Zhe Luo
GenoBERT: A Language Model for Accurate Genotype Imputation
Mar 31, 2026Abstract:Genotype imputation enables dense variant coverage for genome-wide association and risk-prediction studies, yet conventional reference-panel methods remain limited by ancestry bias and reduced rare-variant accuracy. We present Genotype Bidirectional Encoder Representations from Transformers (GenoBERT), a transformer-based, reference-free framework that tokenizes phased genotypes and uses a self-attention mechanism to capture both short- and long-range linkage disequilibrium (LD) dependencies. Benchmarking on two independent datasets including the Louisiana Osteoporosis Study (LOS) and the 1000 Genomes Project (1KGP) across ancestry groups and multiple genotype missingness levels (5-50%) shows that GenoBERT achieves the highest overall accuracy compared to four baseline methods (Beagle5.4, SCDA, BiU-Net, and STICI). At practical sparsity levels (up to 25% missing), GenoBERT attains high overall imputation accuracy ($r^2 approx 0.98$) across datasets, and maintains robust performance ($r^2 > 0.90$) even at 50% missingness. Experimental results across different ancestries confirm consistent gains across datasets, with resilience to small sample sizes and weak LD. A 128-SNP (single-nucleotide polymorphism) context window (approximately 100 Kb) is validated through LD-decay analyses as sufficient to capture local correlation structures. By eliminating reference-panel dependence while preserving high accuracy, GenoBERT provides a scalable and robust solution for genotype imputation and a foundation for downstream genomic modeling.
Cefdet: Cognitive Effectiveness Network Based on Fuzzy Inference for Action Detection
Oct 08, 2024



Abstract:Action detection and understanding provide the foundation for the generation and interaction of multimedia content. However, existing methods mainly focus on constructing complex relational inference networks, overlooking the judgment of detection effectiveness. Moreover, these methods frequently generate detection results with cognitive abnormalities. To solve the above problems, this study proposes a cognitive effectiveness network based on fuzzy inference (Cefdet), which introduces the concept of "cognition-based detection" to simulate human cognition. First, a fuzzy-driven cognitive effectiveness evaluation module (FCM) is established to introduce fuzzy inference into action detection. FCM is combined with human action features to simulate the cognition-based detection process, which clearly locates the position of frames with cognitive abnormalities. Then, a fuzzy cognitive update strategy (FCS) is proposed based on the FCM, which utilizes fuzzy logic to re-detect the cognition-based detection results and effectively update the results with cognitive abnormalities. Experimental results demonstrate that Cefdet exhibits superior performance against several mainstream algorithms on the public datasets, validating its effectiveness and superiority.
DPER: Diffusion Prior Driven Neural Representation for Limited Angle and Sparse View CT Reconstruction
Apr 27, 2024



Abstract:Limited-angle and sparse-view computed tomography (LACT and SVCT) are crucial for expanding the scope of X-ray CT applications. However, they face challenges due to incomplete data acquisition, resulting in diverse artifacts in the reconstructed CT images. Emerging implicit neural representation (INR) techniques, such as NeRF, NeAT, and NeRP, have shown promise in under-determined CT imaging reconstruction tasks. However, the unsupervised nature of INR architecture imposes limited constraints on the solution space, particularly for the highly ill-posed reconstruction task posed by LACT and ultra-SVCT. In this study, we introduce the Diffusion Prior Driven Neural Representation (DPER), an advanced unsupervised framework designed to address the exceptionally ill-posed CT reconstruction inverse problems. DPER adopts the Half Quadratic Splitting (HQS) algorithm to decompose the inverse problem into data fidelity and distribution prior sub-problems. The two sub-problems are respectively addressed by INR reconstruction scheme and pre-trained score-based diffusion model. This combination initially preserves the implicit image local consistency prior from INR. Additionally, it effectively augments the feasibility of the solution space for the inverse problem through the generative diffusion model, resulting in increased stability and precision in the solutions. We conduct comprehensive experiments to evaluate the performance of DPER on LACT and ultra-SVCT reconstruction with two public datasets (AAPM and LIDC). The results show that our method outperforms the state-of-the-art reconstruction methods on in-domain datasets, while achieving significant performance improvements on out-of-domain datasets.
Multi-View Variational Autoencoder for Missing Value Imputation in Untargeted Metabolomics
Oct 12, 2023


Abstract:Background: Missing data is a common challenge in mass spectrometry-based metabolomics, which can lead to biased and incomplete analyses. The integration of whole-genome sequencing (WGS) data with metabolomics data has emerged as a promising approach to enhance the accuracy of data imputation in metabolomics studies. Method: In this study, we propose a novel method that leverages the information from WGS data and reference metabolites to impute unknown metabolites. Our approach utilizes a multi-view variational autoencoder to jointly model the burden score, polygenetic risk score (PGS), and linkage disequilibrium (LD) pruned single nucleotide polymorphisms (SNPs) for feature extraction and missing metabolomics data imputation. By learning the latent representations of both omics data, our method can effectively impute missing metabolomics values based on genomic information. Results: We evaluate the performance of our method on empirical metabolomics datasets with missing values and demonstrate its superiority compared to conventional imputation techniques. Using 35 template metabolites derived burden scores, PGS and LD-pruned SNPs, the proposed methods achieved r2-scores > 0.01 for 71.55% of metabolites. Conclusion: The integration of WGS data in metabolomics imputation not only improves data completeness but also enhances downstream analyses, paving the way for more comprehensive and accurate investigations of metabolic pathways and disease associations. Our findings offer valuable insights into the potential benefits of utilizing WGS data for metabolomics data imputation and underscore the importance of leveraging multi-modal data integration in precision medicine research.
Multi-view information fusion using multi-view variational autoencoders to predict proximal femoral strength
Oct 03, 2022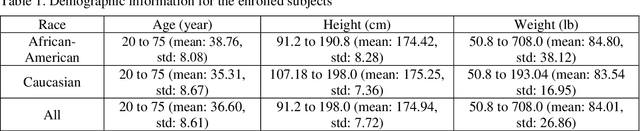
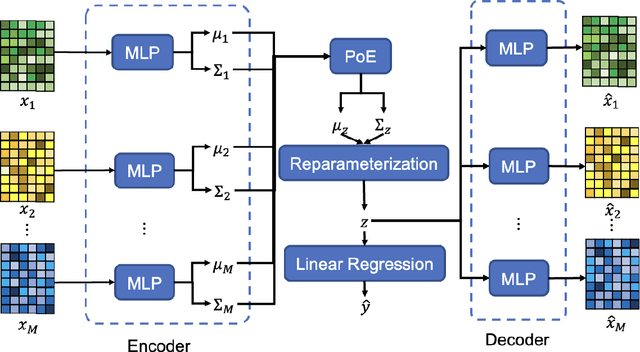
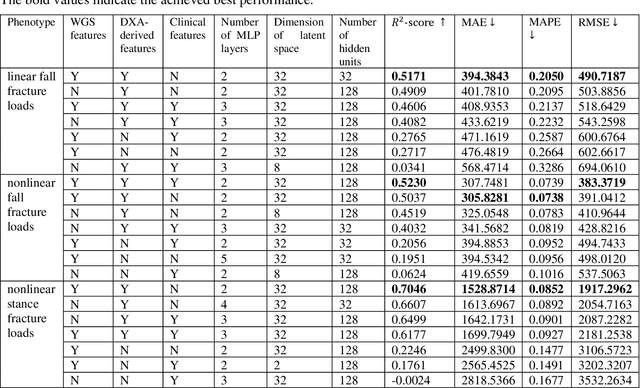
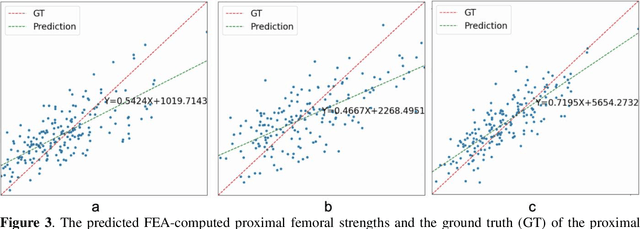
Abstract:Background and aim: Hip fracture can be devastating. The proximal femoral strength can be computed by subject-specific finite element (FE) analysis (FEA) using quantitative CT images. The aim of this paper is to design a deep learning-based model for hip fracture prediction with multi-view information fusion. Method: We developed a multi-view variational autoencoder (MMVAE) for feature representation learning and designed the product of expert model (PoE) for multi-view information fusion.We performed genome-wide association studies (GWAS) to select the most relevant genetic features with proximal femoral strengths and integrated genetic features with DXA-derived imaging features and clinical variables for proximal femoral strength prediction. Results: The designed model achieved the mean absolute percentage error of 0.2050,0.0739 and 0.0852 for linear fall, nonlinear fall and nonlinear stance fracture load prediction, respectively. For linear fall and nonlinear stance fracture load prediction, integrating genetic and DXA-derived imaging features were beneficial; while for nonlinear fall fracture load prediction, integrating genetic features, DXA-derived imaging features as well as clinical variables, the model achieved the best performance. Conclusion: The proposed model is capable of predicting proximal femoral strengths using genetic features, DXA-derived imaging features as well as clinical variables. Compared to performing FEA using QCT images to calculate proximal femoral strengths, the presented method is time-efficient and cost effective, and radiation dosage is limited. From the technique perspective, the final models can be applied to other multi-view information integration tasks.
Robust and Efficient Boosting Method using the Conditional Risk
Jun 21, 2018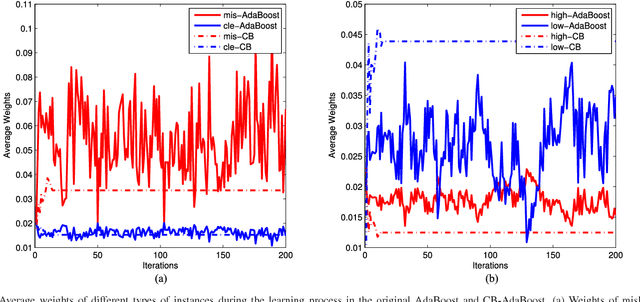
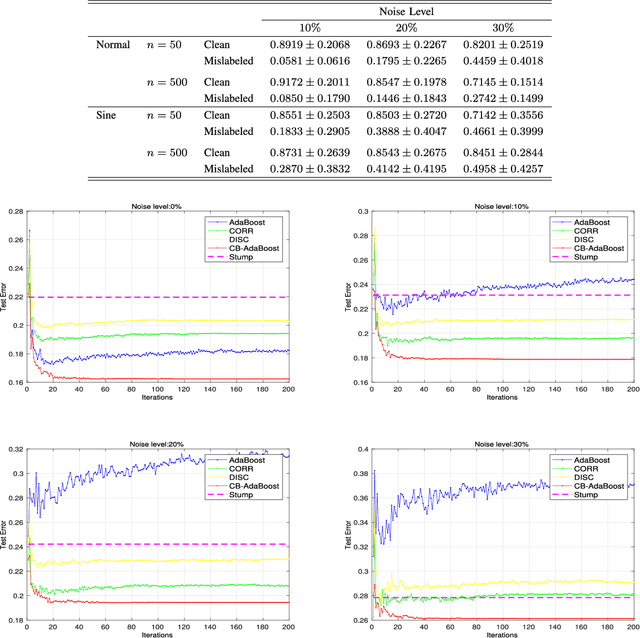
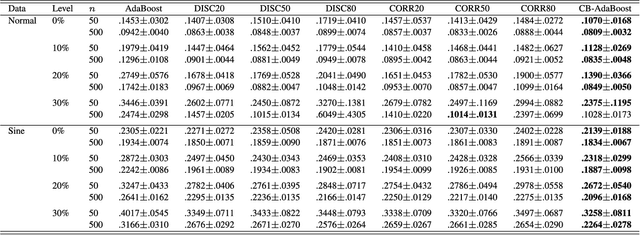
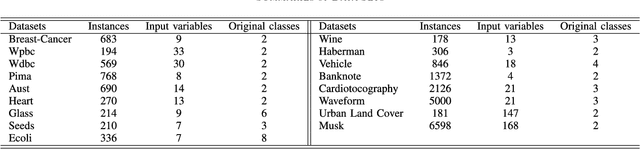
Abstract:Well-known for its simplicity and effectiveness in classification, AdaBoost, however, suffers from overfitting when class-conditional distributions have significant overlap. Moreover, it is very sensitive to noise that appears in the labels. This article tackles the above limitations simultaneously via optimizing a modified loss function (i.e., the conditional risk). The proposed approach has the following two advantages. (1) It is able to directly take into account label uncertainty with an associated label confidence. (2) It introduces a "trustworthiness" measure on training samples via the Bayesian risk rule, and hence the resulting classifier tends to have finite sample performance that is superior to that of the original AdaBoost when there is a large overlap between class conditional distributions. Theoretical properties of the proposed method are investigated. Extensive experimental results using synthetic data and real-world data sets from UCI machine learning repository are provided. The empirical study shows the high competitiveness of the proposed method in predication accuracy and robustness when compared with the original AdaBoost and several existing robust AdaBoost algorithms.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge