Xinke Shen
A geometry aware framework enhances noninvasive mapping of whole human brain dynamics
Apr 28, 2026Abstract:Non-invasive electrophysiology lacks methods that accurately reconstruct whole-brain spatiotemporal dynamics while incorporating individual cortical geometry, leaving current electroencephalography and magnetoencephalography source imaging limited by simplistic or biologically implausible priors. Here, we show that embedding participant-specific Geometric Basis Functions (GBFs), eigenmodes derived from each individual's cortical surface, provides a powerful anatomic constraint that resolves the inverse problem and improves reconstruction fidelity. The method reconstructs neural sources as linear combinations of geometric basis functions, thereby aligning source estimates with the geometric organization of neural dynamics. We validate GBF across the Meta-Source Benchmark, task-evoked data, resting-state networks, intracranial stimulation, and epilepsy data. The results demonstrate that GBF yields high localization accuracy and captures fast spatiotemporal dynamics consistent with anatomical pathways. These findings suggest that both spontaneous and evoked whole-brain activity can be described by hundreds of geometric modes, providing a compact yet accurate representation of neural sources. By linking cortical geometry to electrophysiological dynamics, GBF offers a versatile source imaging tool for both scientific and clinical applications.
DSAINet: An Efficient Dual-Scale Attentive Interaction Network for General EEG Decoding
Apr 20, 2026Abstract:In real-world applications of noninvasive electroencephalography (EEG), specialized decoders often show limited generalizability across diverse tasks under subject-independent settings. One central challenge is that task-relevant EEG signals often follow different temporal organization patterns across tasks, while many existing methods rely on task-tailored architectural designs that introduce task-specific temporal inductive biases. This mismatch makes it difficult to adapt temporal modeling across tasks without changing the model configuration. To address these challenges, we propose DSAINet, an efficient dual-scale attentive interaction network for general EEG decoding. Specifically, DSAINet constructs shared spatiotemporal token representations from raw EEG signals and models diverse temporal dynamics through parallel convolutional branches at fine and coarse scales. The resulting representations are then adaptively refined by intra-branch attention to emphasize salient scale-specific patterns and by inter-branch attention to integrate task-relevant features across scales, followed by adaptive token aggregation to yield a compact representation for prediction. Extensive experiments on five downstream EEG decoding tasks across ten public datasets show that DSAINet consistently outperforms 13 representative baselines under strict subject-independent evaluation. Notably, this performance is achieved using the same architecture hyperparameters across datasets. Moreover, DSAINet achieves a favorable accuracy-efficiency trade-off with only about 77K trainable parameters and provides interpretable neurophysiological insights. The code is publicly available at https://github.com/zy0929/DSAINet.
Brain-DiT: A Universal Multi-state fMRI Foundation Model with Metadata-Conditioned Pretraining
Apr 14, 2026Abstract:Current fMRI foundation models primarily rely on a limited range of brain states and mismatched pretraining tasks, restricting their ability to learn generalized representations across diverse brain states. We present \textit{Brain-DiT}, a universal multi-state fMRI foundation model pretrained on 349,898 sessions from 24 datasets spanning resting, task, naturalistic, disease, and sleep states. Unlike prior fMRI foundation models that rely on masked reconstruction in the raw-signal space or a latent space, \textit{Brain-DiT} adopts metadata-conditioned diffusion pretraining with a Diffusion Transformer (DiT), enabling the model to learn multi-scale representations that capture both fine-grained functional structure and global semantics. Across extensive evaluations and ablations on 7 downstream tasks, we find consistent evidence that diffusion-based generative pretraining is a stronger proxy than reconstruction or alignment, with metadata-conditioned pretraining further improving downstream performance by disentangling intrinsic neural dynamics from population-level variability. We also observe that downstream tasks exhibit distinct preferences for representational scale: ADNI classification benefits more from global semantic representations, whereas age/sex prediction comparatively relies more on fine-grained local structure. Code and parameters of Brain-DiT are available at \href{https://github.com/REDMAO4869/Brain-DiT}{Link}.
LI-DSN: A Layer-wise Interactive Dual-Stream Network for EEG Decoding
Apr 02, 2026Abstract:Electroencephalography (EEG) provides a non-invasive window into brain activity, offering high temporal resolution crucial for understanding and interacting with neural processes through brain-computer interfaces (BCIs). Current dual-stream neural networks for EEG often process temporal and spatial features independently through parallel branches, delaying their integration until a final, late-stage fusion. This design inherently leads to an "information silo" problem, precluding intermediate cross-stream refinement and hindering spatial-temporal decompositions essential for full feature utilization. We propose LI-DSN, a layer-wise interactive dual-stream network that facilitates progressive, cross-stream communication at each layer, thereby overcoming the limitations of late-fusion paradigms. LI-DSN introduces a novel Temporal-Spatial Integration Attention (TSIA) mechanism, which constructs a Spatial Affinity Correlation Matrix (SACM) to capture inter-electrode spatial structural relationships and a Temporal Channel Aggregation Matrix (TCAM) to integrate cosine-gated temporal dynamics under spatial guidance. Furthermore, we employ an adaptive fusion strategy with learnable channel weights to optimize the integration of dual-stream features. Extensive experiments across eight diverse EEG datasets, encompassing motor imagery (MI) classification, emotion recognition, and steady-state visual evoked potentials (SSVEP), consistently demonstrate that LI-DSN significantly outperforms 13 state-of-the-art (SOTA) baseline models, showcasing its superior robustness and decoding performance. The code will be publicized after acceptance.
ChineseEEG-2: An EEG Dataset for Multimodal Semantic Alignment and Neural Decoding during Reading and Listening
Aug 06, 2025Abstract:EEG-based neural decoding requires large-scale benchmark datasets. Paired brain-language data across speaking, listening, and reading modalities are essential for aligning neural activity with the semantic representation of large language models (LLMs). However, such datasets are rare, especially for non-English languages. Here, we present ChineseEEG-2, a high-density EEG dataset designed for benchmarking neural decoding models under real-world language tasks. Building on our previous ChineseEEG dataset, which focused on silent reading, ChineseEEG-2 adds two active modalities: Reading Aloud (RA) and Passive Listening (PL), using the same Chinese corpus. EEG and audio were simultaneously recorded from four participants during ~10.7 hours of reading aloud. These recordings were then played to eight other participants, collecting ~21.6 hours of EEG during listening. This setup enables speech temporal and semantic alignment across the RA and PL modalities. ChineseEEG-2 includes EEG signals, precise audio, aligned semantic embeddings from pre-trained language models, and task labels. Together with ChineseEEG, this dataset supports joint semantic alignment learning across speaking, listening, and reading. It enables benchmarking of neural decoding algorithms and promotes brain-LLM alignment under multimodal language tasks, especially in Chinese. ChineseEEG-2 provides a benchmark dataset for next-generation neural semantic decoding.
Dynamic-Attention-based EEG State Transition Modeling for Emotion Recognition
Nov 07, 2024



Abstract:Electroencephalogram (EEG)-based emotion decoding can objectively quantify people's emotional state and has broad application prospects in human-computer interaction and early detection of emotional disorders. Recently emerging deep learning architectures have significantly improved the performance of EEG emotion decoding. However, existing methods still fall short of fully capturing the complex spatiotemporal dynamics of neural signals, which are crucial for representing emotion processing. This study proposes a Dynamic-Attention-based EEG State Transition (DAEST) modeling method to characterize EEG spatiotemporal dynamics. The model extracts spatiotemporal components of EEG that represent multiple parallel neural processes and estimates dynamic attention weights on these components to capture transitions in brain states. The model is optimized within a contrastive learning framework for cross-subject emotion recognition. The proposed method achieved state-of-the-art performance on three publicly available datasets: FACED, SEED, and SEED-V. It achieved 75.4% accuracy in the binary classification of positive and negative emotions and 59.3% in nine-class discrete emotion classification on the FACED dataset, 88.1% in the three-class classification of positive, negative, and neutral emotions on the SEED dataset, and 73.6% in five-class discrete emotion classification on the SEED-V dataset. The learned EEG spatiotemporal patterns and dynamic transition properties offer valuable insights into neural dynamics underlying emotion processing.
A Contrastive Learning Based Convolutional Neural Network for ERP Brain-Computer Interfaces
Jul 02, 2024


Abstract:ERP-based EEG detection is gaining increasing attention in the field of brain-computer interfaces. However, due to the complexity of ERP signal components, their low signal-to-noise ratio, and significant inter-subject variability, cross-subject ERP signal detection has been challenging. The continuous advancement in deep learning has greatly contributed to addressing this issue. This brief proposes a contrastive learning training framework and an Inception module to extract multi-scale temporal and spatial features, representing the subject-invariant components of ERP signals. Specifically, a base encoder integrated with a linear Inception module and a nonlinear projector is used to project the raw data into latent space. By maximizing signal similarity under different targets, the inter-subject EEG signal differences in latent space are minimized. The extracted spatiotemporal features are then used for ERP target detection. The proposed algorithm achieved the best AUC performance in single-trial binary classification tasks on the P300 dataset and showed significant optimization in speller decoding tasks compared to existing algorithms.
Contrastive Learning of Shared Spatiotemporal EEG Representations Across Individuals for Naturalistic Neuroscience
Feb 22, 2024



Abstract:Neural representations induced by naturalistic stimuli offer insights into how humans respond to peripheral stimuli in daily life. The key to understanding the general neural mechanisms underlying naturalistic stimuli processing involves aligning neural activities across individuals and extracting inter-subject shared neural representations. Targeting the Electroencephalogram (EEG) technique, known for its rich spatial and temporal information, this study presents a general framework for Contrastive Learning of Shared SpatioTemporal EEG Representations across individuals (CL-SSTER). Harnessing the representational capabilities of contrastive learning, CL-SSTER utilizes a neural network to maximize the similarity of EEG representations across individuals for identical stimuli, contrasting with those for varied stimuli. The network employed spatial and temporal convolutions to simultaneously learn the spatial and temporal patterns inherent in EEG. The versatility of CL-SSTER was demonstrated on three EEG datasets, including a synthetic dataset, a speech audio EEG dataset, and an emotional video EEG dataset. CL-SSTER attained the highest inter-subject correlation (ISC) values compared to the state-of-the-art ISC methods. The latent representations generated by CL-SSTER exhibited reliable spatiotemporal EEG patterns, which can be explained by specific aspects of the stimuli. CL-SSTER serves as an interpretable and scalable foundational framework for the identification of inter-subject shared neural representations in the realm of naturalistic neuroscience.
Perturbing a Neural Network to Infer Effective Connectivity: Evidence from Synthetic EEG Data
Jul 19, 2023



Abstract:Identifying causal relationships among distinct brain areas, known as effective connectivity, holds key insights into the brain's information processing and cognitive functions. Electroencephalogram (EEG) signals exhibit intricate dynamics and inter-areal interactions within the brain. However, methods for characterizing nonlinear causal interactions among multiple brain regions remain relatively underdeveloped. In this study, we proposed a data-driven framework to infer effective connectivity by perturbing the trained neural networks. Specifically, we trained neural networks (i.e., CNN, vanilla RNN, GRU, LSTM, and Transformer) to predict future EEG signals according to historical data and perturbed the networks' input to obtain effective connectivity (EC) between the perturbed EEG channel and the rest of the channels. The EC reflects the causal impact of perturbing one node on others. The performance was tested on the synthetic EEG generated by a biological-plausible Jansen-Rit model. CNN and Transformer obtained the best performance on both 3-channel and 90-channel synthetic EEG data, outperforming the classical Granger causality method. Our work demonstrated the potential of perturbing an artificial neural network, learned to predict future system dynamics, to uncover the underlying causal structure.
Contrastive Learning of Subject-Invariant EEG Representations for Cross-Subject Emotion Recognition
Sep 20, 2021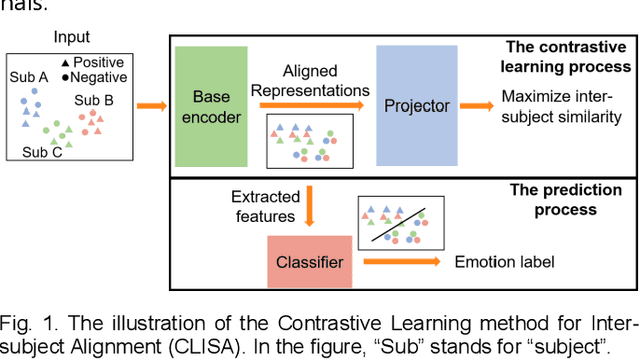
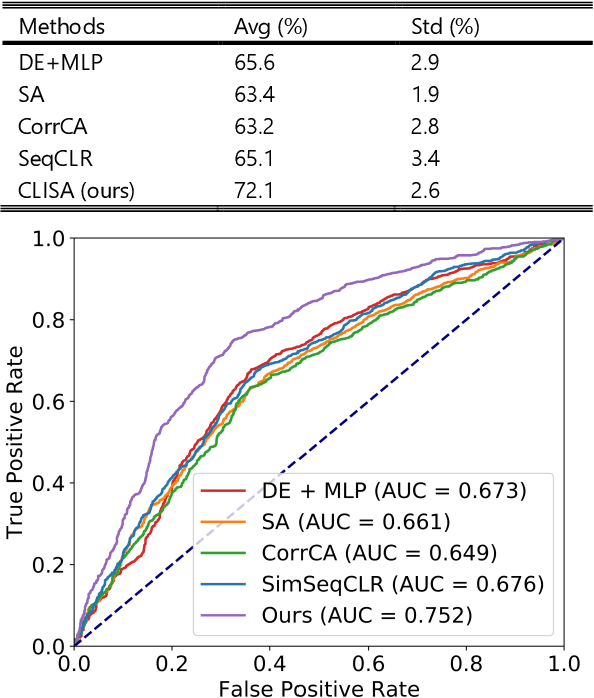
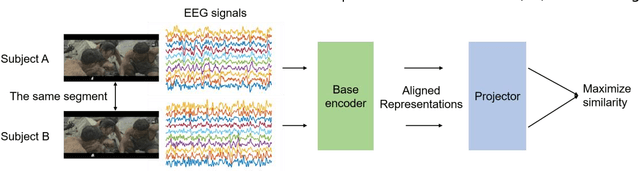
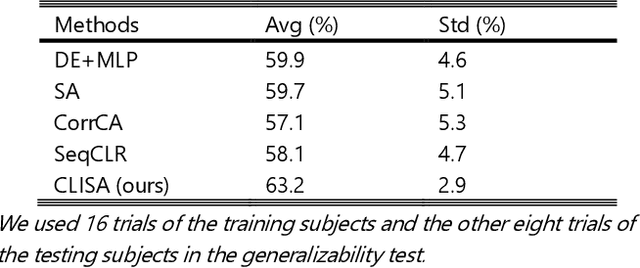
Abstract:Emotion recognition plays a vital role in human-machine interactions and daily healthcare. EEG signals have been reported to be informative and reliable for emotion recognition in recent years. However, the inter-subject variability of emotion-related EEG signals poses a great challenge for the practical use of EEG-based emotion recognition. Inspired by the recent neuroscience studies on inter-subject correlation, we proposed a Contrastive Learning method for Inter-Subject Alignment (CLISA) for reliable cross-subject emotion recognition. Contrastive learning was employed to minimize the inter-subject differences by maximizing the similarity in EEG signals across subjects when they received the same stimuli in contrast to different ones. Specifically, a convolutional neural network with depthwise spatial convolution and temporal convolution layers was applied to learn inter-subject aligned spatiotemporal representations from raw EEG signals. Then the aligned representations were used to extract differential entropy features for emotion classification. The performance of the proposed method was evaluated on our THU-EP dataset with 80 subjects and the publicly available SEED dataset with 15 subjects. Comparable or better cross-subject emotion recognition accuracy (i.e., 72.1% and 47.0% for binary and nine-class classification, respectively, on the THU-EP dataset and 86.3% on the SEED dataset for three-class classification) was achieved as compared to the state-of-the-art methods. The proposed method could be generalized well to unseen emotional stimuli as well. The CLISA method is therefore expected to considerably increase the practicality of EEG-based emotion recognition by operating in a "plug-and-play" manner. Furthermore, the learned spatiotemporal representations by CLISA could provide insights into the neural mechanisms of human emotion processing.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge