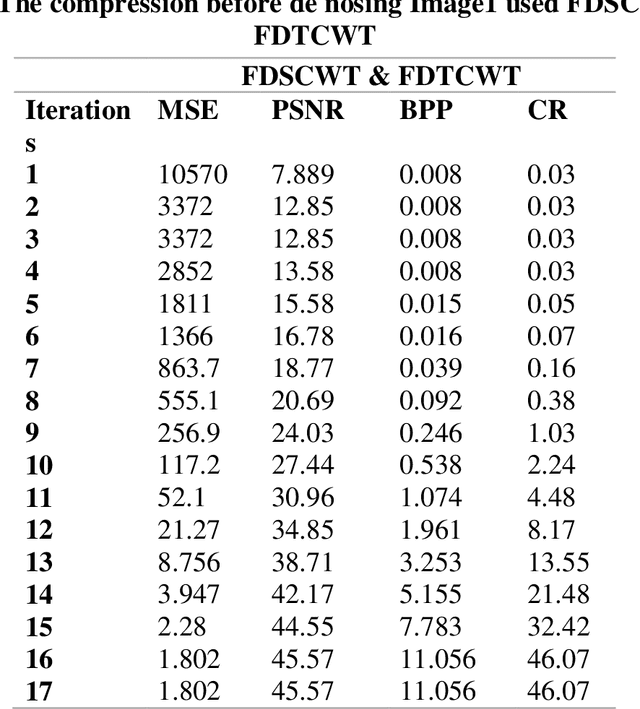
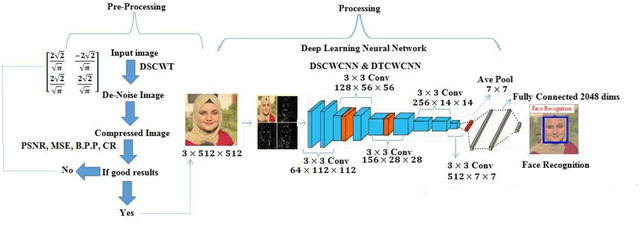
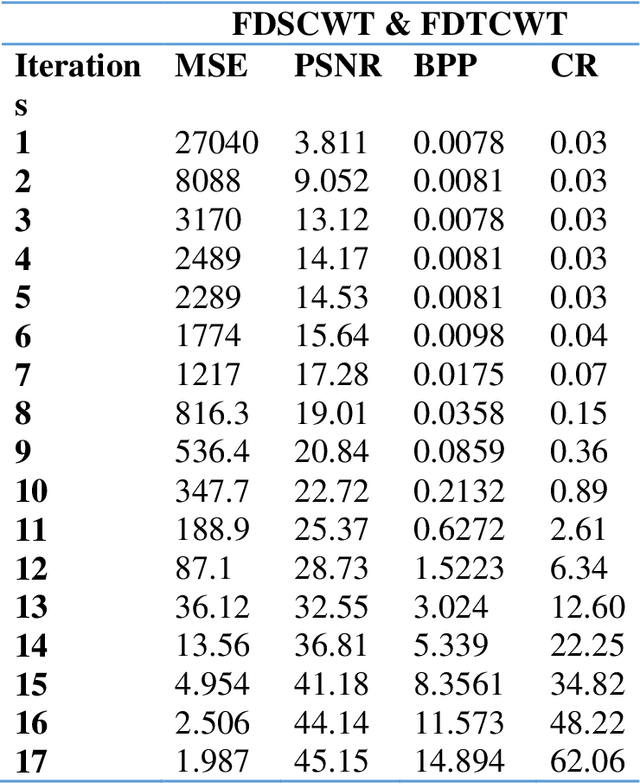
This work is unique in the use of discrete wavelets that were built from or derived from Chebyshev polynomials of the second and third kind, filter the Discrete Second Chebyshev Wavelets Transform (DSCWT), and derive two effective filters. The Filter Discrete Third Chebyshev Wavelets Transform (FDTCWT) is used in the process of analyzing color images and removing noise and impurities that accompany the image, as well as because of the large amount of data that makes up the image as it is taken. These data are massive, making it difficult to deal with each other during transmission. However to address this issue, the image compression technique is used, with the image not losing information due to the readings that were obtained, and the results were satisfactory. Mean Square Error (MSE), Peak Signal Noise Ratio (PSNR), Bit Per Pixel (BPP), and Compression Ratio (CR) Coronavirus is the initial treatment, while the processing stage is done with network training for Convolutional Neural Networks (CNN) with Discrete Second Chebeshev Wavelets Convolutional Neural Network (DSCWCNN) and Discrete Third Chebeshev Wavelets Convolutional Neural Network (DTCWCNN) to create an efficient algorithm for face recognition, and the best results were achieved in accuracy and in the least amount of time. Two samples of color images that were made or implemented were used. The proposed theory was obtained with fast and good results; the results are evident shown in the tables below.
Revisiting model self-interpretability in a decision-theoretic way for binary medical image classification
Mar 13, 2023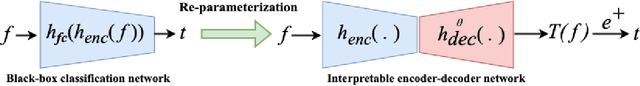
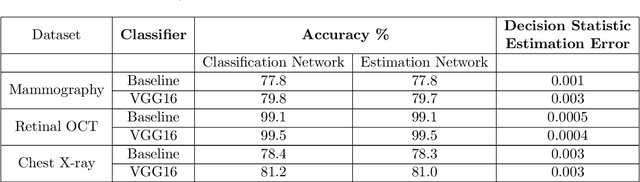

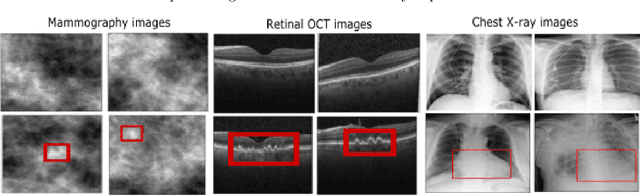
Interpretability is highly desired for deep neural network-based classifiers, especially when addressing high-stake decisions in medical imaging. Commonly used post-hoc interpretability methods may not be always useful because different such methods can produce several plausible but different interpretations of a given model, leading to confusion about which one to choose. {In this work, an {inherently} interpretable encoder-decoder model coupled with a single-layer fully connected network with unity weights is proposed for binary medical image classification problems. The feature extraction component of a trained black-box network for the same task is employed as the pre-trained encoder of the interpretable model. The model is trained to estimate the decision statistic of the given trained black-box deep binary classifier to maintain a similar accuracy.} The decoder output represents a transformed version of the to-be-classified image that, when processed by the fixed fully connected layer, produces the same decision statistic value as the original classifier. This is accomplished by minimizing the mean squared error between the decision statistic values of the black-box model and encoder-decoder based model during training. The decoder output image is referred to as an equivalency map. Because the single-layer network is fully interpretable, the equivalency map provides a visualization of the transformed image features that contribute to the decision statistic value and, moreover, permits quantification of their relative contributions. Unlike the traditional post-hoc interpretability methods, the proposed method is inherently interpretable, quantitative, and fundamentally based on decision theory.
LAMM: Language-Assisted Multi-Modal Instruction-Tuning Dataset, Framework, and Benchmark
Jun 11, 2023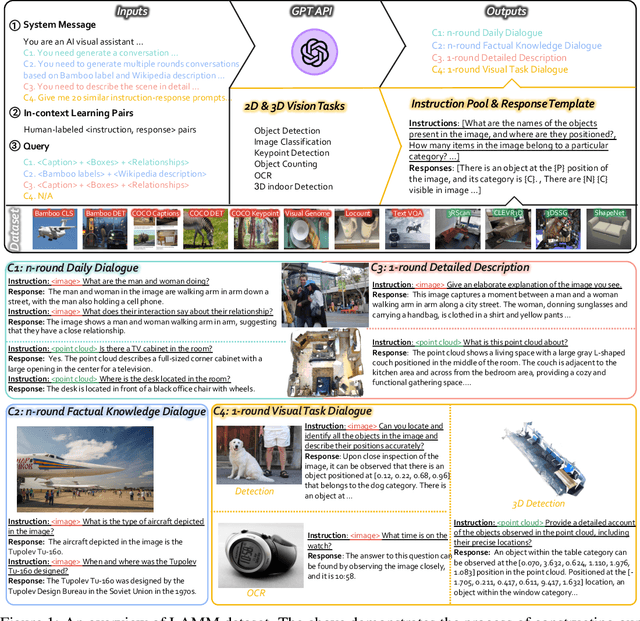
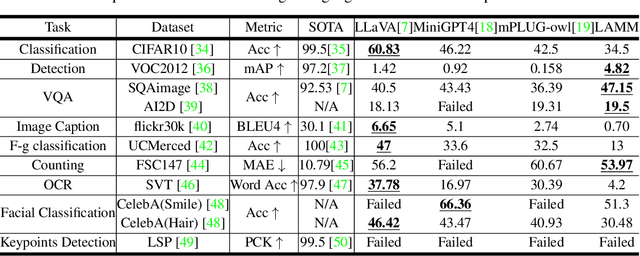
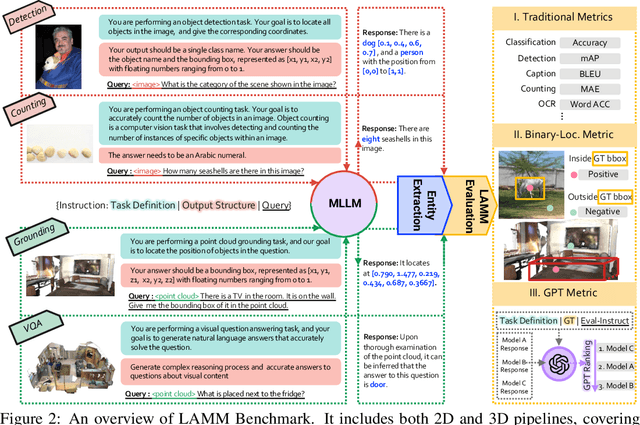
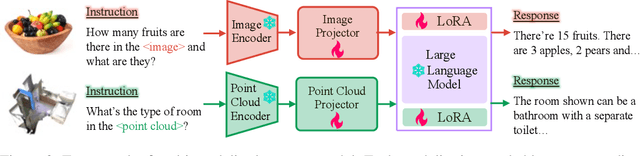
Large language models have become a potential pathway toward achieving artificial general intelligence. Recent works on multi-modal large language models have demonstrated their effectiveness in handling visual modalities. In this work, we extend the research of MLLMs to point clouds and present the LAMM-Dataset and LAMM-Benchmark for 2D image and 3D point cloud understanding. We also establish an extensible framework to facilitate the extension of MLLMs to additional modalities. Our main contribution is three-fold: 1) We present the LAMM-Dataset and LAMM-Benchmark, which cover almost all high-level vision tasks for 2D and 3D vision. Extensive experiments validate the effectiveness of our dataset and benchmark. 2) We demonstrate the detailed methods of constructing instruction-tuning datasets and benchmarks for MLLMs, which will enable future research on MLLMs to scale up and extend to other domains, tasks, and modalities faster. 3) We provide a primary but potential MLLM training framework optimized for modalities' extension. We also provide baseline models, comprehensive experimental observations, and analysis to accelerate future research.
3rd Place Solution for PVUW Challenge 2023: Video Panoptic Segmentation
Jun 11, 2023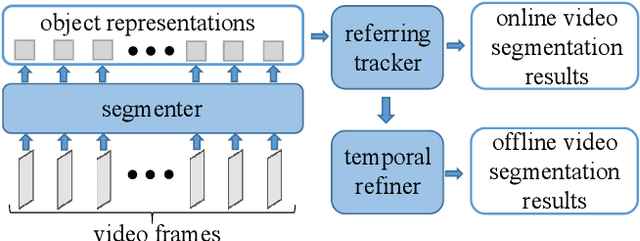
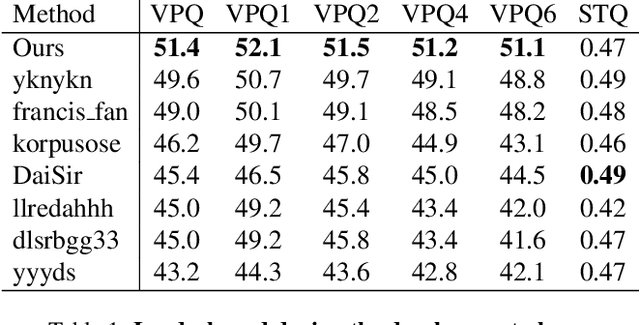
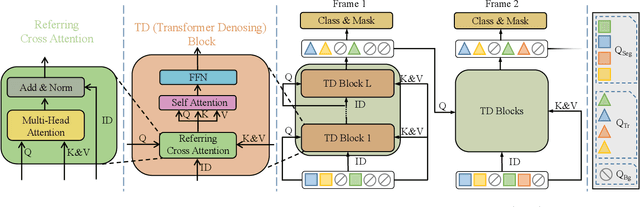
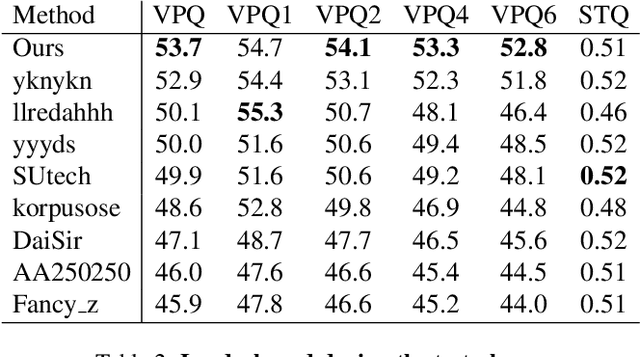
In order to deal with the task of video panoptic segmentation in the wild, we propose a robust integrated video panoptic segmentation solution. In our solution, we regard the video panoptic segmentation task as a segmentation target querying task, represent both semantic and instance targets as a set of queries, and then combine these queries with video features extracted by neural networks to predict segmentation masks. In order to improve the learning accuracy and convergence speed of the solution, we add additional tasks of video semantic segmentation and video instance segmentation for joint training. In addition, we also add an additional image semantic segmentation model to further improve the performance of semantic classes. In addition, we also add some additional operations to improve the robustness of the model. Extensive experiments on the VIPSeg dataset show that the proposed solution achieves state-of-the-art performance with 50.04\% VPQ on the VIPSeg test set, which is 3rd place on the video panoptic segmentation track of the PVUW Challenge 2023.
On the Efficacy of 3D Point Cloud Reinforcement Learning
Jun 11, 2023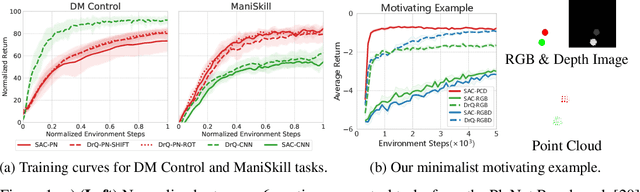
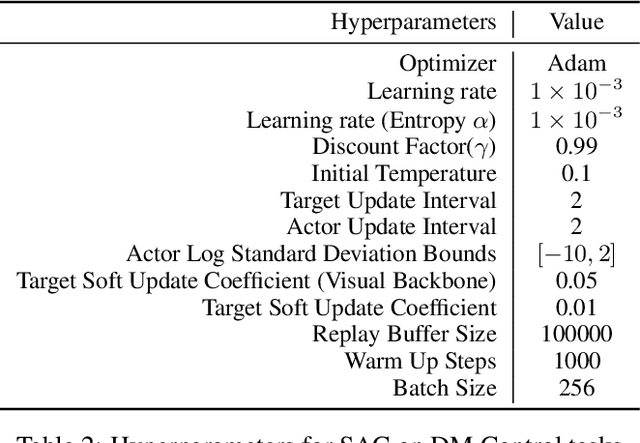
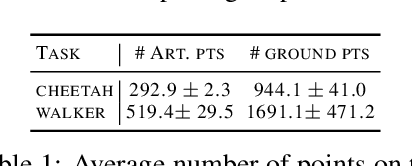


Recent studies on visual reinforcement learning (visual RL) have explored the use of 3D visual representations. However, none of these work has systematically compared the efficacy of 3D representations with 2D representations across different tasks, nor have they analyzed 3D representations from the perspective of agent-object / object-object relationship reasoning. In this work, we seek answers to the question of when and how do 3D neural networks that learn features in the 3D-native space provide a beneficial inductive bias for visual RL. We specifically focus on 3D point clouds, one of the most common forms of 3D representations. We systematically investigate design choices for 3D point cloud RL, leading to the development of a robust algorithm for various robotic manipulation and control tasks. Furthermore, through comparisons between 2D image vs 3D point cloud RL methods on both minimalist synthetic tasks and complex robotic manipulation tasks, we find that 3D point cloud RL can significantly outperform the 2D counterpart when agent-object / object-object relationship encoding is a key factor.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge