Stefan Bauer
Institute for Surgical Technology and Biomechanics, University of Bern, Switzerland
Dexterous Robotic Manipulation using Deep Reinforcement Learning and Knowledge Transfer for Complex Sparse Reward-based Tasks
May 19, 2022

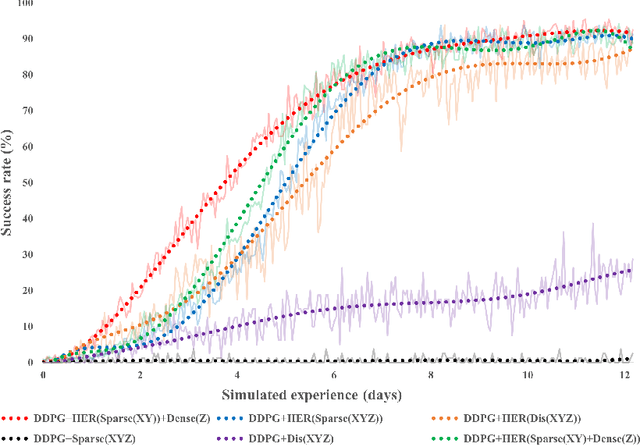
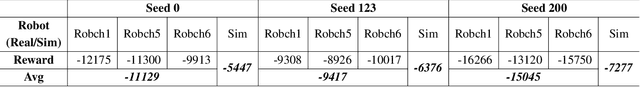
Abstract:This paper describes a deep reinforcement learning (DRL) approach that won Phase 1 of the Real Robot Challenge (RRC) 2021, and then extends this method to a more difficult manipulation task. The RRC consisted of using a TriFinger robot to manipulate a cube along a specified positional trajectory, but with no requirement for the cube to have any specific orientation. We used a relatively simple reward function, a combination of goal-based sparse reward and distance reward, in conjunction with Hindsight Experience Replay (HER) to guide the learning of the DRL agent (Deep Deterministic Policy Gradient (DDPG)). Our approach allowed our agents to acquire dexterous robotic manipulation strategies in simulation. These strategies were then applied to the real robot and outperformed all other competition submissions, including those using more traditional robotic control techniques, in the final evaluation stage of the RRC. Here we extend this method, by modifying the task of Phase 1 of the RRC to require the robot to maintain the cube in a particular orientation, while the cube is moved along the required positional trajectory. The requirement to also orient the cube makes the agent unable to learn the task through blind exploration due to increased problem complexity. To circumvent this issue, we make novel use of a Knowledge Transfer (KT) technique that allows the strategies learned by the agent in the original task (which was agnostic to cube orientation) to be transferred to this task (where orientation matters). KT allowed the agent to learn and perform the extended task in the simulator, which improved the average positional deviation from 0.134 m to 0.02 m, and average orientation deviation from 142{\deg} to 76{\deg} during evaluation. This KT concept shows good generalisation properties and could be applied to any actor-critic learning algorithm.
Federated Learning in Multi-Center Critical Care Research: A Systematic Case Study using the eICU Database
Apr 20, 2022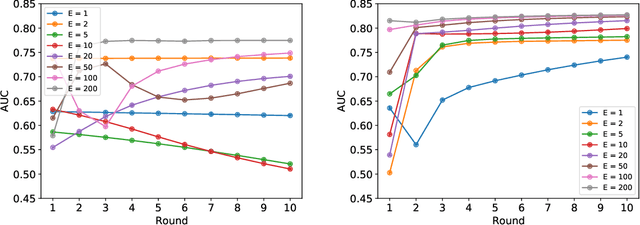
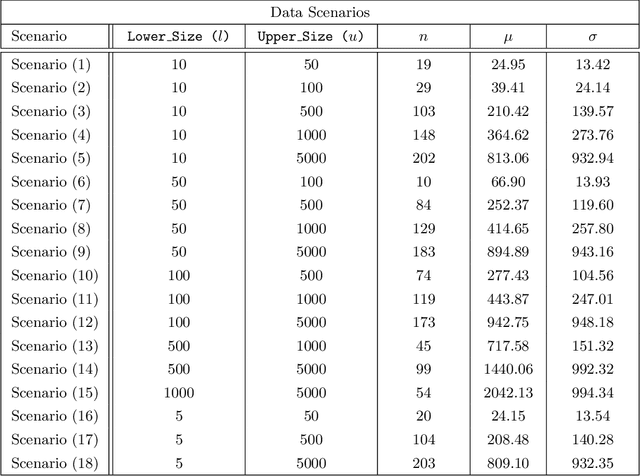
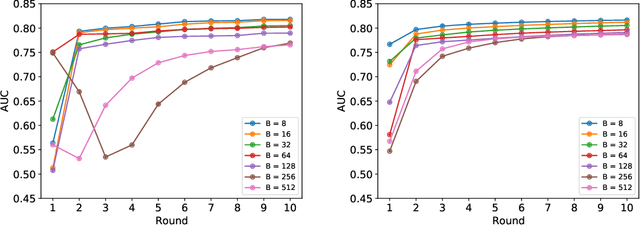
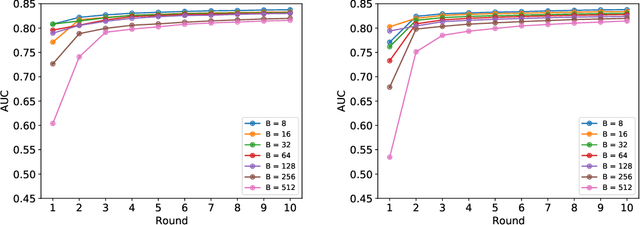
Abstract:Federated learning (FL) has been proposed as a method to train a model on different units without exchanging data. This offers great opportunities in the healthcare sector, where large datasets are available but cannot be shared to ensure patient privacy. We systematically investigate the effectiveness of FL on the publicly available eICU dataset for predicting the survival of each ICU stay. We employ Federated Averaging as the main practical algorithm for FL and show how its performance changes by altering three key hyper-parameters, taking into account that clients can significantly vary in size. We find that in many settings, a large number of local training epochs improves the performance while at the same time reducing communication costs. Furthermore, we outline in which settings it is possible to have only a low number of hospitals participating in each federated update round. When many hospitals with low patient counts are involved, the effect of overfitting can be avoided by decreasing the batchsize. This study thus contributes toward identifying suitable settings for running distributed algorithms such as FL on clinical datasets.
Interventions, Where and How? Experimental Design for Causal Models at Scale
Mar 03, 2022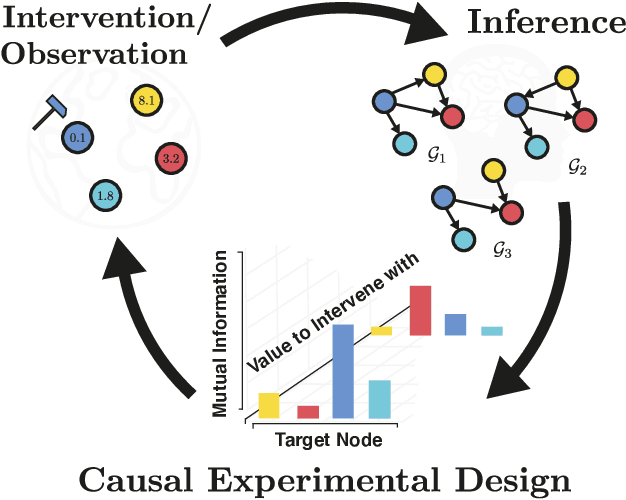
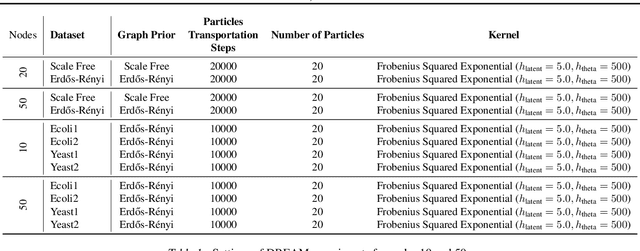
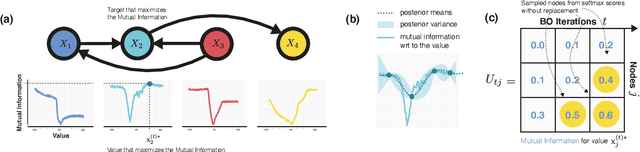
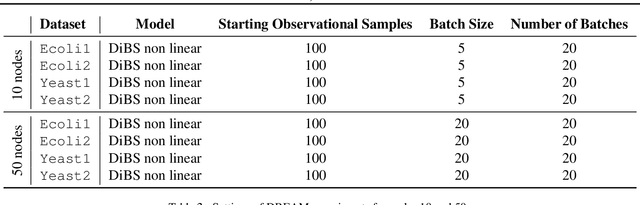
Abstract:Causal discovery from observational and interventional data is challenging due to limited data and non-identifiability which introduces uncertainties in estimating the underlying structural causal model (SCM). Incorporating these uncertainties and selecting optimal experiments (interventions) to perform can help to identify the true SCM faster. Existing methods in experimental design for causal discovery from limited data either rely on linear assumptions for the SCM or select only the intervention target. In this paper, we incorporate recent advances in Bayesian causal discovery into the Bayesian optimal experimental design framework, which allows for active causal discovery of nonlinear, large SCMs, while selecting both the target and the value to intervene with. We demonstrate the performance of the proposed method on synthetic graphs (Erdos-R\`enyi, Scale Free) for both linear and nonlinear SCMs as well as on the in-silico single-cell gene regulatory network dataset, DREAM.
Bayesian Structure Learning with Generative Flow Networks
Feb 28, 2022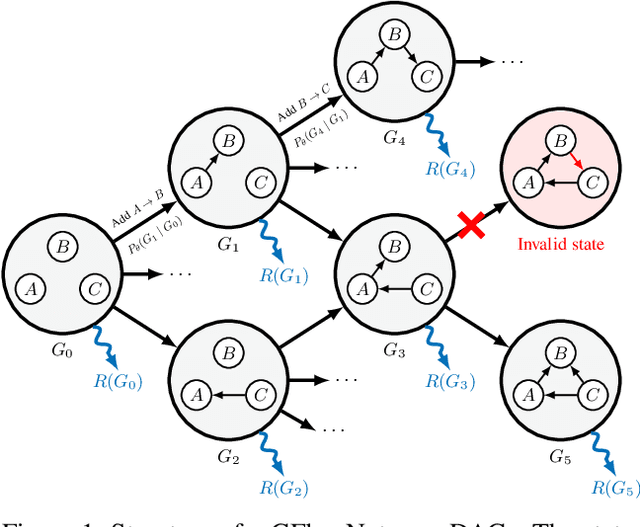
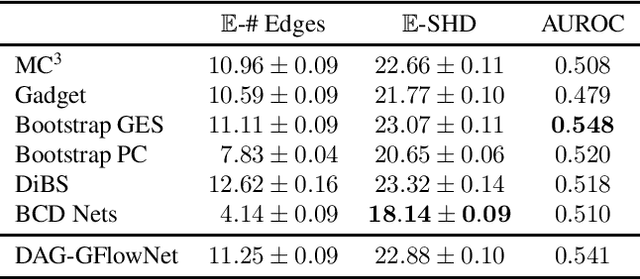
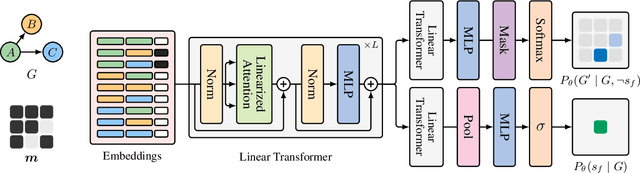
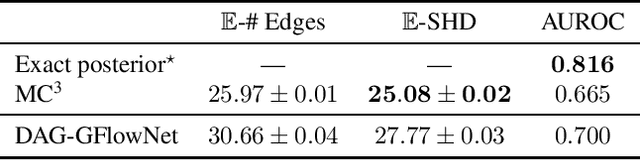
Abstract:In Bayesian structure learning, we are interested in inferring a distribution over the directed acyclic graph (DAG) structure of Bayesian networks, from data. Defining such a distribution is very challenging, due to the combinatorially large sample space, and approximations based on MCMC are often required. Recently, a novel class of probabilistic models, called Generative Flow Networks (GFlowNets), have been introduced as a general framework for generative modeling of discrete and composite objects, such as graphs. In this work, we propose to use a GFlowNet as an alternative to MCMC for approximating the posterior distribution over the structure of Bayesian networks, given a dataset of observations. Generating a sample DAG from this approximate distribution is viewed as a sequential decision problem, where the graph is constructed one edge at a time, based on learned transition probabilities. Through evaluation on both simulated and real data, we show that our approach, called DAG-GFlowNet, provides an accurate approximation of the posterior over DAGs, and it compares favorably against other methods based on MCMC or variational inference.
Compositional Multi-Object Reinforcement Learning with Linear Relation Networks
Jan 31, 2022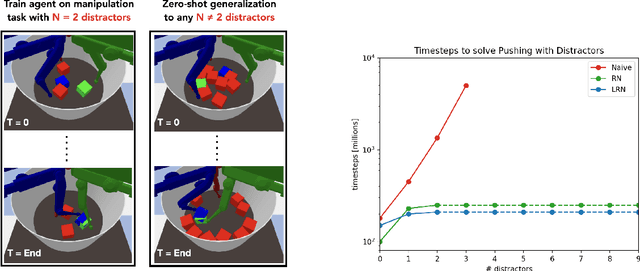
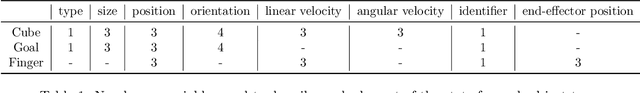

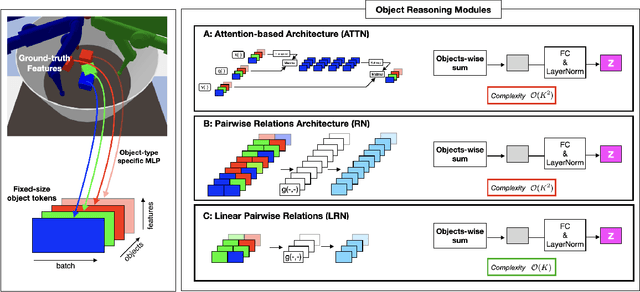
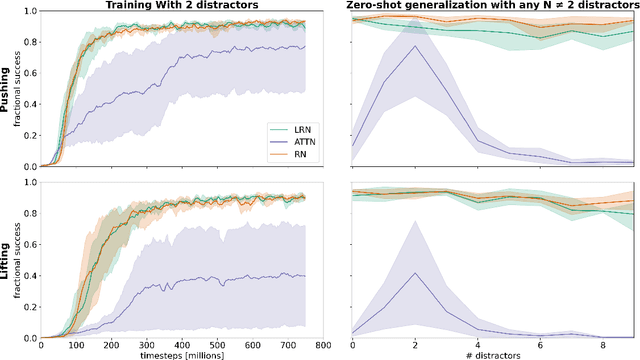
Abstract:Although reinforcement learning has seen remarkable progress over the last years, solving robust dexterous object-manipulation tasks in multi-object settings remains a challenge. In this paper, we focus on models that can learn manipulation tasks in fixed multi-object settings and extrapolate this skill zero-shot without any drop in performance when the number of objects changes. We consider the generic task of bringing a specific cube out of a set to a goal position. We find that previous approaches, which primarily leverage attention and graph neural network-based architectures, do not generalize their skills when the number of input objects changes while scaling as $K^2$. We propose an alternative plug-and-play module based on relational inductive biases to overcome these limitations. Besides exceeding performances in their training environment, we show that our approach, which scales linearly in $K$, allows agents to extrapolate and generalize zero-shot to any new object number.
Conditional Generation of Medical Time Series for Extrapolation to Underrepresented Populations
Jan 20, 2022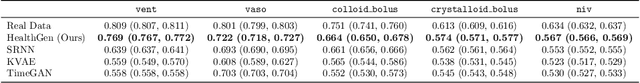
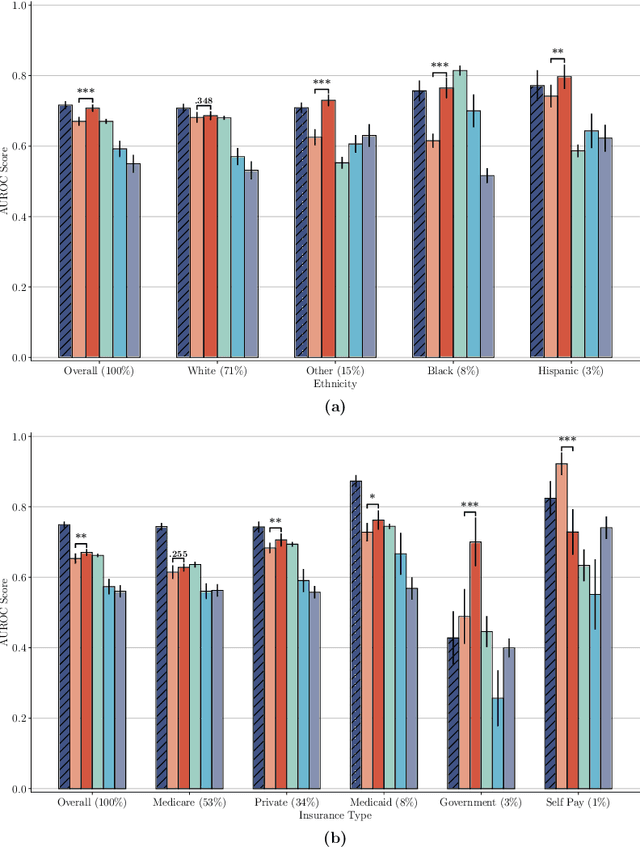
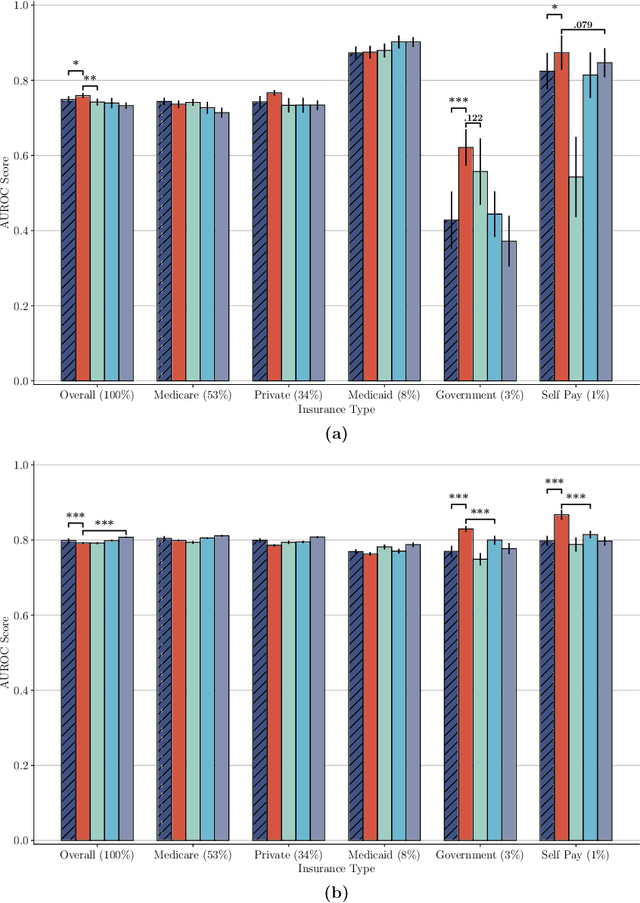
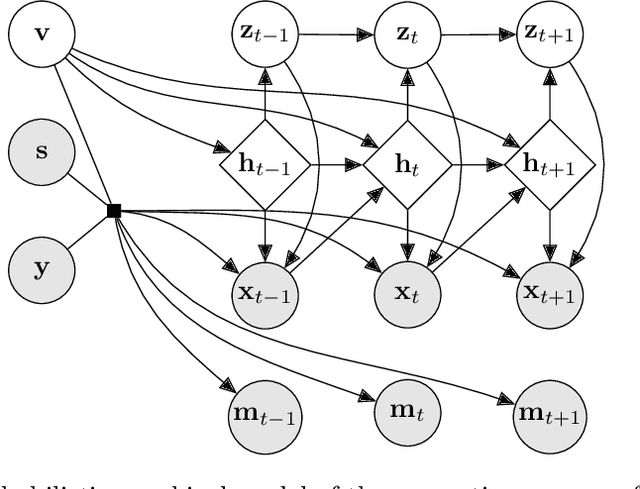
Abstract:The widespread adoption of electronic health records (EHRs) and subsequent increased availability of longitudinal healthcare data has led to significant advances in our understanding of health and disease with direct and immediate impact on the development of new diagnostics and therapeutic treatment options. However, access to EHRs is often restricted due to their perceived sensitive nature and associated legal concerns, and the cohorts therein typically are those seen at a specific hospital or network of hospitals and therefore not representative of the wider population of patients. Here, we present HealthGen, a new approach for the conditional generation of synthetic EHRs that maintains an accurate representation of real patient characteristics, temporal information and missingness patterns. We demonstrate experimentally that HealthGen generates synthetic cohorts that are significantly more faithful to real patient EHRs than the current state-of-the-art, and that augmenting real data sets with conditionally generated cohorts of underrepresented subpopulations of patients can significantly enhance the generalisability of models derived from these data sets to different patient populations. Synthetic conditionally generated EHRs could help increase the accessibility of longitudinal healthcare data sets and improve the generalisability of inferences made from these data sets to underrepresented populations.
Physical Derivatives: Computing policy gradients by physical forward-propagation
Jan 15, 2022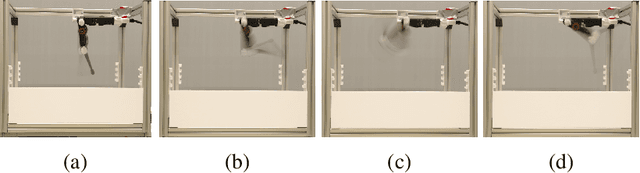
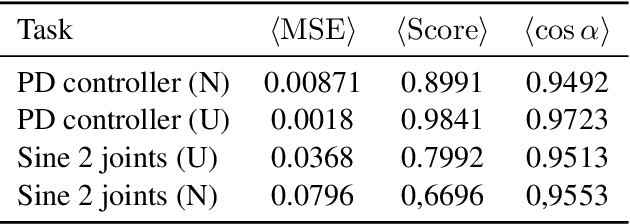
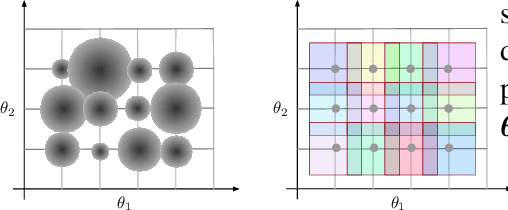
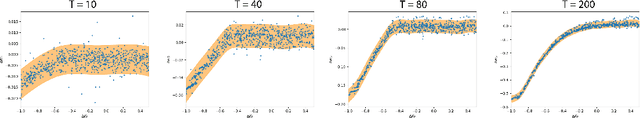
Abstract:Model-free and model-based reinforcement learning are two ends of a spectrum. Learning a good policy without a dynamic model can be prohibitively expensive. Learning the dynamic model of a system can reduce the cost of learning the policy, but it can also introduce bias if it is not accurate. We propose a middle ground where instead of the transition model, the sensitivity of the trajectories with respect to the perturbation of the parameters is learned. This allows us to predict the local behavior of the physical system around a set of nominal policies without knowing the actual model. We assay our method on a custom-built physical robot in extensive experiments and show the feasibility of the approach in practice. We investigate potential challenges when applying our method to physical systems and propose solutions to each of them.
GeneDisco: A Benchmark for Experimental Design in Drug Discovery
Oct 22, 2021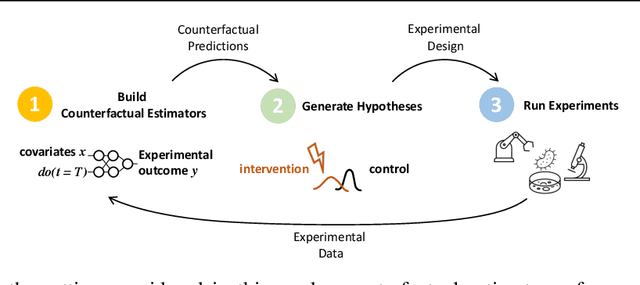
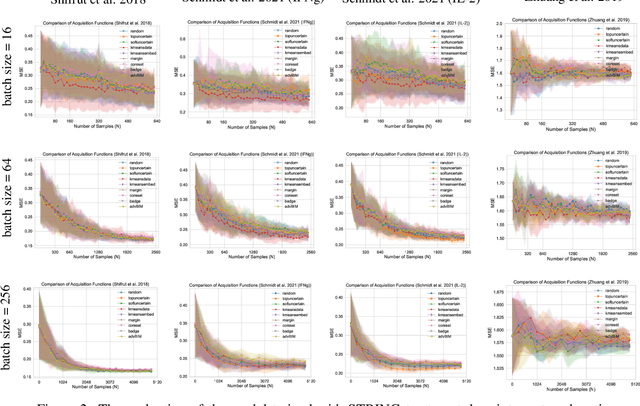
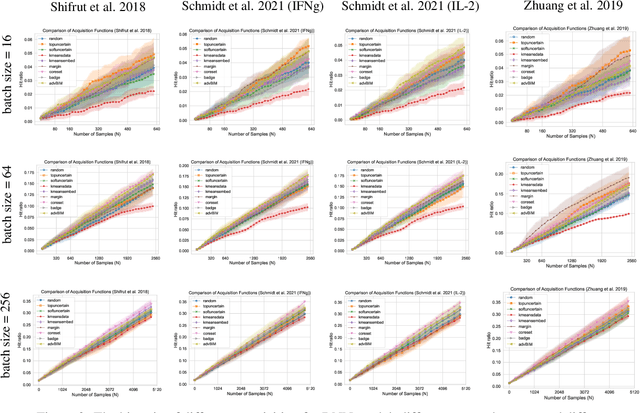
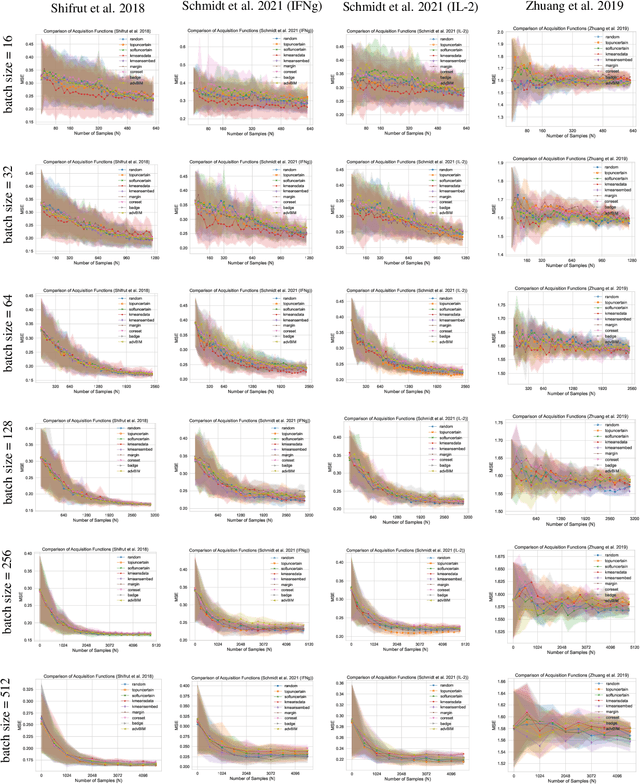
Abstract:In vitro cellular experimentation with genetic interventions, using for example CRISPR technologies, is an essential step in early-stage drug discovery and target validation that serves to assess initial hypotheses about causal associations between biological mechanisms and disease pathologies. With billions of potential hypotheses to test, the experimental design space for in vitro genetic experiments is extremely vast, and the available experimental capacity - even at the largest research institutions in the world - pales in relation to the size of this biological hypothesis space. Machine learning methods, such as active and reinforcement learning, could aid in optimally exploring the vast biological space by integrating prior knowledge from various information sources as well as extrapolating to yet unexplored areas of the experimental design space based on available data. However, there exist no standardised benchmarks and data sets for this challenging task and little research has been conducted in this area to date. Here, we introduce GeneDisco, a benchmark suite for evaluating active learning algorithms for experimental design in drug discovery. GeneDisco contains a curated set of multiple publicly available experimental data sets as well as open-source implementations of state-of-the-art active learning policies for experimental design and exploration.
Boxhead: A Dataset for Learning Hierarchical Representations
Oct 07, 2021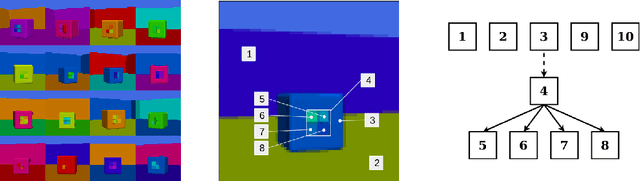
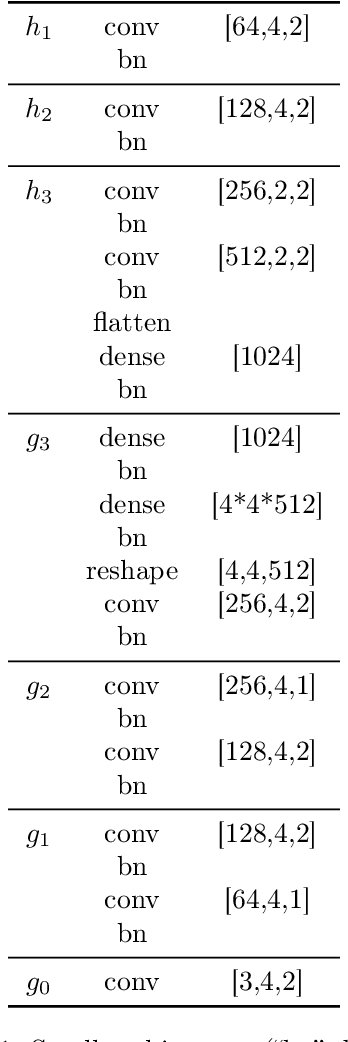
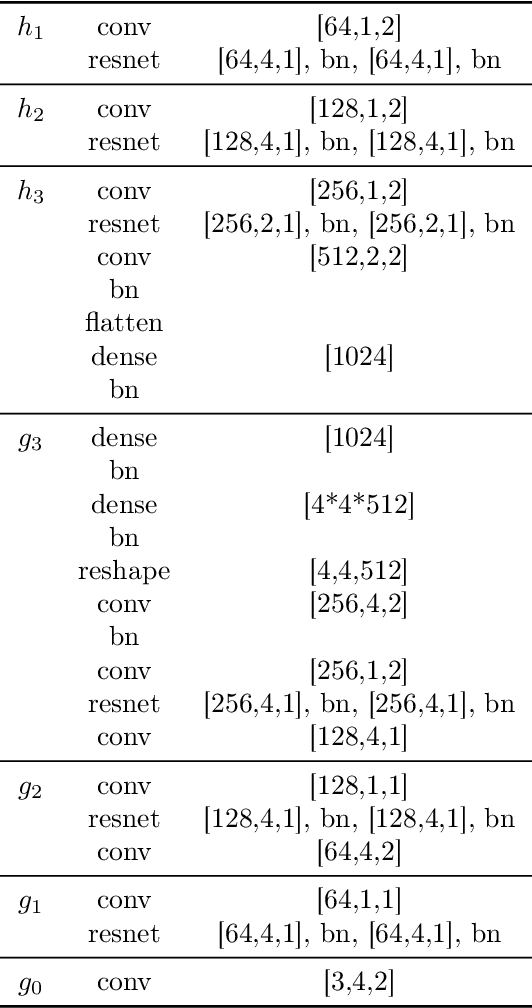

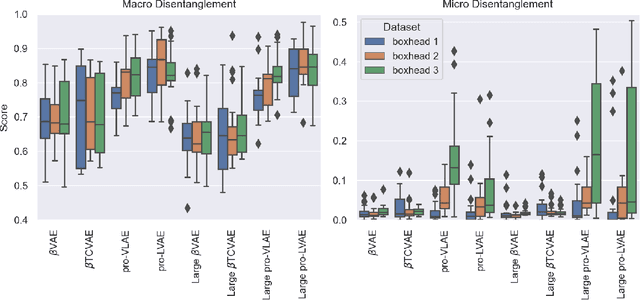

Abstract:Disentanglement is hypothesized to be beneficial towards a number of downstream tasks. However, a common assumption in learning disentangled representations is that the data generative factors are statistically independent. As current methods are almost solely evaluated on toy datasets where this ideal assumption holds, we investigate their performance in hierarchical settings, a relevant feature of real-world data. In this work, we introduce Boxhead, a dataset with hierarchically structured ground-truth generative factors. We use this novel dataset to evaluate the performance of state-of-the-art autoencoder-based disentanglement models and observe that hierarchical models generally outperform single-layer VAEs in terms of disentanglement of hierarchically arranged factors.
A Robot Cluster for Reproducible Research in Dexterous Manipulation
Sep 22, 2021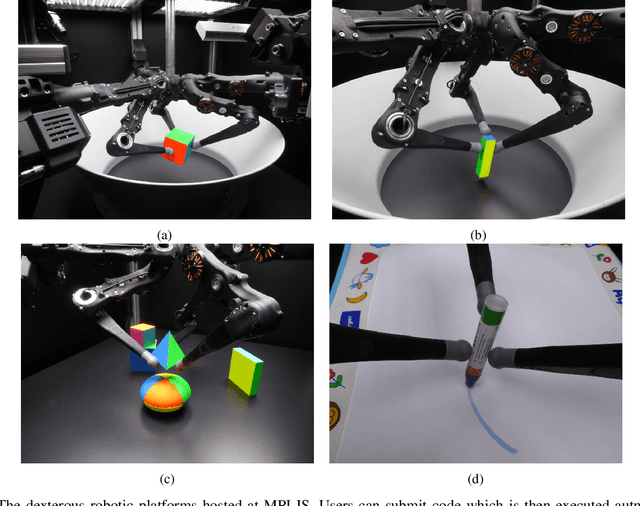
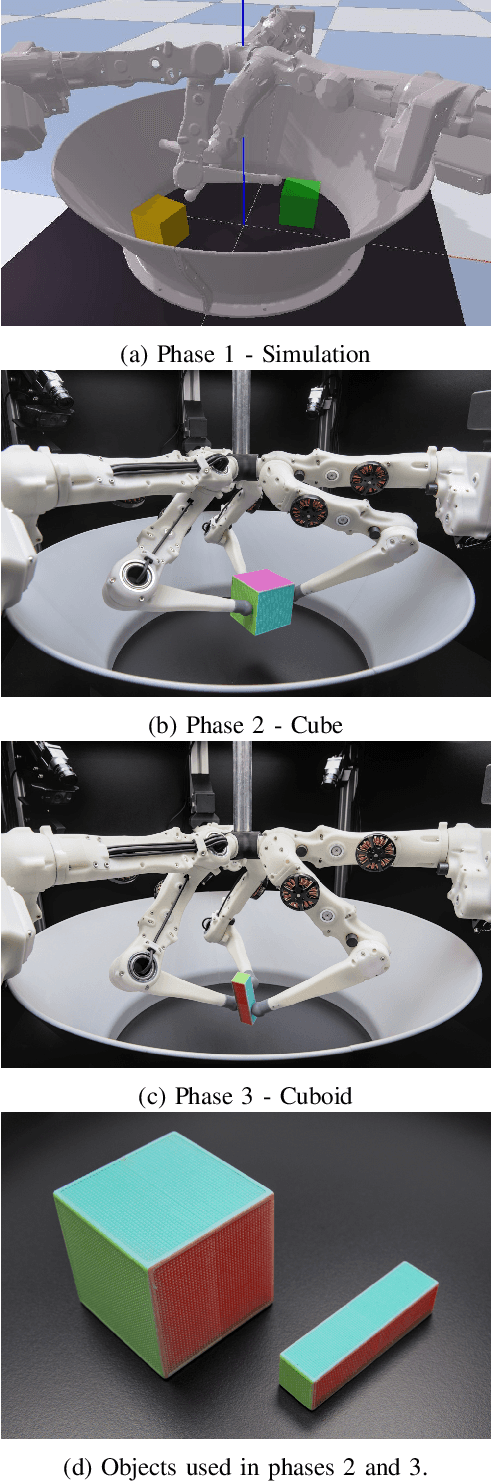

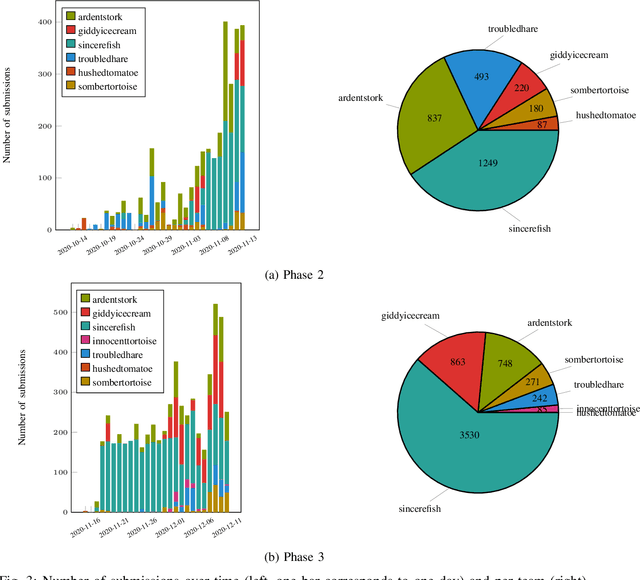
Abstract:Dexterous manipulation remains an open problem in robotics. To coordinate efforts of the research community towards tackling this problem, we propose a shared benchmark. We designed and built robotic platforms that are hosted at the MPI-IS and can be accessed remotely. Each platform consists of three robotic fingers that are capable of dexterous object manipulation. Users are able to control the platforms remotely by submitting code that is executed automatically, akin to a computational cluster. Using this setup, i) we host robotics competitions, where teams from anywhere in the world access our platforms to tackle challenging tasks, ii) we publish the datasets collected during these competitions (consisting of hundreds of robot hours), and iii) we give researchers access to these platforms for their own projects.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge