Xinwei Sun
Causal Hidden Markov Model for Time Series Disease Forecasting
Mar 30, 2021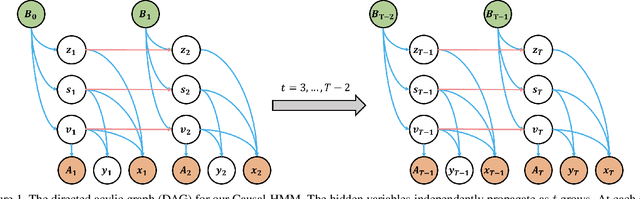
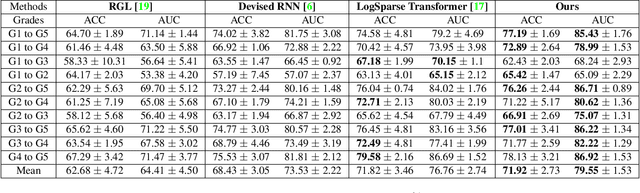
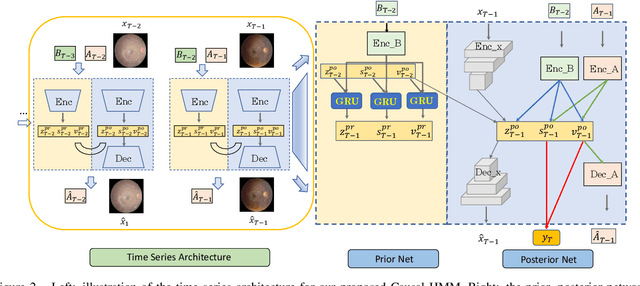
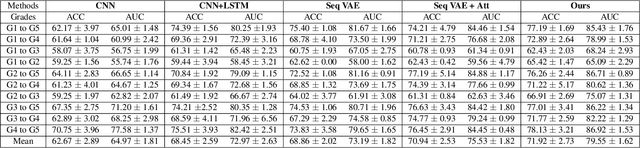
Abstract:We propose a causal hidden Markov model to achieve robust prediction of irreversible disease at an early stage, which is safety-critical and vital for medical treatment in early stages. Specifically, we introduce the hidden variables which propagate to generate medical data at each time step. To avoid learning spurious correlation (e.g., confounding bias), we explicitly separate these hidden variables into three parts: a) the disease (clinical)-related part; b) the disease (non-clinical)-related part; c) others, with only a),b) causally related to the disease however c) may contain spurious correlations (with the disease) inherited from the data provided. With personal attributes and the disease label respectively provided as side information and supervision, we prove that these disease-related hidden variables can be disentangled from others, implying the avoidance of spurious correlation for generalization to medical data from other (out-of-) distributions. Guaranteed by this result, we propose a sequential variational auto-encoder with a reformulated objective function. We apply our model to the early prediction of peripapillary atrophy and achieve promising results on out-of-distribution test data. Further, the ablation study empirically shows the effectiveness of each component in our method. And the visualization shows the accurate identification of lesion regions from others.
Disease Forecast via Progression Learning
Dec 21, 2020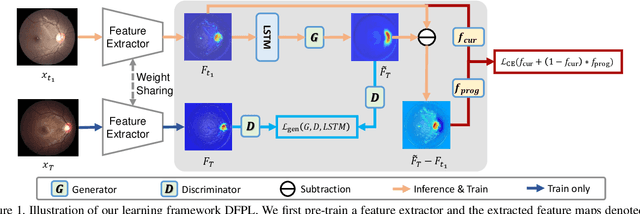
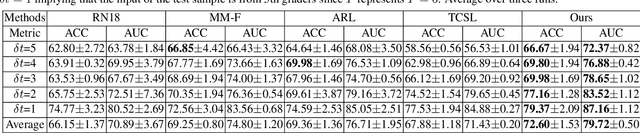
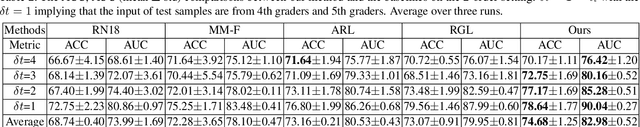
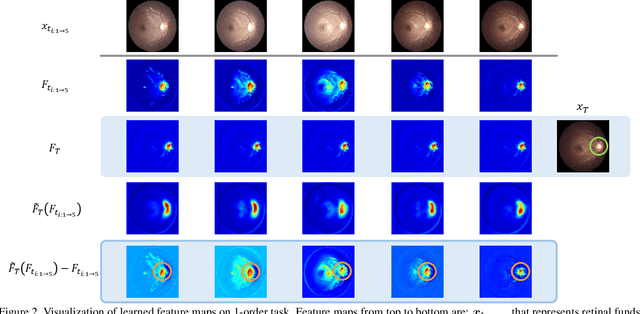
Abstract:Forecasting Parapapillary atrophy (PPA), i.e., a symptom related to most irreversible eye diseases, provides an alarm for implementing an intervention to slow down the disease progression at early stage. A key question for this forecast is: how to fully utilize the historical data (e.g., retinal image) up to the current stage for future disease prediction? In this paper, we provide an answer with a novel framework, namely \textbf{D}isease \textbf{F}orecast via \textbf{P}rogression \textbf{L}earning (\textbf{DFPL}), which exploits the irreversibility prior (i.e., cannot be reversed once diagnosed). Specifically, based on this prior, we decompose two factors that contribute to the prediction of the future disease: i) the current disease label given the data (retinal image, clinical attributes) at present and ii) the future disease label given the progression of the retinal images that from the current to the future. To model these two factors, we introduce the current and progression predictors in DFPL, respectively. In order to account for the degree of progression of the disease, we propose a temporal generative model to accurately generate the future image and compare it with the current one to get a residual image. The generative model is implemented by a recurrent neural network, in order to exploit the dependency of the historical data. To verify our approach, we apply it to a PPA in-house dataset and it yields a significant improvement (\textit{e.g.}, \textbf{4.48\%} of accuracy; \textbf{3.45\%} of AUC) over others. Besides, our generative model can accurately localize the disease-related regions.
Identifying Invariant Texture Violation for Robust Deepfake Detection
Dec 19, 2020
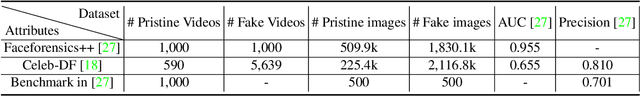
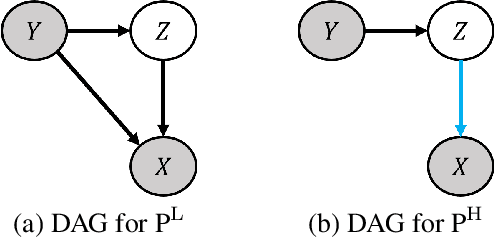
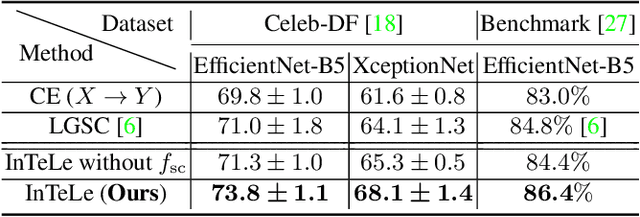
Abstract:Existing deepfake detection methods have reported promising in-distribution results, by accessing published large-scale dataset. However, due to the non-smooth synthesis method, the fake samples in this dataset may expose obvious artifacts (e.g., stark visual contrast, non-smooth boundary), which were heavily relied on by most of the frame-level detection methods above. As these artifacts do not come up in real media forgeries, the above methods can suffer from a large degradation when applied to fake images that close to reality. To improve the robustness for high-realism fake data, we propose the Invariant Texture Learning (InTeLe) framework, which only accesses the published dataset with low visual quality. Our method is based on the prior that the microscopic facial texture of the source face is inevitably violated by the texture transferred from the target person, which can hence be regarded as the invariant characterization shared among all fake images. To learn such an invariance for deepfake detection, our InTeLe introduces an auto-encoder framework with different decoders for pristine and fake images, which are further appended with a shallow classifier in order to separate out the obvious artifact-effect. Equipped with such a separation, the extracted embedding by encoder can capture the texture violation in fake images, followed by the classifier for the final pristine/fake prediction. As a theoretical guarantee, we prove the identifiability of such an invariance texture violation, i.e., to be precisely inferred from observational data. The effectiveness and utility of our method are demonstrated by promising generalization ability from low-quality images with obvious artifacts to fake images with high realism.
Latent Causal Invariant Model
Nov 04, 2020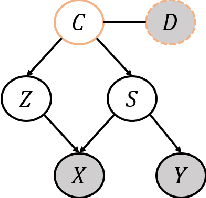
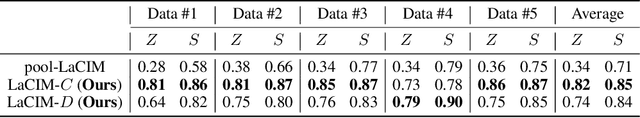
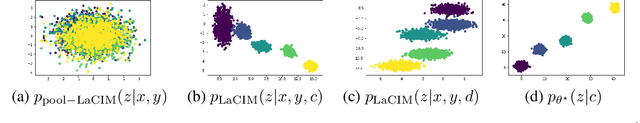
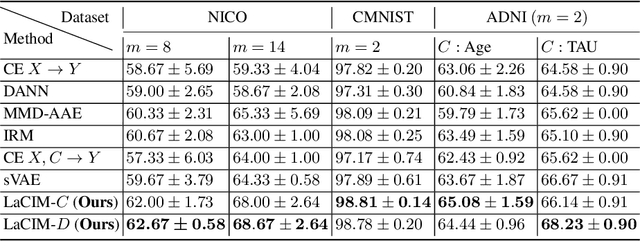
Abstract:Current supervised learning can learn spurious correlation during the data-fitting process, imposing issues regarding interpretability, out-of-distribution (OOD) generalization, and robustness. To avoid spurious correlation, we propose a Latent Causal Invariance Model (LaCIM) which pursues causal prediction. Specifically, we introduce latent variables that are separated into (a) output-causative factors and (b) others that are spuriously correlated to the output via confounders, to model the underlying causal factors. We further assume the generating mechanisms from latent space to observed data to be causally invariant. We give the identifiable claim of such invariance, particularly the disentanglement of output-causative factors from others, as a theoretical guarantee for precise inference and avoiding spurious correlation. We propose a Variational-Bayesian-based method for estimation and to optimize over the latent space for prediction. The utility of our approach is verified by improved interpretability, prediction power on various OOD scenarios (including healthcare) and robustness on security.
Learning Causal Semantic Representation for Out-of-Distribution Prediction
Nov 03, 2020
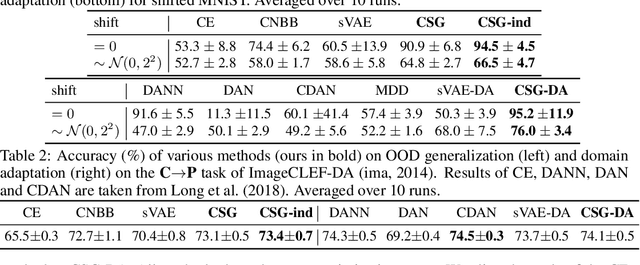
Abstract:Conventional supervised learning methods, especially deep ones, are found to be sensitive to out-of-distribution (OOD) examples, largely because the learned representation mixes the semantic factor with the variation factor due to their domain-specific correlation, while only the semantic factor causes the output. To address the problem, we propose a Causal Semantic Generative model (CSG) based on causality to model the two factors separately, and learn it on a single training domain for prediction without (OOD generalization) or with (domain adaptation) unsupervised data in a test domain. We prove that CSG identifies the semantic factor on the training domain, and the invariance principle of causality subsequently guarantees the boundedness of OOD generalization error and the success of adaptation. We design learning methods for both effective learning and easy prediction, by leveraging the graphical structure of CSG. Empirical study demonstrates the effect of our methods to improve test accuracy for OOD generalization and domain adaptation.
Bilateral Asymmetry Guided Counterfactual Generating Network for Mammogram Classification
Sep 30, 2020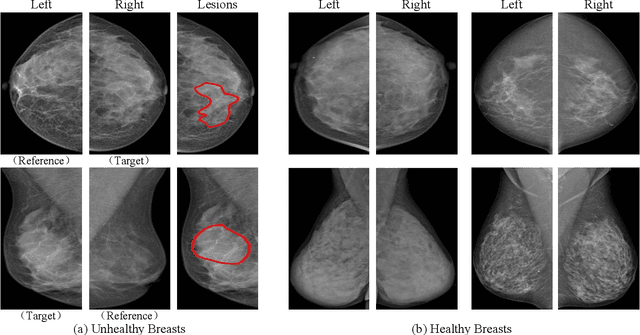
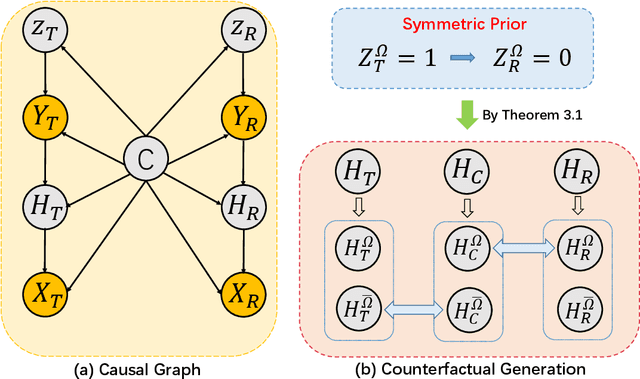
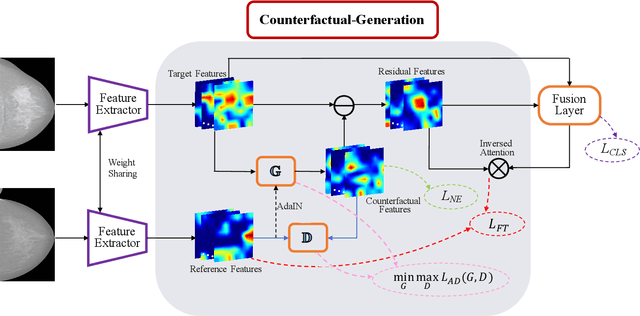
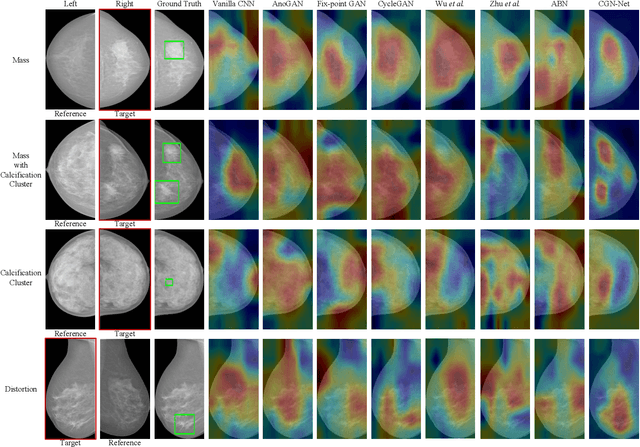
Abstract:Mammogram benign or malignant classification with only image-level labels is challenging due to the absence of lesion annotations. Motivated by the symmetric prior that the lesions on one side of breasts rarely appear in the corresponding areas on the other side, given a diseased image, we can explore a counterfactual problem that how would the features have behaved if there were no lesions in the image, so as to identify the lesion areas. We derive a new theoretical result for counterfactual generation based on the symmetric prior. By building a causal model that entails such a prior for bilateral images, we obtain two optimization goals for counterfactual generation, which can be accomplished via our newly proposed counterfactual generative network. Our proposed model is mainly composed of Generator Adversarial Network and a \emph{prediction feedback mechanism}, they are optimized jointly and prompt each other. Specifically, the former can further improve the classification performance by generating counterfactual features to calculate lesion areas. On the other hand, the latter helps counterfactual generation by the supervision of classification loss. The utility of our method and the effectiveness of each module in our model can be verified by state-of-the-art performance on INBreast and an in-house dataset and ablation studies.
Leveraging both Lesion Features and Procedural Bias in Neuroimaging: An Dual-Task Split dynamics of inverse scale space
Jul 17, 2020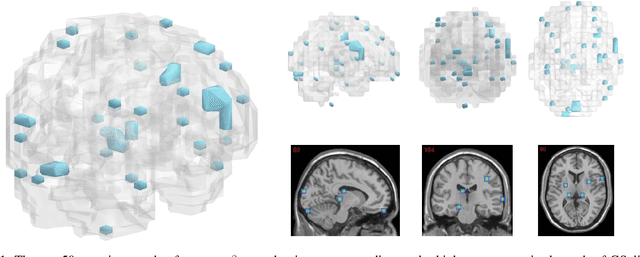
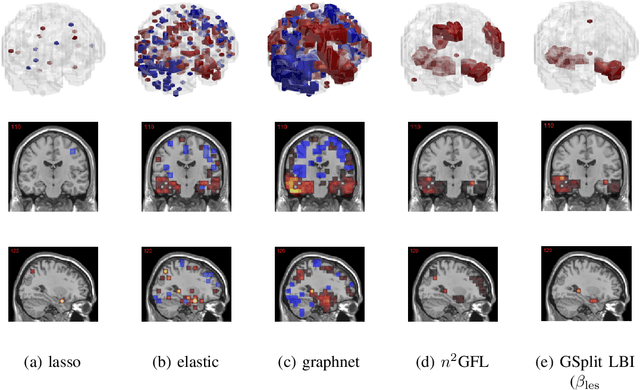
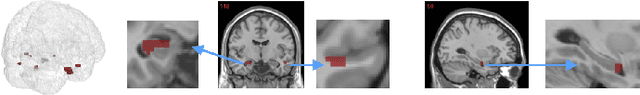
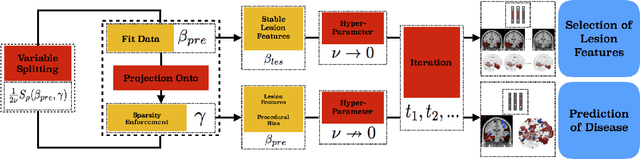
Abstract:The prediction and selection of lesion features are two important tasks in voxel-based neuroimage analysis. Existing multivariate learning models take two tasks equivalently and optimize simultaneously. However, in addition to lesion features, we observe that there is another type of feature, which is commonly introduced during the procedure of preprocessing steps, which can improve the prediction result. We call such a type of feature as procedural bias. Therefore, in this paper, we propose that the features/voxels in neuroimage data are consist of three orthogonal parts: lesion features, procedural bias, and null features. To stably select lesion features and leverage procedural bias into prediction, we propose an iterative algorithm (termed GSplit LBI) as a discretization of differential inclusion of inverse scale space, which is the combination of Variable Splitting scheme and Linearized Bregman Iteration (LBI). Specifically, with a variable the splitting term, two estimators are introduced and split apart, i.e. one is for feature selection (the sparse estimator) and the other is for prediction (the dense estimator). Implemented with Linearized Bregman Iteration (LBI), the solution path of both estimators can be returned with different sparsity levels on the sparse estimator for the selection of lesion features. Besides, the dense the estimator can additionally leverage procedural bias to further improve prediction results. To test the efficacy of our method, we conduct experiments on the simulated study and Alzheimer's Disease Neuroimaging Initiative (ADNI) database. The validity and the benefit of our model can be shown by the improvement of prediction results and the interpretability of visualized procedural bias and lesion features.
TCGM: An Information-Theoretic Framework for Semi-Supervised Multi-Modality Learning
Jul 14, 2020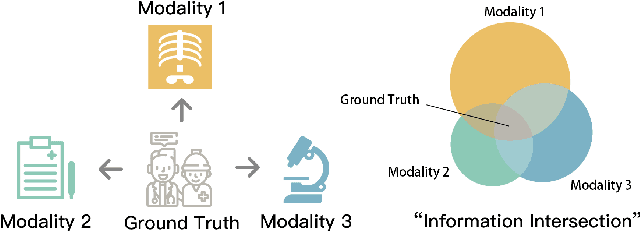
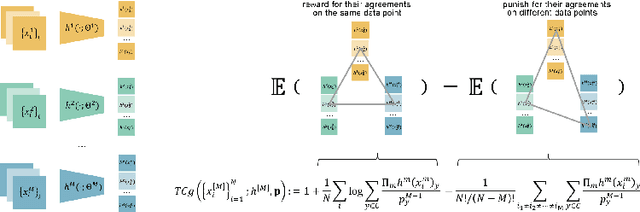
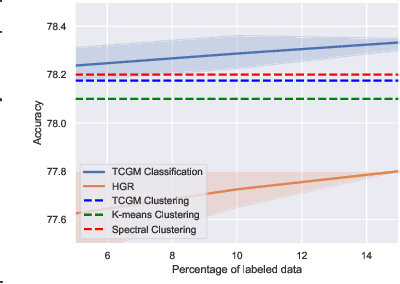
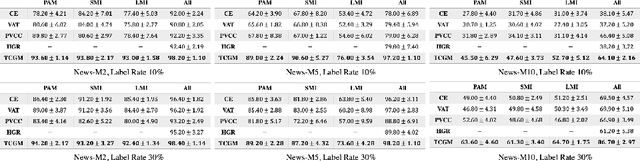
Abstract:Fusing data from multiple modalities provides more information to train machine learning systems. However, it is prohibitively expensive and time-consuming to label each modality with a large amount of data, which leads to a crucial problem of semi-supervised multi-modal learning. Existing methods suffer from either ineffective fusion across modalities or lack of theoretical guarantees under proper assumptions. In this paper, we propose a novel information-theoretic approach, namely \textbf{T}otal \textbf{C}orrelation \textbf{G}ain \textbf{M}aximization (TCGM), for semi-supervised multi-modal learning, which is endowed with promising properties: (i) it can utilize effectively the information across different modalities of unlabeled data points to facilitate training classifiers of each modality (ii) it has theoretical guarantee to identify Bayesian classifiers, i.e., the ground truth posteriors of all modalities. Specifically, by maximizing TC-induced loss (namely TC gain) over classifiers of all modalities, these classifiers can cooperatively discover the equivalent class of ground-truth classifiers; and identify the unique ones by leveraging limited percentage of labeled data. We apply our method to various tasks and achieve state-of-the-art results, including news classification, emotion recognition and disease prediction.
DessiLBI: Exploring Structural Sparsity of Deep Networks via Differential Inclusion Paths
Jul 04, 2020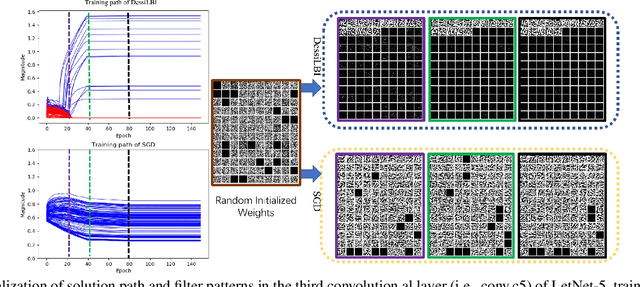
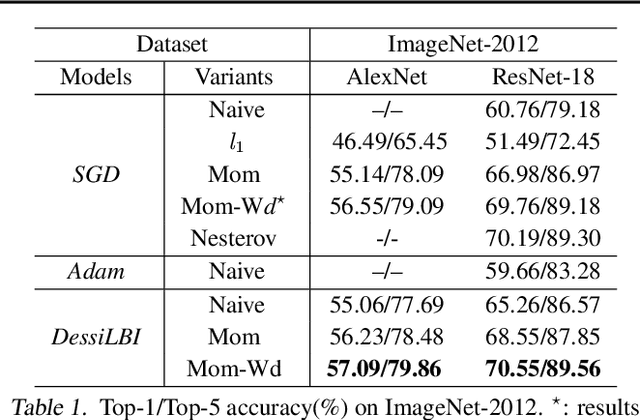

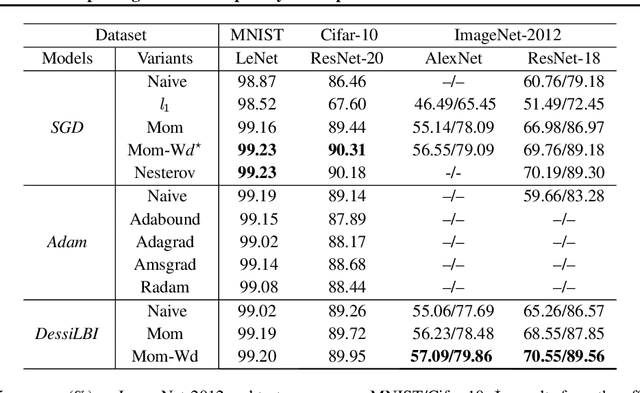
Abstract:Over-parameterization is ubiquitous nowadays in training neural networks to benefit both optimization in seeking global optima and generalization in reducing prediction error. However, compressive networks are desired in many real world applications and direct training of small networks may be trapped in local optima. In this paper, instead of pruning or distilling over-parameterized models to compressive ones, we propose a new approach based on differential inclusions of inverse scale spaces. Specifically, it generates a family of models from simple to complex ones that couples a pair of parameters to simultaneously train over-parameterized deep models and structural sparsity on weights of fully connected and convolutional layers. Such a differential inclusion scheme has a simple discretization, proposed as Deep structurally splitting Linearized Bregman Iteration (DessiLBI), whose global convergence analysis in deep learning is established that from any initializations, algorithmic iterations converge to a critical point of empirical risks. Experimental evidence shows that DessiLBI achieve comparable and even better performance than the competitive optimizers in exploring the structural sparsity of several widely used backbones on the benchmark datasets. Remarkably, with early stopping, DessiLBI unveils "winning tickets" in early epochs: the effective sparse structure with comparable test accuracy to fully trained over-parameterized models.
* conference , 23 pages https://github.com/corwinliu9669/dS2LBI
iSplit LBI: Individualized Partial Ranking with Ties via Split LBI
Oct 14, 2019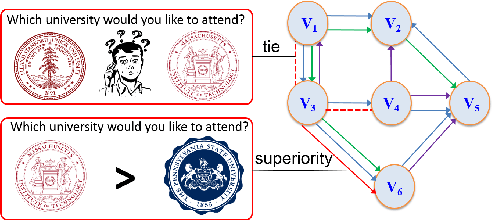
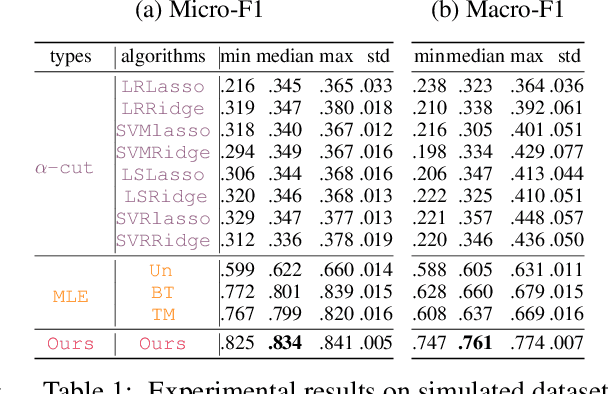
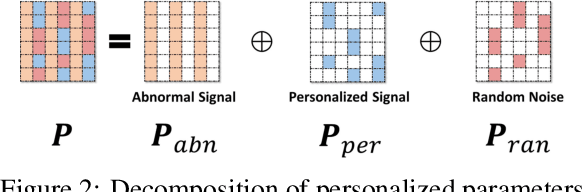
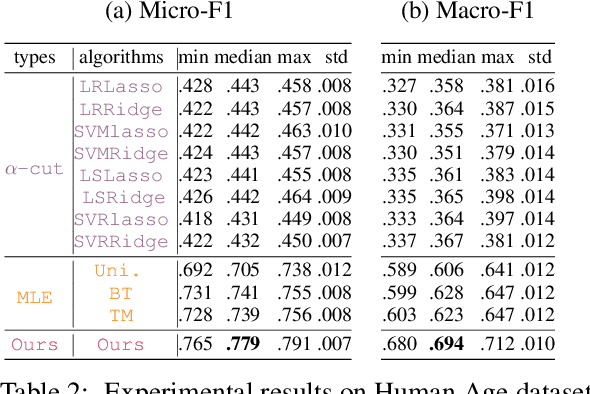
Abstract:Due to the inherent uncertainty of data, the problem of predicting partial ranking from pairwise comparison data with ties has attracted increasing interest in recent years. However, in real-world scenarios, different individuals often hold distinct preferences. It might be misleading to merely look at a global partial ranking while ignoring personal diversity. In this paper, instead of learning a global ranking which is agreed with the consensus, we pursue the tie-aware partial ranking from an individualized perspective. Particularly, we formulate a unified framework which not only can be used for individualized partial ranking prediction, but also be helpful for abnormal user selection. This is realized by a variable splitting-based algorithm called \ilbi. Specifically, our algorithm generates a sequence of estimations with a regularization path, where both the hyperparameters and model parameters are updated. At each step of the path, the parameters can be decomposed into three orthogonal parts, namely, abnormal signals, personalized signals and random noise. The abnormal signals can serve the purpose of abnormal user selection, while the abnormal signals and personalized signals together are mainly responsible for individual partial ranking prediction. Extensive experiments on simulated and real-world datasets demonstrate that our new approach significantly outperforms state-of-the-art alternatives. The code is now availiable at https://github.com/qianqianxu010/NeurIPS2019-iSplitLBI.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge