Wen Tang
Peking University
Augmenting Clinical Decision-Making with an Interactive and Interpretable AI Copilot: A Real-World User Study with Clinicians in Nephrology and Obstetrics
Jan 31, 2026Abstract:Clinician skepticism toward opaque AI hinders adoption in high-stakes healthcare. We present AICare, an interactive and interpretable AI copilot for collaborative clinical decision-making. By analyzing longitudinal electronic health records, AICare grounds dynamic risk predictions in scrutable visualizations and LLM-driven diagnostic recommendations. Through a within-subjects counterbalanced study with 16 clinicians across nephrology and obstetrics, we comprehensively evaluated AICare using objective measures (task completion time and error rate), subjective assessments (NASA-TLX, SUS, and confidence ratings), and semi-structured interviews. Our findings indicate AICare's reduced cognitive workload. Beyond performance metrics, qualitative analysis reveals that trust is actively constructed through verification, with interaction strategies diverging by expertise: junior clinicians used the system as cognitive scaffolding to structure their analysis, while experts engaged in adversarial verification to challenge the AI's logic. This work offers design implications for creating AI systems that function as transparent partners, accommodating diverse reasoning styles to augment rather than replace clinical judgment.
Proving Olympiad Inequalities by Synergizing LLMs and Symbolic Reasoning
Feb 19, 2025



Abstract:Large language models (LLMs) can prove mathematical theorems formally by generating proof steps (\textit{a.k.a.} tactics) within a proof system. However, the space of possible tactics is vast and complex, while the available training data for formal proofs is limited, posing a significant challenge to LLM-based tactic generation. To address this, we introduce a neuro-symbolic tactic generator that synergizes the mathematical intuition learned by LLMs with domain-specific insights encoded by symbolic methods. The key aspect of this integration is identifying which parts of mathematical reasoning are best suited to LLMs and which to symbolic methods. While the high-level idea of neuro-symbolic integration is broadly applicable to various mathematical problems, in this paper, we focus specifically on Olympiad inequalities (Figure~1). We analyze how humans solve these problems and distill the techniques into two types of tactics: (1) scaling, handled by symbolic methods, and (2) rewriting, handled by LLMs. In addition, we combine symbolic tools with LLMs to prune and rank the proof goals for efficient proof search. We evaluate our framework on 161 challenging inequalities from multiple mathematics competitions, achieving state-of-the-art performance and significantly outperforming existing LLM and symbolic approaches without requiring additional training data.
The Potential of LLMs in Medical Education: Generating Questions and Answers for Qualification Exams
Oct 31, 2024Abstract:Recent research on large language models (LLMs) has primarily focused on their adaptation and application in specialized domains. The application of LLMs in the medical field is mainly concentrated on tasks such as the automation of medical report generation, summarization, diagnostic reasoning, and question-and-answer interactions between doctors and patients. The challenge of becoming a good teacher is more formidable than that of becoming a good student, and this study pioneers the application of LLMs in the field of medical education. In this work, we investigate the extent to which LLMs can generate medical qualification exam questions and corresponding answers based on few-shot prompts. Utilizing a real-world Chinese dataset of elderly chronic diseases, we tasked the LLMs with generating open-ended questions and answers based on a subset of sampled admission reports across eight widely used LLMs, including ERNIE 4, ChatGLM 4, Doubao, Hunyuan, Spark 4, Qwen, Llama 3, and Mistral. Furthermore, we engaged medical experts to manually evaluate these open-ended questions and answers across multiple dimensions. The study found that LLMs, after using few-shot prompts, can effectively mimic real-world medical qualification exam questions, whereas there is room for improvement in the correctness, evidence-based statements, and professionalism of the generated answers. Moreover, LLMs also demonstrate a decent level of ability to correct and rectify reference answers. Given the immense potential of artificial intelligence in the medical field, the task of generating questions and answers for medical qualification exams aimed at medical students, interns and residents can be a significant focus of future research.
ColaCare: Enhancing Electronic Health Record Modeling through Large Language Model-Driven Multi-Agent Collaboration
Oct 03, 2024



Abstract:We introduce ColaCare, a framework that enhances Electronic Health Record (EHR) modeling through multi-agent collaboration driven by Large Language Models (LLMs). Our approach seamlessly integrates domain-specific expert models with LLMs to bridge the gap between structured EHR data and text-based reasoning. Inspired by clinical consultations, ColaCare employs two types of agents: DoctorAgent and MetaAgent, which collaboratively analyze patient data. Expert models process and generate predictions from numerical EHR data, while LLM agents produce reasoning references and decision-making reports within the collaborative consultation framework. We additionally incorporate the Merck Manual of Diagnosis and Therapy (MSD) medical guideline within a retrieval-augmented generation (RAG) module for authoritative evidence support. Extensive experiments conducted on four distinct EHR datasets demonstrate ColaCare's superior performance in mortality prediction tasks, underscoring its potential to revolutionize clinical decision support systems and advance personalized precision medicine. The code, complete prompt templates, more case studies, etc. are publicly available at the anonymous link: https://colacare.netlify.app.
Hunting imaging biomarkers in pulmonary fibrosis: Benchmarks of the AIIB23 challenge
Dec 21, 2023



Abstract:Airway-related quantitative imaging biomarkers are crucial for examination, diagnosis, and prognosis in pulmonary diseases. However, the manual delineation of airway trees remains prohibitively time-consuming. While significant efforts have been made towards enhancing airway modelling, current public-available datasets concentrate on lung diseases with moderate morphological variations. The intricate honeycombing patterns present in the lung tissues of fibrotic lung disease patients exacerbate the challenges, often leading to various prediction errors. To address this issue, the 'Airway-Informed Quantitative CT Imaging Biomarker for Fibrotic Lung Disease 2023' (AIIB23) competition was organized in conjunction with the official 2023 International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI). The airway structures were meticulously annotated by three experienced radiologists. Competitors were encouraged to develop automatic airway segmentation models with high robustness and generalization abilities, followed by exploring the most correlated QIB of mortality prediction. A training set of 120 high-resolution computerised tomography (HRCT) scans were publicly released with expert annotations and mortality status. The online validation set incorporated 52 HRCT scans from patients with fibrotic lung disease and the offline test set included 140 cases from fibrosis and COVID-19 patients. The results have shown that the capacity of extracting airway trees from patients with fibrotic lung disease could be enhanced by introducing voxel-wise weighted general union loss and continuity loss. In addition to the competitive image biomarkers for prognosis, a strong airway-derived biomarker (Hazard ratio>1.5, p<0.0001) was revealed for survival prognostication compared with existing clinical measurements, clinician assessment and AI-based biomarkers.
Predict and Interpret Health Risk using EHR through Typical Patients
Dec 18, 2023Abstract:Predicting health risks from electronic health records (EHR) is a topic of recent interest. Deep learning models have achieved success by modeling temporal and feature interaction. However, these methods learn insufficient representations and lead to poor performance when it comes to patients with few visits or sparse records. Inspired by the fact that doctors may compare the patient with typical patients and make decisions from similar cases, we propose a Progressive Prototypical Network (PPN) to select typical patients as prototypes and utilize their information to enhance the representation of the given patient. In particular, a progressive prototype memory and two prototype separation losses are proposed to update prototypes. Besides, a novel integration is introduced for better fusing information from patients and prototypes. Experiments on three real-world datasets demonstrate that our model brings improvement on all metrics. To make our results better understood by physicians, we developed an application at http://ppn.ai-care.top. Our code is released at https://github.com/yzhHoward/PPN.
M$^3$Fair: Mitigating Bias in Healthcare Data through Multi-Level and Multi-Sensitive-Attribute Reweighting Method
Jun 07, 2023
Abstract:In the data-driven artificial intelligence paradigm, models heavily rely on large amounts of training data. However, factors like sampling distribution imbalance can lead to issues of bias and unfairness in healthcare data. Sensitive attributes, such as race, gender, age, and medical condition, are characteristics of individuals that are commonly associated with discrimination or bias. In healthcare AI, these attributes can play a significant role in determining the quality of care that individuals receive. For example, minority groups often receive fewer procedures and poorer-quality medical care than white individuals in US. Therefore, detecting and mitigating bias in data is crucial to enhancing health equity. Bias mitigation methods include pre-processing, in-processing, and post-processing. Among them, Reweighting (RW) is a widely used pre-processing method that performs well in balancing machine learning performance and fairness performance. RW adjusts the weights for samples within each (group, label) combination, where these weights are utilized in loss functions. However, RW is limited to considering only a single sensitive attribute when mitigating bias and assumes that each sensitive attribute is equally important. This may result in potential inaccuracies when addressing intersectional bias. To address these limitations, we propose M3Fair, a multi-level and multi-sensitive-attribute reweighting method by extending the RW method to multiple sensitive attributes at multiple levels. Our experiments on real-world datasets show that the approach is effective, straightforward, and generalizable in addressing the healthcare fairness issues.
Multi-site, Multi-domain Airway Tree Modeling : A Public Benchmark for Pulmonary Airway Segmentation
Mar 10, 2023



Abstract:Open international challenges are becoming the de facto standard for assessing computer vision and image analysis algorithms. In recent years, new methods have extended the reach of pulmonary airway segmentation that is closer to the limit of image resolution. Since EXACT'09 pulmonary airway segmentation, limited effort has been directed to quantitative comparison of newly emerged algorithms driven by the maturity of deep learning based approaches and clinical drive for resolving finer details of distal airways for early intervention of pulmonary diseases. Thus far, public annotated datasets are extremely limited, hindering the development of data-driven methods and detailed performance evaluation of new algorithms. To provide a benchmark for the medical imaging community, we organized the Multi-site, Multi-domain Airway Tree Modeling (ATM'22), which was held as an official challenge event during the MICCAI 2022 conference. ATM'22 provides large-scale CT scans with detailed pulmonary airway annotation, including 500 CT scans (300 for training, 50 for validation, and 150 for testing). The dataset was collected from different sites and it further included a portion of noisy COVID-19 CTs with ground-glass opacity and consolidation. Twenty-three teams participated in the entire phase of the challenge and the algorithms for the top ten teams are reviewed in this paper. Quantitative and qualitative results revealed that deep learning models embedded with the topological continuity enhancement achieved superior performance in general. ATM'22 challenge holds as an open-call design, the training data and the gold standard evaluation are available upon successful registration via its homepage.
Mortality Prediction with Adaptive Feature Importance Recalibration for Peritoneal Dialysis Patients: a deep-learning-based study on a real-world longitudinal follow-up dataset
Jan 17, 2023



Abstract:Objective: Peritoneal Dialysis (PD) is one of the most widely used life-supporting therapies for patients with End-Stage Renal Disease (ESRD). Predicting mortality risk and identifying modifiable risk factors based on the Electronic Medical Records (EMR) collected along with the follow-up visits are of great importance for personalized medicine and early intervention. Here, our objective is to develop a deep learning model for a real-time, individualized, and interpretable mortality prediction model - AICare. Method and Materials: Our proposed model consists of a multi-channel feature extraction module and an adaptive feature importance recalibration module. AICare explicitly identifies the key features that strongly indicate the outcome prediction for each patient to build the health status embedding individually. This study has collected 13,091 clinical follow-up visits and demographic data of 656 PD patients. To verify the application universality, this study has also collected 4,789 visits of 1,363 hemodialysis dialysis (HD) as an additional experiment dataset to test the prediction performance, which will be discussed in the Appendix. Results: 1) Experiment results show that AICare achieves 81.6%/74.3% AUROC and 47.2%/32.5% AUPRC for the 1-year mortality prediction task on PD/HD dataset respectively, which outperforms the state-of-the-art comparative deep learning models. 2) This study first provides a comprehensive elucidation of the relationship between the causes of mortality in patients with PD and clinical features based on an end-to-end deep learning model. 3) This study first reveals the pattern of variation in the importance of each feature in the mortality prediction based on built-in interpretability. 4) We develop a practical AI-Doctor interaction system to visualize the trajectory of patients' health status and risk indicators.
A Comprehensive Benchmark for COVID-19 Predictive Modeling Using Electronic Health Records in Intensive Care: Choosing the Best Model for COVID-19 Prognosis
Sep 16, 2022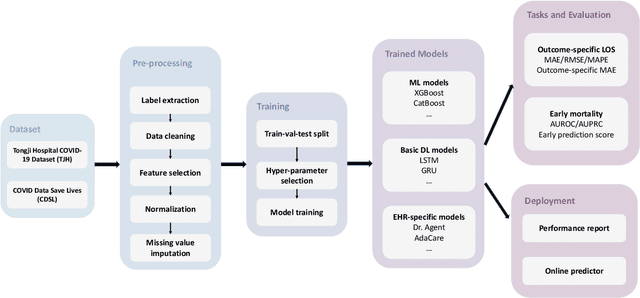
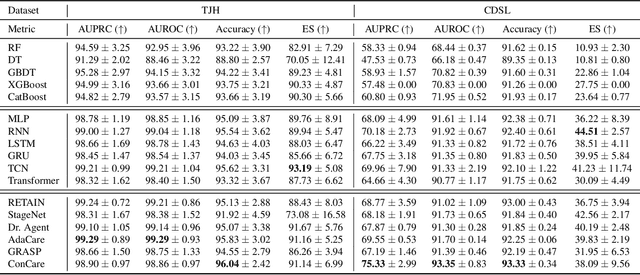
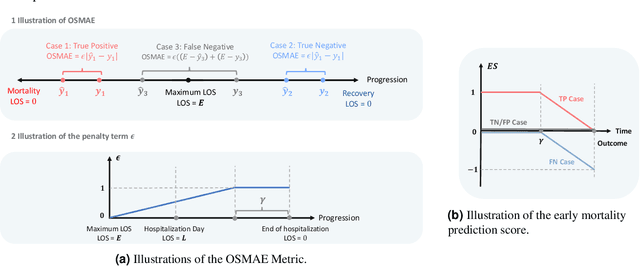
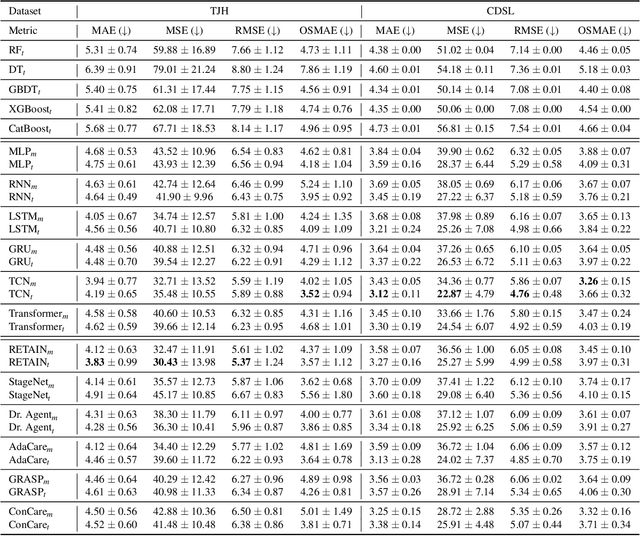
Abstract:The COVID-19 pandemic has posed a heavy burden to the healthcare system worldwide and caused huge social disruption and economic loss. Many deep learning models have been proposed to conduct clinical predictive tasks such as mortality prediction for COVID-19 patients in intensive care units using Electronic Health Record (EHR) data. Despite their initial success in certain clinical applications, there is currently a lack of benchmarking results to achieve a fair comparison so that we can select the optimal model for clinical use. Furthermore, there is a discrepancy between the formulation of traditional prediction tasks and real-world clinical practice in intensive care. To fill these gaps, we propose two clinical prediction tasks, Outcome-specific length-of-stay prediction and Early mortality prediction for COVID-19 patients in intensive care units. The two tasks are adapted from the naive length-of-stay and mortality prediction tasks to accommodate the clinical practice for COVID-19 patients. We propose fair, detailed, open-source data-preprocessing pipelines and evaluate 17 state-of-the-art predictive models on two tasks, including 5 machine learning models, 6 basic deep learning models and 6 deep learning predictive models specifically designed for EHR data. We provide benchmarking results using data from two real-world COVID-19 EHR datasets. Both datasets are publicly available without needing any inquiry and one dataset can be accessed on request. We provide fair, reproducible benchmarking results for two tasks. We deploy all experiment results and models on an online platform. We also allow clinicians and researchers to upload their data to the platform and get quick prediction results using our trained models. We hope our efforts can further facilitate deep learning and machine learning research for COVID-19 predictive modeling.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge